Abstract
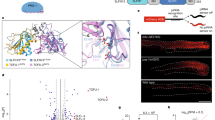
PIWI-interacting RNAs (piRNAs) silence transposons to maintain genome integrity in animal germ lines1,2,3,4. piRNAs are classified as primary and secondary piRNAs, depending on their biogenesis machinery5,6,7,8,9,10. Primary piRNAs are processed from long non-coding RNA precursors transcribed from piRNA clusters in the genome through the primary processing pathway5,8,9,10. Although the existence of a ribonuclease participating in this pathway has been predicted, its molecular identity remained unknown. Here we show that Zucchini (Zuc), a mitochondrial phospholipase D (PLD) superfamily member11, is an endoribonuclease essential for primary piRNA biogenesis. We solved the crystal structure of Drosophila melanogaster Zuc (DmZuc) at 1.75 Å resolution. The structure revealed that DmZuc has a positively charged, narrow catalytic groove at the dimer interface, which could accommodate a single-stranded, but not a double-stranded, RNA. DmZuc and the mouse homologue MmZuc (also known as Pld6 and MitoPLD)12,13,14 showed endoribonuclease activity for single-stranded RNAs in vitro. The RNA cleavage products bear a 5′-monophosphate group, a hallmark of mature piRNAs. Mutational analyses revealed that the conserved active-site residues of DmZuc are critical for the ribonuclease activity in vitro, and for piRNA maturation and transposon silencing in vivo. We propose a model for piRNA biogenesis in animal germ lines, in which the Zuc endoribonuclease has a key role in primary piRNA maturation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Malone, C. D. & Hannon, G. J. Small RNAs as guardians of the genome. Cell 136, 656–668 (2009)
Siomi, M. C., Sato, K., Pezic, C. & Aravin, A. A. PIWI-interacting small RNAs: the vanguard of genome defence. Nature Rev. Mol. Cell Biol. 12, 246–258 (2011)
Pillai, R. S. & Chuma, S. piRNAs and their involvement in male germline development in mice. Dev. Growth Differ. 54, 78–92 (2012)
Senti, K. A. & Brennecke, J. The piRNA pathway: a fly’s perspective on the guardian of the genome. Trends Genet. 26, 499–509 (2010)
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103 (2007)
Vagin, V. V. et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006)
Gunawardane, L. S. et al. A slicer-mediated mechanism for repeat-associated siRNA 5′ end formation in Drosophila. Science 315, 1587–1590 (2007)
Malone, C. D. et al. Specialized piRNA pathways act in germline and somatic tissues of the Drosophila ovary. Cell 137, 522–535 (2009)
Li, C. et al. Collapse of germline piRNAs in the absence of Argonaute3 reveals somatic piRNAs in flies. Cell 137, 509–521 (2009)
Saito, K. et al. A regulatory circuit for piwi by the large Maf gene traffic jam in Drosophila. Nature 461, 1296–1299 (2009)
Pane, A., Wehr, K. & Schüpbach, T. zucchini and squash encode two putative nucleases required for rasiRNA production in the Drosophila germline. Dev. Cell 12, 851–862 (2007)
Choi, S. Y. et al. A common lipid links Mfn-mediated mitochondrial fusion and SNARE-regulated exocytosis. Nature Cell Biol. 8, 1255–1262 (2006)
Watanabe, T. et al. MITOPLD is a mitochondrial protein essential for nuage formation and piRNA biogenesis in the mouse germline. Dev. Cell 20, 364–375 (2011)
Huang, H. et al. piRNA-associated germline nuage formation and spermatogenesis require MitoPLD profusogenic mitochondrial-surface lipid signaling. Dev. Cell 20, 376–387 (2011)
Saito, K. et al. Roles for the Yb body components Armitage and Yb in primary piRNA biogenesis in Drosophila. Genes Dev. 24, 2493–2498 (2010)
Olivieri, D., Sykora, M. M., Sachidanandam, R., Mechtler, K. & Brennecke, J. An in vivo RNAi assay identifies major genetic and cellular requirements for primary piRNA biogenesis in Drosophila. EMBO J. 29, 3301–3317 (2010)
Ponting, C. P. & Kerr, I. D. A novel family of phospholipase D homologues that includes phospholipid synthases and putative endonucleases: Identification of duplicated repeats and potential active site residues. Protein Sci. 5, 914–922 (1996)
Pohlman, R. F. et al. Genetic and biochemical analysis of an endonuclease encoded by the IncN plasmid pKM101. Nucleic Acids Res. 21, 4867–4872 (1993)
