Abstract
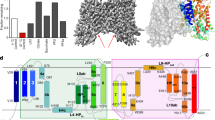
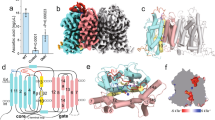
In human cells, cytosolic citrate is a chief precursor for the synthesis of fatty acids, triacylglycerols, cholesterol and low-density lipoprotein. Cytosolic citrate further regulates the energy balance of the cell by activating the fatty-acid-synthesis pathway while downregulating both the glycolysis and fatty-acid β-oxidation pathways1,2,3,4. The rate of fatty-acid synthesis in liver and adipose cells, the two main tissue types for such synthesis, correlates directly with the concentration of citrate in the cytosol2,3,4,5, with the cytosolic citrate concentration partially depending on direct import across the plasma membrane through the Na+-dependent citrate transporter (NaCT)6,7. Mutations of the homologous fly gene (Indy; I’m not dead yet) result in reduced fat storage through calorie restriction8. More recently, Nact (also known as Slc13a5)-knockout mice have been found to have increased hepatic mitochondrial biogenesis, higher lipid oxidation and energy expenditure, and reduced lipogenesis, which taken together protect the mice from obesity and insulin resistance9. To understand the transport mechanism of NaCT and INDY proteins, here we report the 3.2 Å crystal structure of a bacterial INDY homologue. One citrate molecule and one sodium ion are bound per protein, and their binding sites are defined by conserved amino acid motifs, forming the structural basis for understanding the specificity of the transporter. Comparison of the structures of the two symmetrical halves of the transporter suggests conformational changes that propel substrate translocation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Spencer, A. F. & Lowenstein, J. M. Supply of precursors for synthesis of fatty acids. J. Biol. Chem. 237, 3640–3648 (1962)
Bloch, K. & Vance, D. Control mechanisms in synthesis of saturated fatty-acids. Annu. Rev. Biochem. 46, 263–298 (1977)
Ruderman, N. B., Saha, A. K., Vavvas, D. & Witters, L. A. Malonyl-CoA, fuel sensing, and insulin resistance. Am. J. Physiol. 276, E1–E18 (1999)
Sul, H. S. & Smith, S. in Biochemistry of Lipids, Lipoproteins and Membranes 5th edn (eds Vance, D. E. & Vance, J. E. ) 155–190 (Elsevier, 2008)
Nishikori, K., Iritani, N. & Numa, S. Levels of acetyl coenzyme a carboxylase and its effectors in rat-liver after short-term fat-free refeeding. FEBS Lett. 32, 19–21 (1973)
Inoue, K., Zhuang, L., Maddox, D. M., Smith, S. B. & Ganapathy, V. Structure, function, and expression pattern of a novel sodium-coupled citrate transporter (NaCT) cloned from mammalian brain. J. Biol. Chem. 277, 39469–39476 (2002)
Gopal, E. et al. Expression and functional features of NaCT, a sodium-coupled citrate transporter, in human and rat livers and cell lines. Am. J. Physiol. Gastrointest. Liver Physiol. 292, G402–G408 (2007)
Rogina, B., Reenan, R. A., Nilsen, S. P. & Helfand, S. L. Extended life-span conferred by cotransporter gene mutations in Drosophila . Science 290, 2137–2140 (2000)
Birkenfeld, A. L. et al. Deletion of the mammalian INDY homolog mimics aspects of dietary restriction and protects against adiposity and insulin resistance in mice. Cell Metab. 14, 184–195 (2011)
Markovich, D. & Murer, H. The SLC13 gene family of sodium sulphate/carboxylate cotransporters. Pflügers Arch. 447, 594–602 (2004)
Pajor, A. M. Molecular properties of the SLC13 family of dicarboxylate and sulfate transporters. Pflügers Arch. 451, 597–605 (2006)
Wright, S. H., Kippen, I., Klinenberg, J. R. & Wright, E. M. Specificity of the transport system for tricarboxylic acid cycle intermediates in renal brush borders. J. Membr. Biol. 57, 73–82 (1980)
Wright, S. H., Hirayama, B., Kaunitz, J. D., Kippen, I. & Wright, E. M. Kinetics of sodium succinate cotransport across renal brush-border membranes. J. Biol. Chem. 258, 5456–5462 (1983)
