Abstract
Identifying and understanding changes in cancer genomes is essential for the development of targeted therapeutics1. Here we analyse systematically more than 70 pairs of primary human colon tumours by applying next-generation sequencing to characterize their exomes, transcriptomes and copy-number alterations. We have identified 36,303 protein-altering somatic changes that include several new recurrent mutations in the Wnt pathway gene TCF7L2, chromatin-remodelling genes such as TET2 and TET3 and receptor tyrosine kinases including ERBB3. Our analysis for significantly mutated cancer genes identified 23 candidates, including the cell cycle checkpoint kinase ATM. Copy-number and RNA-seq data analysis identified amplifications and corresponding overexpression of IGF2 in a subset of colon tumours. Furthermore, using RNA-seq data we identified multiple fusion transcripts including recurrent gene fusions involving R-spondin family members RSPO2 and RSPO3 that together occur in 10% of colon tumours. The RSPO fusions were mutually exclusive with APC mutations, indicating that they probably have a role in the activation of Wnt signalling and tumorigenesis. Consistent with this we show that the RSPO fusion proteins were capable of potentiating Wnt signalling. The R-spondin gene fusions and several other gene mutations identified in this study provide new potential opportunities for therapeutic intervention in colon cancer.
Similar content being viewed by others
Main
Colorectal cancer (CRC) is the fourth most prevalent cancer, accounting for over 50,000 deaths per year in the United States2. Approximately 15% of CRCs have microsatellite instability (MSI) arising from defects in the DNA mismatch-repair (MMR) system3, whereas the other ∼85% of microsatellite-stable (MSS) CRCs are the result of chromosomal instability3. Genomic studies have identified mutations in genes, chromosomal structural variants and pathway alterations that probably contribute to CRC development3,4,5,6.
In an effort to understand better the pathogenesis of human CRCs we generated exome sequences (72 tumour–normal pairs, 15 MSI and 57 MSS; Supplementary Fig. 1 and Supplementary Table 2 ), RNA-seq (68 tumour–normal pairs), Illumina 2.5 M single-nucleotide-polymorphism (SNP) array data (74 tumour–normal pairs) and whole-genome sequences (1 MSI and 1 MSS tumour–normal pair) for a set of 74 colon tumour–normal pairs (Supplementary Table 1).
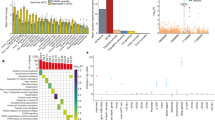
Exome sequencing identified 95,075 somatic mutations, of which 36,303 were protein altering (Fig. 1a). Two MSS samples were hypermutated (24,830 and 5,780 mutations, of which 9,479 and 2,332 were protein-altering mutations, respectively; Fig. 1a) and were not included in background mutation-rate calculations. We found 12,153 somatic mutations in 55 MSS samples studied (3,922 missense, 289 nonsense, 6 stop lost, 69 essential splice site, 20 protein-altering insertions/deletions (indels), 1,584 synonymous, 4,375 intronic and 1,888 others) and 52,312 somatic mutations in the 15 MSI samples (18,436 missense, 929 nonsense, 22 stop lost, 436 essential splice site, 363 protein-altering indels, 8,065 synonymous, 16,675 intronic and 7,386 others) (Fig. 1a and Supplementary Tables 3, 4 and 5). About 98% (35,524 out of 36,303) of the protein-altering single-nucleotide variants identified are new and are not reported in COSMIC v58 (ref. 7). We further confirmed a subset of the mutations reported to be somatic using RNA-seq data and/or mass spectrometry genotyping (Supplementary Methods, Supplementary Fig. 2 and Supplementary Table 3). In addition, we confirmed the presence of aberrant/alternate transcripts for 112 genes with canonical splice-site mutations using RNA-seq data (Supplementary Table 4). We observed a mean non-synonymous mutation frequency of 2.8 mutations per 106bases (31–149 coding-region mutations in 55 samples) for the MSS samples and 47 mutations per 106bases (764–3,113 coding-region mutations in 15 samples) for the MSI samples, consistent with the MMR defect in the latter (Fig. 1a) and the presence of alterations in one or more of the MMR genes in ∼80% of the samples (Supplementary Fig. 3 and Supplementary Table 6). Consistent with previous mutation reports4,6,8, analysis of base-level transitions and transversions at mutated sites using both exome and whole-genome sequencing (WGS) showed that C to T transitions were predominant in CRCs, regardless of MMR status (Fig. 1b, c), except for the two hypermutated tumour samples, which also showed a higher proportion of C to A and T to G transversions (Fig. 1b, c).
a, Number of mutations in each of the MSS, MSI and hypermutated subtypes. b, Transitions and transversions at the individual sample level for the three groups in the same order as shown in a. c, Transitions and transversions averaged across each of the three sample groups. d, Circos plot representation of the whole genome of an MSS and an MSI colon tumour. The outer blue band shows copy number as measured using WGS reads. Red dots indicate somatic non-synonymous, splice-site and stop-gained mutations (MSS genome, 17,651 mutations, average of 6.2 mutations per 106bases, 3,257 mutated genes; MSI genome, 97,968 mutations, average of 34.5 mutations per 106 bases, 9,851 mutated genes). Grey dots indicate all other somatic mutation types. Orange and grey lines show predicted intra- and interchromosomal rearrangements, respectively.
