Abstract
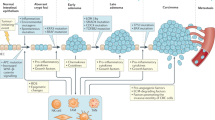
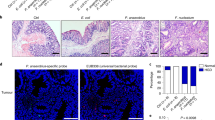
Approximately 2% of colorectal cancer is linked to pre-existing inflammation known as colitis-associated cancer, but most develops in patients without underlying inflammatory bowel disease. Colorectal cancer often follows a genetic pathway whereby loss of the adenomatous polyposis coli (APC) tumour suppressor and activation of β-catenin are followed by mutations in K-Ras, PIK3CA and TP53, as the tumour emerges and progresses1,2. Curiously, however, ‘inflammatory signature’ genes characteristic of colitis-associated cancer are also upregulated in colorectal cancer3,4. Further, like most solid tumours, colorectal cancer exhibits immune/inflammatory infiltrates5, referred to as ‘tumour-elicited inflammation’6. Although infiltrating CD4+ TH1 cells and CD8+ cytotoxic T cells constitute a positive prognostic sign in colorectal cancer7,8, myeloid cells and T-helper interleukin (IL)-17-producing (TH17) cells promote tumorigenesis5,6, and a ‘TH17 expression signature’ in stage I/II colorectal cancer is associated with a drastic decrease in disease-free survival9. Despite its pathogenic importance, the mechanisms responsible for the appearance of tumour-elicited inflammation are poorly understood. Many epithelial cancers develop proximally to microbial communities, which are physically separated from immune cells by an epithelial barrier10. We investigated mechanisms responsible for tumour-elicited inflammation in a mouse model of colorectal tumorigenesis, which, like human colorectal cancer, exhibits upregulation of IL-23 and IL-17. Here we show that IL-23 signalling promotes tumour growth and progression, and development of a tumoural IL-17 response. IL-23 is mainly produced by tumour-associated myeloid cells that are likely to be activated by microbial products, which penetrate the tumours but not adjacent tissue. Both early and late colorectal neoplasms exhibit defective expression of several barrier proteins. We propose that barrier deterioration induced by colorectal-cancer-initiating genetic lesions results in adenoma invasion by microbial products that trigger tumour-elicited inflammation, which in turn drives tumour growth.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
07 November 2012
The FACS image in Fig. 1e was corrected; scale bars in Figs 3b, e and f and 4a, b and f were amended.
References
Vogelstein, B. & Kinzler, K. W. Cancer genes and the pathways they control. Nature Med. 10, 789–799 (2004)
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990)
Wood, L. D. et al. The genomic landscapes of human breast and colorectal cancers. Science 318, 1108–1113 (2007)
Reichling, T. et al. Transcriptional profiles of intestinal tumors in Apc(Min) mice are unique from those of embryonic intestine and identify novel gene targets dysregulated in human colorectal tumors. Cancer Res. 65, 166–176 (2005)
Mantovani, A., Allavena, P., Sica, A. & Balkwill, F. Cancer-related inflammation. Nature 454, 436–444 (2008)
Grivennikov, S. I., Greten, F. R. & Karin, M. Immunity, inflammation, and cancer. Cell 140, 883–899 (2010)
Schreiber, R. D., Old, L. J. & Smyth, M. J. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 331, 1565–1570 (2011)
Galon, J. et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006)
Tosolini, M. et al. Clinical impact of different classes of infiltrating T cytotoxic and helper cells (Th1, th2, treg, th17) in patients with colorectal cancer. Cancer Res. 71, 1263–1271 (2011)
Brenchley, J. M. & Douek, D. C. Microbial translocation across the GI tract. Annu. Rev. Immunol. 30, 149–173 (2012)
McKenzie, B. S., Kastelein, R. A. & Cua, D. J. Understanding the IL-23-IL-17 immune pathway. Trends Immunol. 27, 17–23 (2006)
Langowski, J. L. et al. IL-23 promotes tumour incidence and growth. Nature 442, 461–465 (2006)
