Abstract
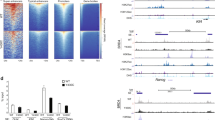
Cornelia de Lange syndrome (CdLS) is a dominantly inherited congenital malformation disorder, caused by mutations in the cohesin-loading protein NIPBL1,2 for nearly 60% of individuals with classical CdLS3,4,5, and by mutations in the core cohesin components SMC1A (∼5%) and SMC3 (<1%) for a smaller fraction of probands6,7. In humans, the multisubunit complex cohesin is made up of SMC1, SMC3, RAD21 and a STAG protein. These form a ring structure that is proposed to encircle sister chromatids to mediate sister chromatid cohesion8 and also has key roles in gene regulation9. SMC3 is acetylated during S-phase to establish cohesiveness of chromatin-loaded cohesin10,11,12,13, and in yeast, the class I histone deacetylase Hos1 deacetylates SMC3 during anaphase14,15,16. Here we identify HDAC8 as the vertebrate SMC3 deacetylase, as well as loss-of-function HDAC8 mutations in six CdLS probands. Loss of HDAC8 activity results in increased SMC3 acetylation and inefficient dissolution of the ‘used’ cohesin complex released from chromatin in both prophase and anaphase. SMC3 with retained acetylation is loaded onto chromatin, and chromatin immunoprecipitation sequencing analysis demonstrates decreased occupancy of cohesin localization sites that results in a consistent pattern of altered transcription seen in CdLS cell lines with either NIPBL or HDAC8 mutations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Sequence Read Archive
Data deposits
ChIP-Seq and RNA-Seq data from this study are available from the Sequence Read Archive (SRA) database (http://www.ncbi.nlm.nih.gov/sra) under the accession number SRP011927.
References
Gillespie, P. J. & Hirano, T. Scc2 couples replication licensing to sister chromatid cohesion in Xenopus egg extracts. Curr. Biol. 14, 1598–1603 (2004)
Takahashi, T. S., Yiu, P., Chou, M. F., Gygi, S. & Walter, J. C. Recruitment of Xenopus Scc2 and cohesin to chromatin requires the pre-replication complex. Nature Cell Biol. 6, 991–996 (2004)
Gillis, L. A. et al. NIPBL mutational analysis in 120 individuals with Cornelia de Lange syndrome and evaluation of genotype-phenotype correlations. Am. J. Hum. Genet. 75, 610–623 (2004)
Krantz, I. D. et al. Cornelia de Lange syndrome is caused by mutations in NIPBL, the human homolog of Drosophila melanogaster Nipped-B. Nature Genet. 36, 631–635 (2004)
Tonkin, E. T., Wang, T. J., Lisgo, S., Bamshad, M. J. & Strachan, T. NIPBL, encoding a homolog of fungal Scc2-type sister chromatid cohesion proteins and fly Nipped-B, is mutated in Cornelia de Lange syndrome. Nature Genet. 36, 636–641 (2004)
Deardorff, M. A. et al. Mutations in cohesin complex members SMC3 and SMC1A cause a mild variant of Cornelia de Lange syndrome with predominant mental retardation. Am. J. Hum. Genet. 80, 485–494 (2007)
Musio, A. et al. X-linked Cornelia de Lange syndrome owing to SMC1L1 mutations. Nature Genet. 38, 528–530 (2006)
Nasmyth, K. & Haering, C. H. Cohesin: its roles and mechanisms. Annu. Rev. Genet. 43, 525–558 (2009)
Dorsett, D. Cohesin: genomic insights into controlling gene transcription and development. Curr. Opin. Genet. Dev. 21, 199–206 (2011)
Rolef Ben-Shahar, T. et al. Eco1-dependent cohesin acetylation during establishment of sister chromatid cohesion. Science 321, 563–566 (2008)
Unal, E. et al. A molecular determinant for the establishment of sister chromatid cohesion. Science 321, 566–569 (2008)
Zhang, B. et al. Dosage effects of cohesin regulatory factor PDS5 on mammalian development: implications for cohesinopathies. PLoS ONE 4, e5232 (2009)
Heidinger-Pauli, J. M., Unal, E. & Koshland, D. Distinct targets of the Eco1 acetyltransferase modulate cohesion in S phase and in response to DNA damage. Mol. Cell 34, 311–321 (2009)
Beckouët, F. et al. An Smc3 acetylation cycle is essential for establishment of sister chromatid cohesion. Mol. Cell 39, 689–699 (2010)