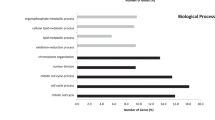
In an effort to understand the impact of the mutations on gene function we applied SIFT9, PolyPhen10 and mCluster11 and found, based on at least two of the three methods, that 34% of the somatic mutations probably have a functional consequence (Supplementary Table 3). We applied a previously described Q score metric to rank significantly mutated cancer genes12. In MSS samples, we identified 23 significantly mutated cancer genes (Q score, ≥ 1; false discovery rate, ≤ 5%; Supplementary Table 7). The MSS CRC genes identified from this analysis included previously reported genes such as KRAS, APC, TP53, SMAD4, FBXW7 and PIK3CA and several new genes including the cell cycle checkpoint gene ATM (Fig. 2a and Supplementary Table 7). KRAS and TP53 were among the top mutated MSI CRC genes; however, owing to the limited number of samples analysed, none of the genes achieved statistical significance.
a, Genes evaluated for significance using Q score criteria for MSS samples are shown. Each circle represents a gene and the size of the circle is proportional to the mutation count for that gene. The genes are represented in order of increasing number of expected mutations from left to right on the x axis. Genes with a statistically significant Q score are labelled. b, TCF12 helix–loop–helix (HLH) domain structure bound to DNA showing the Arg 603 hotspot and a schematic diagram depicting the various TCF12 mutations identified in this study. c, Somatic mutations in the TET gene family including TET1, TET2 and TET3 shown on the domain architecture of the TET genes. Recurrent mutations found in this study for TET3 and for TET2 by comparison to COSMIC data are shown in red. CD, Cysteine-rich domain; CXXC, Cys-X-X-Cys-type domain; DSBH, double-stranded β-helix 2OG-Fe(II)-dependent dioxygenase domain. Asterisks in b and c denote nonsense mutations that result in premature stop codons.
Of the 432 candidate CRC genes identified in colon cancer mouse model screens13,14, 356 were mutated in this study (Supplementary Table 8). Frequently mutated genes that overlapped included KRAS, APC, SMAD4, FBXW7 and EP400, as well as genes involved in chromatin remodelling such as SIN3A, SMARCA5, NCOR1 and histone-modifying enzyme JARID2 (Supplementary Table 8). TCF12, a CRC gene identified in a mouse model screen, was mutated in five of the MSI samples. A hotspot mutation in TCF12 at Arg 603 (three out of five mutations) was found within the TCF12 helix–loop–helix domain and probably abolishes its DNA-binding ability (Fig. 2b).
Mutational hotspots are indicative of functionally relevant driver genes. In this study we have identified 274 genes with hotspot mutations (Supplementary Table 9 and Supplementary Figs 4 and 5), 40 of which were not previously reported in COSMIC7. Comparison with mutations reported in COSMIC7 identified an additional 435 hotspot mutations in 361 genes (Supplementary Table 10), 18 of which were also identified in mouse CRC screens13,14 (Supplementary Table 8). Mutations in the TET family of genes that encode methylcytosine dioxygenases have not been reported in CRC4,15. We found that all three family members—TET1, TET2 and TET3—were mutated in our study (Fig. 2c). In addition, ERBB3, which was mutated in 8% (6 out of 72) of the samples, contained multiple hotspot mutations that were oncogenic, as reported in a separate study (B.S.J. et al., submitted).
The top differentially expressed genes identified using RNA-seq (Supporting Methods, Supplementary Figs 6 and 7 and Supplementary Table 11) included FOXQ1 and CLDN1, both of which have been implicated in colon tumorigenesis16. Notably, we also identified IGF2 upregulation in 12% (8 out of 68) of the samples (Supplementary Fig. 7). Most (7 out of 8) of the tumours with IGF2 overexpression also showed focal amplification of the IGF2 locus as measured by Illumina 2.5 M arrays (Supplementary Fig. 8). Overall, the differentially expressed genes affect multiple signalling pathways, including Wnt and glutamate receptor signalling (Supplementary Fig. 26 and Supplementary Table 12).