Hinoi, T. et al. Mouse model of colonic adenoma-carcinoma progression based on somatic Apc inactivation. Cancer Res. 67, 9721–9730 (2007)
Sawa, S. et al. RORγt+ innate lymphoid cells regulate intestinal homeostasis by integrating negative signals from the symbiotic microbiota. Nature Immunol. 12, 320–326 (2011)
Medzhitov, R. Recognition of microorganisms and activation of the immune response. Nature 449, 819–826 (2007)
Rakoff-Nahoum, S. & Medzhitov, R. Regulation of spontaneous intestinal tumorigenesis through the adaptor protein MyD88. Science 317, 124–127 (2007)
Van der Sluis, M. et al. Muc2-deficient mice spontaneously develop colitis, indicating that MUC2 is critical for colonic protection. Gastroenterology 131, 117–129 (2006)
Velcich, A. et al. Colorectal cancer in mice genetically deficient in the mucin Muc2. Science 295, 1726–1729 (2002)
Rothwell, P. M. et al. Effect of daily aspirin on long-term risk of death due to cancer: analysis of individual patient data from randomised trials. Lancet 377, 31–41 (2011)
Wang, X., Tully, O., Ngo, B., Zitin, M. & Mullin, J. M. Epithelial tight junctional changes in colorectal cancer tissues. Sci. World J. 11, 826–841 (2011)
Tanaka, T. et al. Dextran sodium sulfate strongly promotes colorectal carcinogenesis in Apc(Min/+) mice: inflammatory stimuli by dextran sodium sulfate results in development of multiple colonic neoplasms. Int. J. Cancer 118, 25–34 (2006)
Meira, L. B. et al. DNA damage induced by chronic inflammation contributes to colon carcinogenesis in mice. J. Clin. Invest. 118, 2516–2525 (2008)
Newman, J. V., Kosaka, T., Sheppard, B. J., Fox, J. G. & Schauer, D. B. Bacterial infection promotes colon tumorigenesis in Apc(Min/+) mice. J. Infect. Dis. 184, 227–230 (2001)
Wu, S. et al. A human colonic commensal promotes colon tumorigenesis via activation of T helper type 17 T cell responses. Nature Med. 15, 1016–1022 (2009)
Dove, W. F. et al. Intestinal neoplasia in the ApcMin mouse: independence from the microbial and natural killer (beige locus) status. Cancer Res. 57, 812–814 (1997)
Yang, L. & Pei, Z. Bacteria, inflammation, and colon cancer. World J. Gastroenterol. 12, 6741–6746 (2006)
Garrett, W. S. et al. Colitis-associated colorectal cancer driven by T-bet deficiency in dendritic cells. Cancer Cell 16, 208–219 (2009)
Zhou, L. et al. IL-6 programs T(H)-17 cell differentiation by promoting sequential engagement of the IL-21 and IL-23 pathways. Nature Immunol. 8, 967–974 (2007)
Ivanov, I. I. et al. Specific microbiota direct the differentiation of IL-17-producing T-helper cells in the mucosa of the small intestine. Cell Host Microbe 4, 337–349 (2008)
Barker, N. et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 457, 608–611 (2009)
Lennard-Jones, J. E. Classification of inflammatory bowel disease. Scand. J. Gastroenterol. Suppl. 1702–6 (1989)
Bond, J. H. Polyp guideline: diagnosis, treatment, and surveillance for patients with colorectal polyps. Practice Parameters Committee of the American College of Gastroenterology. Am. J. Gastroenterol. 95, 3053–3063 (2000)
Winawer, S. J. & Zauber, A. G. The advanced adenoma as the primary target of screening. Gastrointest. Endosc. Clin. N. Am. 12, 1–9 (2002)
Liu, P., Jenkins, N. A. & Copeland, N. G. A highly efficient recombineering-based method for generating conditional knockout mutations. Genome Res. 13, 476–484 (2003)
Ghilardi, N. et al. Compromised humoral and delayed-type hypersensitivity responses in IL-23-deficient mice. J. Immunol. 172, 2827–2833 (2004)
Ye, P. et al. Requirement of interleukin 17 receptor signaling for lung CXC chemokine and granulocyte colony-stimulating factor expression, neutrophil recruitment, and host defense. J. Exp. Med. 194, 519–527 (2001)
Awasthi, A. et al. Cutting edge: IL-23 receptor gfp reporter mice reveal distinct populations of IL-17-producing cells. J. Immunol. 182, 5904–5908 (2009)