Xiong, B., Lu, S. & Gerton, J. L. Hos1 is a lysine deacetylase for the Smc3 subunit of cohesin. Curr. Biol. 20, 1660–1665 (2010)
Borges, V. et al. Hos1 deacetylates Smc3 to close the cohesin acetylation cycle. Mol. Cell 39, 677–688 (2010)
Zhang, J. et al. Acetylation of Smc3 by Eco1 is required for S phase sister chromatid cohesion in both human and yeast. Mol. Cell 31, 143–151 (2008)
Nishiyama, T. et al. Sororin mediates sister chromatid cohesion by antagonizing Wapl. Cell 143, 737–749 (2010)
Wendt, K. S. et al. Cohesin mediates transcriptional insulation by CCCTC-binding factor. Nature 451, 796–801 (2008)
Liu, J. et al. Transcriptional dysregulation in NIPBL and cohesin mutant human cells. PLoS Biol. 7, e1000119 (2009)
The 1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010)
Vannini, A. et al. Substrate binding to histone deacetylases as shown by the crystal structure of the HDAC8-substrate complex. EMBO Rep. 8, 879–884 (2007)
Dowling, D. P., Gantt, S. L., Gattis, S. G., Fierke, C. A. & Christianson, D. W. Structural studies of human histone deacetylase 8 and its site-specific variants complexed with substrate and inhibitors. Biochemistry 47, 13554–13563 (2008)
Haering, C. H., Lowe, J., Hochwagen, A. & Nasmyth, K. Molecular architecture of SMC proteins and the yeast cohesin complex. Mol. Cell 9, 773–788 (2002)
Waizenegger, I. C., Hauf, S., Meinke, A. & Peters, J. M. Two distinct pathways remove mammalian cohesin from chromosome arms in prophase and from centromeres in anaphase. Cell 103, 399–410 (2000)
Giménez-Abian, J. F. et al. Regulation of sister chromatid cohesion between chromosome arms. Curr. Biol. 14, 1187–1193 (2004)
Gandhi, R., Gillespie, P. J. & Hirano, T. Human Wapl is a cohesin-binding protein that promotes sister-chromatid resolution in mitotic prophase. Curr. Biol. 16, 2406–2417 (2006)
Kueng, S. et al. Wapl controls the dynamic association of cohesin with chromatin. Cell 127, 955–967 (2006)
Schmitz, J., Watrin, E., Lenart, P., Mechtler, K. & Peters, J. M. Sororin is required for stable binding of cohesin to chromatin and for sister chromatid cohesion in interphase. Curr. Biol. 17, 630–636 (2007)
Hauf, S., Waizenegger, I. C. & Peters, J. M. Cohesin cleavage by separase required for anaphase and cytokinesis in human cells. Science 293, 1320–1323 (2001)
Watrin, E. et al. Human Scc4 is required for cohesin binding to chromatin, sister-chromatid cohesion, and mitotic progression. Curr. Biol. 16, 863–874 (2006)
Eddy, S. R. Multiple alignment using hidden Markov models. Proc. Int. Conf. Intell. Syst. Mol. Biol. 3, 114–120 (1995)
Wang, Y., Geer, L. Y., Chappey, C., Kans, J. A. & Bryant, S. H. Cn3D: sequence and structure views for Entrez. Trends Biochem. Sci. 25, 300–302 (2000)
The. PyMOL Molecular Graphics System v. 1.5.0.1 (Schrödinger, LLC, 2012)
Geiss, G. K. et al. Direct multiplexed measurement of gene expression with color-coded probe pairs. Nature Biotechnol. 26, 317–325 (2008)
Gantt, S. L., Gattis, S. G. & Fierke, C. A. Catalytic activity and inhibition of human histone deacetylase 8 is dependent on the identity of the active site metal ion. Biochemistry 45, 6170–6178 (2006)
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 72, 248–254 (1976)
Allen, R. C., Zoghbi, H. Y., Moseley, A. B., Rosenblatt, H. M. & Belmont, J. W. Methylation of HpaII and HhaI sites near the polymorphic CAG repeat in the human androgen-receptor gene correlates with X chromosome inactivation. Am. J. Hum. Genet. 51, 1229–1239 (1992)
Wang, X. et al. Mutations in X-linked PORCN, a putative regulator of Wnt signaling, cause focal dermal hypoplasia. Nature Genet. 39, 836–838 (2007)
Amos-Landgraf, J. M. et al. X chromosome-inactivation patterns of 1,005 phenotypically unaffected females. Am. J. Hum. Genet. 79, 493–499 (2006)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Katou, Y. et al. S-phase checkpoint proteins Tof1 and Mrc1 form a stable replication-pausing complex. Nature 424, 1078–1083 (2003)