We assessed copy-number alterations in 74 tumour–normal pairs by applying GISTIC17 to the circular binary segmented (CBS)18 copy-number data. In addition to the IGF2 amplifications, we found known amplifications involving KRAS (13%; 10 out of 74) and MYC (23%; 17 out of 74) located in a broad amplicon on chromosome 8q (Supplementary Fig. 9 and Supplementary Table 13). Deletion involving FHIT, a tumour-suppressor gene19, was observed in 30% (22 out of 74) of the samples (Supplementary Table 14 and Supplementary Fig. 9). We also detected deletion of APC (8%; 6 out of 74), PTEN (4%; 3 out of 74) and SMAD3 (9%; 10 out of 74). SMAD4 and SMAD2 are both altered in 27% (20 out of 74) of the samples and are located within 3 megabases (Mb) of each other on chromosome 18q, which is frequently lost (Supplementary Fig. 9). Finally, we found chromosome 20q to be frequently gained (Supplementary Fig. 9).
Using RNA-seq data we identified 36 rearrangements that result in gene fusions20, including two recurrent ones (Supplementary Fig. 10 and Supplementary Table 15). The recurrent fusions found in MSS samples involve the R-spondin family members, RSPO2 (3%; 2 out of 68) and RSPO3 (8%; 5 out of 68). R-spondins are secreted proteins that can potentiate canonical Wnt signalling21. The recurrent RSPO2 fusion identified in two tumour samples involves EIF3E exon 1 and RSPO2 exon 2 (Fig. 3b). This fusion transcript is expected to produce a functional RSPO2 protein driven by the EIF3E promoter (Fig. 3d). A second RSPO2 fusion that was not expected to produce a functional protein, EIF3E(e1)–RSPO2(e3), was also detected in the same samples (Supplementary Table 15). To confirm the nature of the alteration at the genome level, we performed WGS of the tumours containing RSPO2 fusions. Analysis of junction-spanning reads, mate-pair reads and copy-number data derived from the WGS data identified a 158 kilobases (kb) deletion in one sample and a 113 kb deletion in the second sample, both of which places exon 1 of EIF3E in close proximity to the 5′ end of RSPO2 (Supplementary Figs 11 and 12), further supporting the fusions observed.
a, List of type and frequency of R-spondin gene fusions in colon cancer. b, Cartoon depicting the location, orientation and exon–intron architecture of EIF3E–RSPO2 fusion on the genome. The read evidence for the EIF3E(e1)–RSPO2(e2) fusion identified using RNA-seq data is shown. c, Independent reverse transcriptase (RT)–PCR-derived products confirming the EIF3E(e1)–RSPO2(e2) somatic fusion resolved on an agarose gel. The RT–PCR product was Sanger-sequenced confirming the fusion junction and a representative chromatogram is shown in b. d, Schematic of the resulting EIF3E(e1)–RSPO2(e2) fusion protein. e, Tumours harbouring R-spondin fusions show elevated expression of the corresponding RSPO gene as depicted by the heat-map. f, Cartoon depicting the location, orientation and exon–intron architecture of the PTPRK–RSPO3 gene fusion on the genome. The read evidence for PTPRK(e1)–RSPO3(e2) fusion identified using RNA-seq data is shown. g, Independent RT–PCR-derived products confirming the PTPRK(e1)–RSPO3(e2) somatic fusion resolved on an agarose gel. The RT–PCR product was Sanger-sequenced confirming the fusion junction, and a representative chromatogram is show in panel f. h, Schematic of PTPRK, RSPO3 and the resulting PTPRK(e1)–RSPO3(e2) fusion proteins. FN3, fibronectin type 3 domain; FU, furin-like repeats; IG, immunoglobulin; MAM, meprin A5 receptor protein tyrosine phosphatase mu domain; N, normal; PTPc, protein tyrosine phosphatase catalytic domain; T, tumour; TM, transmembrane domain; TSP1, thrombospondin type 1 repeats.
The RSPO3 gene fusions were observed in 5 out of 68 tumours and involve PTPRK as the 5′ partner. We identified two different RSPO3 fusion variants consisting of either exon 1 or exon 7 of PTPRK and exon 2 of RSPO3 (Fig. 3f–h and Supplementary Fig. 13). WGS reads from the five tumours expressing the RSPO3 fusions showed rearrangements involving a simple (three samples) or a complex (two samples) inversion that places RSPO3 in proximity to PTPRK on the same strand as PTPRK on chromosome 6q (Supplementary Figs 14–18). The PTPRK(e1)–RSPO3(e2) transcript found in four samples is an in-frame fusion that preserves the entire coding sequence of RSPO3 and replaces its secretion signal sequence with that of PTPRK (Fig. 3h). The PTPRK(e7)–RSPO3(e2) fusion, detected in one sample, is also an in-frame fusion that encodes a ∼70 kDa protein consisting of the first 387 amino acids of PTPRK, including its secretion signal sequence, and the RSPO3 amino acids 34–272 lacking its native signal peptide (Supplementary Fig. 13c). Interestingly, RNA-seq data showed that the expression of RSPO2 and RSPO3 in colon tumour samples containing the fusions was elevated compared with the tumour samples lacking R-spondin fusions (Fig. 3e). Furthermore, all of the RSPO-positive fusion tumours expressed the potential R-spondin receptors LGR4, LGR5 and LGR6 (refs 22, 23, 24), although LGR6 expression was lower compared with LGR4 and LGR5 (Supplementary Fig. 19).