Kirkland, D. et al. B cell-intrinsic MyD88 signaling prevents the lethal dissemination of commensal bacteria during colonic damage. Immunity 36, 228–238 (2012)
Kawai, T., Adachi, O., Ogawa, T., Takeda, K. & Akira, S. Unresponsiveness of MyD88-deficient mice to endotoxin. Immunity 11, 115–122 (1999)
Hoshino, K. et al. Cutting edge: generation of IL-18 receptor-deficient mice: evidence for IL-1 receptor-related protein as an essential IL-18 binding receptor. J. Immunol. 162, 5041–5044 (1999)
Grivennikov, S. et al. IL-6 and Stat3 are required for survival of intestinal epithelial cells and development of colitis-associated cancer. Cancer Cell 15, 103–113 (2009)
Kim, P. et al. In vivo wide-area cellular imaging by side-view endomicroscopy. Nature Methods 7, 303–305 (2010)
Hung, K. E. et al. Development of a mouse model for sporadic and metastatic colon tumors and its use in assessing drug treatment. Proc. Natl Acad. Sci. USA 107, 1565–1570 (2010)
Chen, G. Y., Shaw, M. H., Redondo, G. & Nunez, G. The innate immune receptor Nod1 protects the intestine from inflammation-induced tumorigenesis. Cancer Res. 68, 10060–10067 (2008)
Acknowledgements
We thank eBioscience, GeneTex, Santa Cruz, BioLegend and Cell Signaling for antibodies; Genentech and Amgen for Il23−/− and Il17ra−/− mice, respectively, and S. Reid and E. Southon for the help in generating Il23rF/F mice. This work was supported by Crohn’s and Colitis Foundation of America (Career Development Award number 2693), NIH/National Institute of Diabetes and Digestive and Kidney Diseases (K99-DK088589) and a University of California, San Diego, Digestive Disease Research Development Center Pilot Grant (DK080506) to S.I.G.; Croucher Foundation and China Postdoctoral Science Foundation (20110490919) to K.W.; Strategic Young Researcher Overseas Visits Program for Accelerating Brain Circulation to K.T.; SPAR Austria to C.D.; NIH (R01CA082223) to E.R.F.; and NIH (AI043477; DK035108) and American Association for Cancer Research (07-60-21-KARI) grants to M.K., who is an American Cancer Society Research Professor. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
S.G. and M.K. conceived the project. S.I.G., K.W., D.M., B.S., D.J., K.T., G.Y.Y., C.O., Y.F. and K.E.H. performed the experiments. S.I.G., K.W., D.M., D.J., H.C., L.E. and M.K. analysed data. C.A.S., V.C., L.T. and G.T. generated Il23rF/F mice. M.O. and F.Y. provided Il23rgfp/gfp and Tlr2,4,9−/− bone marrow, respectively, and Y.F. and E.R.F. provided CPC-APC mice and tissues from Cdx2ERT-Cre-APC mice. C.A.S., E.R.F., H.C. and G.T. provided conceptual advice. C.D. collected and provided human specimens. S.I.G., K.W. and M.K. wrote the manuscript, with all authors contributing to the writing and providing advice.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1 and Supplementary Figures 1-11. Supplementary Figs 1e, 2b and 8a were corrected on 07 November 2012. (PDF 2192 kb)
Rights and permissions
About this article
Cite this article
Grivennikov, S., Wang, K., Mucida, D. et al. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature 491, 254–258 (2012). https://doi.org/10.1038/nature11465
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11465
This article is cited by
-
Network pharmacology prediction and molecular docking-based strategy to explore the potential mechanism of Radix Astragali against hypopharyngeal carcinoma
Scientific Reports (2024)
-
Intratumoural microbiota: a new frontier in cancer development and therapy
Signal Transduction and Targeted Therapy (2024)
-
Deciphering the pharmacological mechanisms of Fraxini Cortex for ulcerative colitis treatment based on network pharmacology and in vivo studies
BMC Complementary Medicine and Therapies (2023)
-
Intratumoural microbiota: from theory to clinical application
Cell Communication and Signaling (2023)
-
Intestinal barrier functions in hematologic and oncologic diseases
Journal of Translational Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.