Stuckey, J. A. & Dixon, J. E. Crystal structure of a phospholipase D family member. Nature Struct. Biol. 6, 278–284 (1999)
Leiros, I., Secundo, F., Zambonelli, C., Servi, S. & Hough, E. The first crystal structure of a phospholipase D. Structure 8, 655–667 (2000)
Gottlin, E. B., Rudolph, A. E., Zhao, Y., Matthews, H. R. & Dixon, J. E. Catalytic mechanism of the phospholipase D superfamily proceeds via a covalent phosphohistidine intermediate. Proc. Natl Acad. Sci. USA 95, 9202–9207 (1998)
Zhao, Y., Studkey, J. A., Lohse, D. L. & Dixon, J. E. Expression, characterization, and crystallization of a member of the novel phospholipase D family of phosphodiesterases. Protein Sci. 6, 2655–2658 (1997)
Raae, A. J., Kleppe, R. K. & Kleppe, K. Kinetics and effect of salts and polyamines on T4 polynucleotide ligase. Eur. J. Biochem. 60, 437–443 (1975)
Rotondo, G. & Frendewey, D. Purification and characterization of the Pac1 ribonuclease of Schizosaccharomyces pombe. Nucleic Acids Res. 24, 2377–2386 (1996)
Saito, K. et al. Specific association of Piwi with rasiRNAs derived from retrotransposon and heterochromatic regions in the Drosophila genome. Genes Dev. 20, 2214–2222 (2006)
Fukuhara, S. et al. Expression, purification, crystallization and preliminary X-ray crystallographic analysis of Zucchini from Drosophila melanogaster. Acta Crystallogr. F (in the press)
Hayashi, K. & Kojima, C. pCold-GST vector: a novel cold-shock vector containing GST tag for soluble protein production. Protein Expr. Purif. 62, 120–127 (2008)
Sheldrick, G. M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008)
de La Fortelle, E. & Bricogne, G. SHARP: a maximum likelihood heavy-atom parameter refinement program for the MIR and MAD methods. Methods Enzymol. 276, 472–494 (1997)
Terwilliger, T. C. & Berendzen, J. Automated MAD and MIR structure solution. Acta Crystallogr. D 55, 849–861 (1999)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D 66, 22–25 (2010)
Acknowledgements
We thank A. Kurabayashi, H. Kotani and Y. Tamaki for technical assistance; other members of the Siomi laboratory for discussions and comments on the manuscript; the beamline staff at BL32XU of SPring-8 for assistance in data collection; and T. Suzuki and H. Suga for technical suggestions and comments. This work was supported by a grant from the Japan Society for the Promotion of Science (JSPS) through its ‘Funding Program for World-Leading Innovative R&D on Science and Technology (FIRST) program’ to O.N., by the Core Research for Evolutional Science and Technology (CREST) program ‘The Creation of Basic Medical Technologies to Clarify and Control the Mechanisms Underlying Chronic Inflammation’ of the Japan Science and Technology Agency (JST) to O.N. and M.C.S., and by a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology (MEXT) of Japan to H.N., H.I., H.S., M.C.S. and O.N.. K.S. is supported by the JSPS.
Author information
Authors and Affiliations
Contributions
H.N., H.I., K.S., H.S., M.C.S. and O.N. conceived and designed the experiments and wrote the manuscript. H.N., S.F., L.B., N.M., T.N. and R.I. performed the structural analyses. H.I., K.S. and M.K.K. performed biochemical and biological analyses. K.N. and J.A. performed PLD activity assays. All authors discussed the data and the manuscript. M.C.S. and O.N. supervised all the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1, Supplementary Figures 1-10 and an additional reference. (PDF 4268 kb)
Rights and permissions
About this article
Cite this article
Nishimasu, H., Ishizu, H., Saito, K. et al. Structure and function of Zucchini endoribonuclease in piRNA biogenesis. Nature 491, 284–287 (2012). https://doi.org/10.1038/nature11509
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11509
This article is cited by
-
The burgeoning importance of PIWI-interacting RNAs in cancer progression
Science China Life Sciences (2024)
-
The epigenetic regulatory mechanism of PIWI/piRNAs in human cancers
Molecular Cancer (2023)
-
Themes and variations on piRNA-guided transposon control
Mobile DNA (2023)
-
The emerging role of the piRNA/PIWI complex in respiratory tract diseases
Respiratory Research (2023)
-
An evolutionarily conserved stop codon enrichment at the 5′ ends of mammalian piRNAs
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.