To confirm the functional relevance of the R-spondin fusion proteins, we first expressed R-spondin fusion constructs in 293T cells and confirmed that the fusion proteins were expressed and secreted as predicted (Fig. 4a). We then examined the biological activity of the R-spondin fusions using a Wnt-responsive luciferase reporter, TOPbrite22,23,24,25. As observed with wild-type RSPO2 and RSPO3, conditioned media from cells transfected with RSPO fusion expression constructs led to activation of the Wnt pathway (Fig. 4b). The observed activation, although apparent in the absence of exogenous Wnt, was further potentiated in the presence of recombinant Wnt, consistent with the known role of R-spondins in Wnt signalling21,22,23,24. In addition to 293T cells, we confirmed the ability of RSPO fusion proteins to activate Wnt signalling in HT-29, a human colon cancer cell line (Supplementary Fig. 20).
a, Secreted RSPO fusion proteins detected by western blot in media from 293T cells transfected with expression constructs encoding the fusion proteins. The arrowhead indicates the expected product for RSPO 1–387. b, RSPO fusion proteins activate and potentiate Wnt signalling as measured using a luciferase reporter assay (see Supplementary Methods). Data shown are from conditioned media derived from cells transfected with the fusion constructs. Representative data from at least three experiments are shown. Error bars represent mean ± s.d. from three replicate experiments. EGFP, enhanced green fluorescent protein. c, Cartoon representing R-spondin-mediated Wnt signalling pathway activation. d, Quilt plot depicting RSPO fusions, somatic mutations, copy-number alteration and expression changes across a select set of Wnt-signalling pathway genes. Each column represents a tumour sample. Het, heterozygous; Hom, homozygous; WT, wild type.
To characterize further the RSPO gene fusions we analysed them in the context of other mutations (Fig. 4c, d, Supplementary Figs 21 and 25 and Supplementary Table 16). We found that most of the colon tumours carried alterations in Wnt, TGF-β, Rtk/Ras and TP53 pathways (Supplementary Figs 22–26). Interestingly, the RSPO2 and RSPO3 fusions were mutually exclusive and occurred in tumours that did not contain APC mutations or copy loss (Fig. 4d; P = 0.038, Fisher’s exact test), with the exception of one tumour that contained an RSPO2 fusion and a single APC allele (Fig. 4d). In addition, the RSPO gene fusions were mutually exclusive with mutations in CTNNB1, another Wnt pathway gene that is mutated in CRC. However, all the samples with RSPO gene fusions also carried a mutation in either KRAS or BRAF (Supplementary Fig. 21). Most APC mutant samples also have Ras pathway gene mutations, indicating that the RSPO gene fusions may function analogously to APC mutations in promoting Wnt signalling during colon tumour development. This is further supported by the fact that the RSPO fusion-positive tumours showed upregulation of Wnt pathway target genes similar to tumours carrying APC mutations (Supplementary Fig. 27). Interestingly, transposon-based insertional mutagenesis screens in mouse models of CRC identified Rspo2 as a top-candidate cancer gene14 (Supplementary Table 8). Furthermore, analysis of these mouse colon tumours showed that the transposon insertions were in most cases mutually exclusive between Apc and Rspo2 (Supplementary Fig. 28a), confirming the mutual exclusivity observed in human samples. Most of the transposon insertions in the mouse Rspo2 gene occur near the transcriptional start site in the same orientation as the Rspo2 promoter, and probably lead to elevated Rspo2 expression (Supplementary Fig. 28b). Consistent with this and with the elevated expression of RSPO genes observed in human tumours, we found a ∼20-fold increase in Rspo2 messenger RNA expression in a mouse tumour carrying a transposon insertion near the Rspo2 transcription start site, compared with adjacent normal tissue (Supplementary Fig. 28c). Taken together, our findings indicate that the R-spondins probably function as drivers in human CRCs. Although further studies will be required to fully understand the role of R-spondin fusions in CRC development, they represent attractive targets for antibody-based therapy in CRC patients positive for R-spondin fusions. In addition, other therapeutic strategies that target downstream components of the Wnt signalling cascade will probably be effective against tumours positive for R-spondin fusions.
Methods Summary
Fresh–frozen primary human colon tumours and their matched normal tissues were obtained from commercial sources with appropriate institutional approval. DNA and RNA isolated from tumours with >70% tumour content and their corresponding matched normal tissue were subject to exome sequencing, RNA-seq and SNP array analysis. A complete description of the materials and methods is provided in the Supplementary Information.