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nature Methods 5, 621–628 (2008)
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010)
Acknowledgements
We are grateful to the individuals and families with Cornelia de Lange syndrome who participated in this study, as well as to the referring physicians and colleagues who have contributed samples and clinical information. We thank Riken Omics Science Center, K. Nakagawa, S. Watanabe, M. Albrecht and J. Eckhold for technical support. We thank J.-M. Peters for the sororin and RAD21 antibodies. We thank F. Beckouët and K. Nasmyth for sharing unpublished results. We are indebted to the continued support of the US and International Cornelia de Lange Syndrome Foundations. This work was supported by National Institutes of Health grants K08HD055488 (NICHD, M.A.D.), GM49758 (D.W.C.) and P01 HD052860 (NICHD; I.D.K.), research grants from the US CdLS Foundation, institutional funds from the Children’s Hospital of Philadelphia, Intramural funding from the University of Lübeck (Schwerpunktprogramm, Medizinische Genetik: Von seltenen Varianten zur Krankheitsentstehung; F.J.K., G.G.-K.), Research Program of Innovative Cell Biology by Innovative Technology, Grant-in-Aid for Scientific Research (S) and for innovative science from MEXT (K.Sh.).
Author information
Authors and Affiliations
Contributions
M.B. and K.Sh. designed and performed the biochemical analyses; M.B., E.W., C.J., L.M.-J. and K.Sh. performed HDAC screening. M.B., K.Sa., E.W. and L.M.-J. performed chromatin spreading assays; N.N. monoclonal isolation; M.B., R.N., T.I., M.K., Y.K. and K.Sh. the ChIP sequencing and analysis; M.B., R.N. and Y.S. RNA sequencing; M.M., K.T. and T.H. microscopy. M.A.D., G.G.-K., L.G.J., F.J.K. and I.D.K. initiated the human studies. M.A.D., D.C., E.D.B., G.R.M., V.M.S., P.W., N.T., Y.G., M.B.P., G.G.-K. and I.D.K. identified and characterized clinical data. M.A.D., M.K., S.E., L.J.F., M.H. and F.J.K. performed mutation screening, inactivation and lymphoblastoid expression studies. K.H., T.K. and H.S. established fibroblast cell lines. M.A.D., J.J.W., K.E.C., P.M.L., C.D. and D.W.C. performed enzymatic and structural analysis. M.A.D., M.B. and K.Sh. drafted the manuscript. All authors analysed data, discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-10, a Supplementary Discussion, Supplementary Tables 1-6 and 8-9, Supplementary References and full scan images for all Western-blotting data presented in both the main article figures and supplementary figures. (PDF 5189 kb)
Supplementary Data
This file contains Nanostring data and calculations for 32-gene CdLS lymphoblastoid expression profiling. (XLS 174 kb)
Supplementary Data
This file contains Supplementary Table 7, which shows RNA sequencing data from Fibroblast experiments. (XLS 1594 kb)
Rights and permissions
About this article
Cite this article
Deardorff, M., Bando, M., Nakato, R. et al. HDAC8 mutations in Cornelia de Lange syndrome affect the cohesin acetylation cycle. Nature 489, 313–317 (2012). https://doi.org/10.1038/nature11316
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11316
This article is cited by
-
Neurobehavioral and developmental profiles: genotype–phenotype correlations in individuals with Cornelia de Lange syndrome
Orphanet Journal of Rare Diseases (2024)
-
Genetic determinants of micronucleus formation in vivo
Nature (2024)
-
Context-dependent perturbations in chromatin folding and the transcriptome by cohesin and related factors
Nature Communications (2023)
-
HDAC8-mediated inhibition of EP300 drives a transcriptional state that increases melanoma brain metastasis
Nature Communications (2023)
-
Enhanced performance of gene expression predictive models with protein-mediated spatial chromatin interactions
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.