Griffith, D. A. & Pajor, A. M. Acidic residues involved in cation and substrate interactions in the Na+/dicarboxylate cotransporter, NaDC-1. Biochemistry 38, 7524–7531 (1999)
Yao, X. & Pajor, A. M. The transport properties of the human renal Na+-dicarboxylate cotransporter under voltage-clamp conditions. Am. J. Physiol. Renal Physiol. 279, F54–F64 (2000)
Pajor, A. M. Conformationally sensitive residues in transmembrane domain 9 of the Na+/dicarboxylate co-transporter. J. Biol. Chem. 276, 29961–29968 (2001)
Joshi, A. D. & Pajor, A. M. Role of conserved prolines in the structure and function of the Na+/dicarboxylate cotransporter 1, NaDC1. Biochemistry 45, 4231–4239 (2006)
Prakash, S., Cooper, G., Singhi, S. & Saier, M. H. The ion transporter superfamily. Biochim. Biophys. Acta. 1618, 79–92 (2003)
Hall, J. A. & Pajor, A. M. Functional characterization of a Na+-coupled dicarboxylate carrier protein from Staphylococcus aureus . J. Bacteriol. 187, 5189–5194 (2005)
Hall, J. A. & Pajor, A. M. Functional reconstitution of SdcS, a Na+-coupled dicarboxylate carrier protein from Staphylococcus aureus . J. Bacteriol. 189, 880–885 (2007)
Youn, J. W., Jolkver, E., Kramer, R., Marin, K. & Wendisch, V. F. Identification and characterization of the dicarboxylate uptake system DccT in Corynebacterium glutamicum . J. Bacteriol. 190, 6458–6466 (2008)
Strickler, M. A., Hall, J. A., Gaiko, O. & Pajor, A. M. Functional characterization of a Na+-coupled dicarboxylate transporter from Bacillus licheniformis . Biochim. Biophys. Acta 1788, 2489–2496 (2009)
Liu, Q., Zhang, Z. & Hendrickson, W. A. Multi-crystal anomalous diffraction for low-resolution macromolecular phasing. Acta Crystallogr. D 67, 45–59 (2011)
Zhang, F. F. & Pajor, A. M. Topology of the Na+/dicarboxylate cotransporter: the N-terminus and hydrophilic loop 4 are located intracellularly. Biochim. Biophys. Acta 1511, 80–89 (2001)
Hunte, C. et al. Structure of a Na+/H+ antiporter and insights into mechanism of action and regulation by pH. Nature 435, 1197–1202 (2005)
Yamashita, A., Singh, S. K., Kawate, T., Jin, Y. & Gouaux, E. Crystal structure of a bacterial homologue of Na+/Cl−-dependent neurotransmitter transporters. Nature 437, 215–223 (2005)
Faham, S. et al. The crystal structure of a sodium galactose transpoter reveals mechanic insights into Na+/sugar symport. Science 321, 810–814 (2008)
Johnson, Z. L., Cheong, C. G. & Lee, S. Y. Crystal structure of a concentrative nucleoside transporter from Vibrio cholerae at 2.4 Å. Nature 483, 489–493 (2012)
Inoue, K., Zhuang, L., Maddox, D. M., Smith, S. B. & Ganapathy, V. Human sodium-coupled citrate transporter, the orthologue of Drosophila Indy, as a novel target for lithium action. Biochem. J. 374, 21–26 (2003)
Pajor, A. M. & Sun, N. N. Single nucleotide polymorphisms in the human Na+-dicarboxylate cotransporter affect transport activity and protein expression. Am. J. Physiol. Renal Physiol. 299, F704–F711 (2010)
Auer, M. et al. High-yield expression and functional analysis of Escherichia coli glycerol-3-phosphate transporter. Biochemistry 40, 6628–6635 (2001)
Li, X. D. et al. Monomeric state and ligand binding of recombinant GABA transporter from Escherichia coli . FEBS Lett. 494, 165–169 (2001)
Wang, D. N. et al. Practical aspects of overexpressing bacterial secondary membrane transporters for structural studies. Biochim. Biophys. Acta 1610, 23–36 (2003)
Love, J. et al. The New York Consortium on Membrane Protein Structure (NYCOMPS): a high-throughput platform for structural genomics of integral membrane proteins. J. Struct. Funct. Genomics 11, 191–199 (2010)
Safferling, M. et al. The TetL tetracycline efflux protein from Bacillus subtilis is a dimer in the membrane and in detergent solution. Biochemistry 42, 13969–13976 (2003)