Online Methods
Samples, DNA and RNA preparation and MSI testing
Patient-matched fresh–frozen primary colon tumours and normal tissue samples with appropriate Institutional Review Board approval and patient-informed consent were obtained from commercial sources (Supplementary Table 1) and subjected to genomic analysis described below. The human tissue samples used in the study were de-identified (double-coded) before their use and hence the study using these samples is not considered human subject research under the US Department of Human and Health Services regulations and related guidance (45 CFR Part 46). All tumour and normal tissues were subject to pathology review. From a set of 90 paired tumour–normal samples we identified 74 sample pairs for further analysis. Tumour DNA and RNA were extracted using the Qiagen AllPrep DNA/RNA kit (Qiagen). Tumour samples were assessed for microsatellite instability using an MSI detection kit (Promega).
Exome capture and sequencing
In total, 72 tumour samples and matched normal tissues were analysed by exome sequencing (Supplementary Table 1). Exome capture was performed using SeqCap EZ human exome library v2.0 (NimbleGen) consisting of 2.1 million empirically optimized long oligonucleotides that target ∼30,000 coding genes (∼300,000 exons, total size 36.5 Mb). The library is capable of capturing a total of 44.1 Mb of the genome, including genes and exons represented in RefSeq (Jan 2010), Consensus CDS Project (CCDS; Sept 2009) and miRBase (v.14, Sept 2009). Exome-capture libraries generated were sequenced on HiSeq 2000 (Illumina). One lane of 2 × 75 base pair (bp) paired-end data were collected for each sample. The targeted regions had a mean coverage of 179× with 97.4% bases covered at ≥ 10 times (Supplementary Fig. 1 and Supplementary Table 2).
RNA-seq
RNA from 68 colon tumour and matched normal sample pairs was used to generate RNA-seq libraries using TruSeq RNA Sample Preparation kit (Illumina). RNA-seq libraries were multiplexed (two per lane) and sequenced on HiSeq 2000 as per manufacturer’s recommendation. We generated ∼30 million paired-end (2 × 75 bp) sequencing reads per sample (Supplementary Fig. 5).
Sequence data processing
All short-read data were evaluated for quality control using the Bioconductor ShortRead package26. To confirm that all samples were identified correctly, all exome and RNA-seq data variants that overlapped with the Illumina 2.5 M array data were compared and checked for consistency. An all-by-all sample comparison was done on germline variants to check that all pairs were matched correctly between the respective patient tumour and matched normal sample and correspondingly did not match with any other patient pair above a cutoff of 90%.
Variant calling
Sequencing reads were mapped to the University of California, Santa Cruz (UCSC) human genome (GRCh37/hg19) using Burrows–Wheeler alignment (BWA) software27 set to default parameters. Local realignment, duplicate marking and raw variant calling were performed as described previously28. Known germline variations represented in dbSNP Build 13129, but not represented in COSMIC v548, were filtered out. In addition, variants that were present in both the tumour and normal samples were removed as germline variations. Remaining variations present in the tumour sample but absent in the matched normal were predicted to be somatic. Predicted somatic variations were additionally filtered to include only positions with a minimum of 10× coverage in both the tumour and matched normal as well as an observed variant allele frequency of <3% in the matched normal and a significant difference in variant allele counts using Fisher’s exact test. To evaluate the performance of this algorithm we randomly selected 807 protein-altering variants and validated them using Sequenom nucleic acid technology as described previously13. Of these, 93% (753) validated as cancer specific, with the invalidated variants being equally split between not being seen in the tumour and also being seen in the adjacent normal (germ line). In addition, we also targeted and validated 164 mutations belonging to significant Q score genes and 212 hotspot mutations using Sequenom mass spectrometry technology. All variants that invalidated were removed from the final set. Indels were called using the GATK Indel Genotyper Version 2, which reads both the tumour and normal BAM file for a given pair28.
To identify additional variants that had a lower mean allele frequency or were affected by a specific mapper, we included Sequenom validated variants using the following algorithm. Reads were mapped to the UCSC human genome (GRCh37/hg19) using GSNAP30. Variants seen at least twice at a given position and greater than 10% allele frequency were selected. These variants were additionally filtered for significant biases in strand and position using Fisher’s exact test. In addition, variants that did not have adequate coverage in the adjacent normal as determined as at least a 1% chance of being missed using a binomial distribution at a normal allele frequency of 12.5% were excluded. All new protein-altering variants identified by the second algorithm were validated by Sequenom, which resulted in a total of 515 additional variants. The effect of all non-synonymous somatic mutations on gene function was predicted using SIFT10 and PolyPhen11. All variants were annotated using Ensembl (release 59, http://www.ensembl.org).
Validation of somatic mutations and indels
We used single base pair extension followed by nucleic acid mass spectrometry (Sequenom) to validate the predicted somatic mutations as described previously13. Tumour and matched normal DNA were whole-genome amplified using the REPLI-g Whole Genome Amplification Midi kit (Qiagen) and cleaned up as per manufacturer’s recommendations. Variants found as expected in the tumour but absent in the normal DNA were designated somatic. Those that were present in both tumour and normal samples were classified as germline. Variants that could not be validated in tumour or normal samples were designated as failed. For indel validation we designed primers for PCR that generated an amplicon of ∼300 bp that contained the indel region. We PCR-amplified the region in both tumour and matched normal samples using Phusion (New England Biolabs) as per manufacturer’s instructions. We then purified the PCR products on a gel, isolated the relevant fragments and Sanger-sequenced them. The sequencing trace files were analysed using Mutation Surveyor (SoftGenetics). Indels that were present in the tumour and absent in the normal sample were designated somatic and are reported in Supplementary Table 4.
Mutational significance and pathway analysis
We evaluated the mutational significance of genes using a previously described method13. In brief, this method can identify genes that have more statistically significant protein-altering mutations than what would be expected based on a calculated background-mutation rate. The background-mutation rate is calculated for six different nucleotide mutation categories (A, C, G, T, CG1, CG2), in which there was sufficient coverage (≥10×) in both the tumour and matched normal sample. A non-synonymous to synonymous ratio, ri, was calculated using a simulation of mutating all protein-coding nucleotides and seeing if the resulting change would result in a synonymous or non-synonymous change. The background mutation rate, fi, was determined by multiplying the number of synonymous somatic variants by ri and normalizing by the total number of protein-coding nucleotides. The number of expected mutations for a given gene was determined as the number of protein-coding bases multiplied by fi and integrated across all mutation categories. A P value was calculated using a Poisson probability function given the expected and observed number of mutations for each gene. P values were corrected for multiple testing using the Benjamini–Hochberg method and the resulting Q values were converted to Q scores by taking the negative log10 of the Q values with a maximum possible value of 16. Given that different mutation rates existed for the MSI and MSS samples, Q scores were calculated separately for each, with the two hypermutated samples being removed completely. To avoid underestimating the background mutation rates, samples with less than 50% tumour content were excluded from the analysis. Pathway mutational significance was also calculated as previously described13, with the exception that the BioCarta Pathway database was used, which was downloaded as part of the MSigDB31. For quilt plots a fourfold increase or decrease compared with the matched normal was used as the cutoff (Fig. 4d and Supplementary Figs 20–24). A log2 tumour to normal ratio of 0.585 and −0.7 was used for copy-number gain and loss, respectively, for these plots. Samples where exome data were not collected were excluded from the plot (Fig. 4d and Supplementary Figs 20–24), Only samples with either a copy number change and/or a mutation are depicted. To help differentiate single-copy from double-copy loss in MMR genes a cutoff of −1 was used for copy-loss events (Supplementary Fig. 3)
WGS and analysis
Paired-end sequence reads were aligned to GRCh37 using BWA. Further processing of the alignments to obtain mutation calls was done in a similar manner to the exome-sequencing analysis using the GATK pipeline. Copy number was calculated by computing the number of reads in 10-kb non-overlapping bins and taking the tumour–normal ratio of these counts. Chromosomal breakpoints were predicted using BreakDancer32. Genome plots were created using Circos33. For detection of genomic alteration in tumours with RSPO fusion, reads aligning to the genomic region containing the RSPO, EIF3E and PTPRK genes were analysed further to reveal the deletions and insertions.
RNA-seq data analysis
RNA-seq reads were aligned to the human genome version GRCh37 using GSNAP30. Expression counts per gene were obtained by counting the number of reads aligning concordantly within a pair and uniquely to each gene locus as defined by CCDS. The gene counts were then normalized for library size and subsequently variance stabilized using the DESeq Bioconductor software package34. Differential gene expression was computed by pairwise t-tests on the variance-stabilized counts followed by correction for multiple testing using the Benjamini–Hochberg method.
SNP array data generation and analysis
Illumina HumanOmni2.5_4v1 arrays were used to assay 74 colon tumours and matched normal for genotype, DNA copy and loss of heterozygosity at ∼2.5 million SNP positions. These samples all passed our quality-control metrics for sample identity and data quality (see below). A subset of 2,295,239 high-quality SNPs was selected for all analyses.
After making modifications to permit use with Illumina array data, we applied the PICNIC35 algorithm to estimate total copy-number and allele-specific copy number/loss of heterozygosity. Modifications included replacement of the segment-initialization component with the CBS algorithm36, and adjustment of the prior distribution for background raw copy-number signal (adjusted mean of 0.7393 and s.d. of 0.05). For the pre-processing required by PICNIC’s hidden Markov model, we used a Bayesian model to estimate cluster centroids for each SNP. For SNP k and genotype g, observed data in normal samples were modelled as following a bivariate Gaussian distribution. Cluster centres for the three diploid genotypes were modelled jointly by a six-dimensional Gaussian distribution, with mean treated as a hyperparameter and set empirically based on a training set of 156 normal samples. Cluster centre and within-genotype covariance matrices were modelled as inverse Wishart, with scale matrix hyperparameters also set empirically, and with degrees of freedom manually tuned to provide satisfactory results for a wide range of probe behaviour and minor allele frequencies. Finally, signal for SNP k (for the A and B alleles separately) was transformed with a nonlinear function  , with parameters selected on the basis of the posterior distributions computed above.