Boulter, J. M. & Wang, D. N. Purification and characterization of human erythrocyte glucose transporter in decylmaltoside detergent solution. Protein Expr. Purif. 22, 337–348 (2001)
Hirato, T., Shinagawa, M., Ishiguro, N. & Sato, G. Polypeptide involved in the Escherichia coli plasmid-mediated citrate transport system. J. Bacteriol. 160, 421–426 (1984)
Law, C. J., Yang, Q., Soudant, C., Maloney, P. C. & Wang, D. N. Kinetic evidence is consistent with the rocker-switch mechanism of membrane transport by GlpT. Biochemistry 46, 12190–12197 (2007)
Law, C. J. et al. Salt-bridge dynamics control substrate-induced conformational change in the membrane transporter GlpT. J. Mol. Biol. 378, 828–839 (2008)
Liu, Q. et al. Structures from anomalous diffraction of native biological macromolecules. Science 336, 1033–1037 (2012)
Kabsch, W. Xds. Acta Crystallogr. D 66, 125–132 (2010)
Evans, P. R. An introduction to data reduction: space-group determination, scaling and intensity statistics. Acta Crystallogr. D 67, 282–292 (2011)
Sheldrick, G. M. Experimental phasing with SHELXC/D/E: combining chain tracing with density modification. Acta Crystallogr. D 66, 479–485 (2010)
Read, R. J. & McCoy, A. J. Using SAD data in Phaser . Acta Crystallogr. D 67, 338–344 (2011)
Cowtan, K. D. & Zhang, K. Y. Density modification for macromolecular phase improvement. Prog. Biophys. Mol. Biol. 72, 245–270 (1999)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
DeLano, W. L. The PyMOL User’s Manual (DeLano Scientific, 2002)
Acknowledgements
We are grateful to M. Punta and B. Rost for bioinformatics analysis of membrane transporters, to J. Love and B. Kloss for assistance in cloning, and to the staff at beamlines X4, X25 and X29 of the National Synchrotron Light Source in the Brookhaven National Laboratory and at the 23ID at the Advanced Photon Source at the Argonne National Laboratory for assistance in X-ray diffraction experiments, and to J. Llodra for help with artwork. We thank B. K. Czyzewski, W. A. Hendrickson, N. K. Karpowich, F. Mancia and J. J. Marden for discussions and for participating in synchrotron trips. This work was financially supported by National Institutes of Health grants U54-GM075026, R01-DK073973, R01-GM093825 and R01-MH083840.
Author information
Authors and Affiliations
Contributions
R.M. and D.-N.W. designed the project. R.M. did all the experiments, with assistance from G.G.G. in diffraction data processing, phasing and structure refinement, and from Q.L. in phasing. R.M. and D.-N.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion, Supplementary References, Supplementary Tables 1-3 and Supplementary Figures 1-17. (PDF 11214 kb)
Rights and permissions
About this article
Cite this article
Mancusso, R., Gregorio, G., Liu, Q. et al. Structure and mechanism of a bacterial sodium-dependent dicarboxylate transporter. Nature 491, 622–626 (2012). https://doi.org/10.1038/nature11542
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11542
This article is cited by
-
Role of sodium dependent SLC13 transporter inhibitors in various metabolic disorders
Molecular and Cellular Biochemistry (2023)
-
Lsi2: A black box in plant silicon transport
Plant and Soil (2021)
-
Energy landscapes and dynamics of ion translocation through membrane transporters: a meeting ground for physics, chemistry, and biology
Journal of Biological Physics (2021)
-
Undaria pinnatifida improves obesity-related outcomes in association with gut microbiota and metabolomics modulation in high-fat diet-fed mice
Applied Microbiology and Biotechnology (2020)
-
Mitochondrial Citrate Transport System in the Fungus Mucor circinelloides: Identification, Phylogenetic Analysis, and Expression Profiling During Growth and Lipid Accumulation
Current Microbiology (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.