, with parameters selected on the basis of the posterior distributions computed above.
Sample identity was verified using genotype concordance: pairs of tumours from the same patient were expected to have >90% concordance, and all other pairs <80% concordance. Samples failing these criteria were excluded. Quality of the overall HMM fit was assessed by root mean squared (r.m.s.) error with respect to fitted values; samples with a r.m.s. error of >1.5 were excluded. Finally, to account for two commonly observed artefacts, fitted copy number values were set to ‘NA’ for singletons with a fitted total copy number of 0, or when the observed and fitted means differed by more than 2 for regions of inferred copy gain.
Recurrent DNA copy-number gain and loss
Genomic regions with recurrent DNA copy gain and loss were identified using GISTIC, version 2.017. Probe integer total copy-number values obtained from PICNIC, c, were converted to log2 ratio values, y, as y = log2(ctumour + 0.1) – log2(ccontrol + 0.1). These ratio values were then segmented using CBS18. Cut-offs of +0.584 and −0.7 were used to categorize log2 ratio values as gain or loss, respectively. A minimum segment length of 20 SNPs and a log2 ratio ‘cap’ value of 3 were used. The same CBS segmented data and thresholds were used to compute a frequency of gain or loss for each gene found to be significant by GISTIC, which is reported in Supplementary Tables 13 and 14.
Fusion detection and validation
Putative fusions were identified using a computational pipeline we have developed called GSTRUCT-fusions. Our pipeline is based on a generate-and-test strategy that is fundamentally similar to our methodology reported previously for finding read-through fusions37. Paired-end reads were aligned using our alignment program GSNAP37. GSNAP has the ability to detect splices representing translocations, inversions and other distant fusions within a single read end.
These distant splices provided one set of candidate fusions for the subsequent testing stage. The other set of candidate fusions derived from unpaired unique alignments, in which each end of the paired-end read aligned uniquely to a different chromosome, and also from paired, but discordant unique alignments, in which each end aligned uniquely to the same chromosome, but with an apparent genomic distance that exceeded 200,000 bp or with genomic orientations that suggested an inversion or scrambling event.
Candidate fusions were then filtered against known transcripts from RefSeq and aligned to the genome using GMAP38. We required that both fragments flanking a distant splice, or both ends of an unpaired or discordant paired-end alignment, map to known exon regions. This filtering step eliminated approximately 90% of the candidates. We further eliminated candidate inversions and deletions that suggested rearrangements of the same gene, as well as apparent read-through fusion events involving adjacent genes in the genome, which our previous research indicated were likely to have a transcriptional rather than a genomic origin.
For the remaining candidate fusion events, we constructed artificial exon–exon junctions consisting of the exons distal to the supported donor exon and the exons proximal to the supported acceptor exon. The exons included in the proximal and distal computations were limited so that the cumulative length along each gene was within an estimated maximum insert length of 200 bp. As a control, we also constructed all exon–exon junctions consisting of combinations of exons within the same gene, for all genes contributing to a candidate fusion event.
In the testing stage of our pipeline, we constructed a genomic index from the artificial exon–exon junctions and controls using the GMAP_BUILD program included as part of the GMAP and GSNAP package. We used this genomic index and the GSNAP program with splice detection turned off to re-align the original read ends that were not concordant to the genome. We extracted reads that aligned to an intergenic junction corresponding to a candidate fusion, but not to a control intragenic junction.
We filtered the results of the re-alignment to require that each candidate fusion have at least one read with an overhang of 20 bp. We also required that each candidate fusion have at least ten supporting reads. For each remaining candidate fusion, we aligned the two component genes against each other using GMAP and eliminated the fusion if the alignment had any region containing 60 matches in a window of 75 bp. We also aligned the exon–exon junction against each of the component genes using GMAP and eliminated the fusion if the alignment had coverage greater than 90% of the junction and identity greater than 95%.
Validation of gene fusions was done using an RT–PCR approach using both colon tumour and matched normal samples. 500 ng of total RNA was reverse transcribed to complementary DNA with a High Capacity cDNA Reverse Transcription kit (Life Technologies) following manufacturer’s instructions. 50 ng of cDNA was amplified in a 25 μl reaction containing 400 pM of each primer, 300 μM of each deoxynucleoside triphosphate and 2.5 U of LongAmp Taq DNA polymerase (New England Biolabs). PCR was performed with an initial denaturation at 95 °C for 3 min followed by 35 cycles of 95 °C for 10 s, 56 °C for 1 min and 68 °C for 30 s, and a final extension step at 68 °C for 10 min. 3 μl of PCR product was run on 1.2% agarose gel to identify samples containing the gene fusion. Specific PCR products were purified with either a QIAquick PCR Purification kit or Gel Extraction kit (Qiagen). The purified DNA was either sequenced directly with PCR primers specific to each fusion (Supplementary Table 14) or cloned into TOPO cloning vector pCR2.1 (Life Technologies) before Sanger sequencing. The clones were sequenced using Sanger sequencing on an ABI 3730 XL (Life Technologies) as per manufacturer’s instructions. The Sanger sequencing trace files were analysed using Sequencher (Gene Codes Corp.).
RSPO fusion activity testing
A eukaryotic expression plasmid pRK5E driving the expression of carboxy-terminal Flag-tagged EIF3E, PTPKR (amino acids 1–387), RSPO2, RSPO3, EIF3E(e1)–RSPO2(e2), PTPRK(e1)–RSPO3(e2) and PTPRK(e7)–RSPO3(e2) was generated using standard PCR and cloning strategies.
Cells, conditioned media, immunoprecipitation and western blotting
HEK293T and HT-29 cells were obtained from Genentech’s cell banking facility. HEK 293T cells were maintained in DMEM buffer supplemented with 10% FBS. HT-29 cells were maintained in RPMI 1640 supplemented with 10% FBS. For expression analysis and condition media generation 3 × 105 HEK29T3 cells were plated in six-well plates in 1.5 ml DMEM containing 10% FBS. Cells were transfected with 1 μg DNA using Fugene 6 (Roche) according to manufacturer’s instructions. Media was conditioned for 48 h, collected, centrifuged and used to stimulate the luciferase reporter assay (final concentration, ×0.1–0.4). For expression analysis, media was collected, centrifuged to remove debris and used for immunoprecipitation.
Luciferase reporter assays
HEK293T (50,000 cells per ml) or HT-29 cells (110,000 cells per ml) were plated in 90 μl of media containing 2.5% FBS per well of a 96-well plate. After 24 h, cells were transfected using Fugene 6 according to manufacturer’s instructions (Roche) with the following DNA per well: 0.04 μg (HEK293T) or 0.08 μg (HT-29) TOPbrite Firefly reporter25 and 0.02 μg (HEK293T) and 0.04 μg (HT-29) pRL-SV40 Renilla (Promega) constructs. Cells were stimulated with 25 μl of either fresh or conditioned media (described above) containing 10% FBS with or without recombinant murine Wnt3a (20–100 ng per ml (final), R&D Systems). After 24 h stimulation, 50 μl of media was removed and replaced with Dual-Glo luciferase detection reagents (Promega) according to manufacturer’s instructions. An Envision Luminometer (Perkin–Elmer) was used to detect luminescence. To control for transfection efficiency, firefly luciferase levels were normalized to Renilla luciferase levels to generate the measure of relative luciferase units. Experimental data are presented as mean ± s.d. from three independent wells.
Immunoprecipitation and western blotting
To confirm that the RSPO wild-type and RSPO fusion proteins were secreted, Flag-tagged proteins were immunoprecipitated from the media using anti-Flag–M2 antibody-coupled beads (Sigma), boiled in SDS–PAGE loading buffer, resolved on 4–20% SDS–PAGE (Invitrogen) and transferred onto a nitrocellulose membrane. RSPO and other Flag-tagged proteins expressed in cells were detected from cell lysates using western blot as described previously. In brief, immunoprecipitated proteins and proteins from cell lysates were detected by western blot using Flag–horseradish peroxidase-conjugated antibody and Super signal West Dura chemiluminescence detection substrate (Thermo Fisher Scientific).
Transposon-based genetic screen
In brief, a cohort of triple-transgenic mice (Rosa26–LsL–Sb11, T2/Onc, Villin–Cre) were generated to allow tissue-restricted mobilization of a mutagenic Sleeping Beauty transposon in the gastrointestinal epithelium6. Triple-transgenic mice developed intestinal intraepithelial neoplasias, adenomas and adenocarcinomas. In total, 135 tumours from 47 mice were analysed for transposon-insertion sites by DNA extraction, linker-mediated PCR to amplify genomic DNA-transposon junctions, and 454 pyrosequencing. 16,000 non-redundant transposon insertion sites were mapped, and a Monte Carlo method was used to define common insertion sites (CISs) that were recurrently targeted by transposon insertions at a higher frequency than predicted by chance.
Quantitative RT–PCR
Quantitative RT–PCR reactions were performed twice in duplicate using Fast Start Universal SYBR Master (Roche) and a Mastercycler ep realplex thermal cycler (Eppendorf). Primers sequences are listed below (5′ to 3′):
Mouse Actb_F1: TCCAGCCTTCCTTCTTGGGTATGGA
Mouse Actb_R1: CGCAGCTCAGTAACAGTCCGCC
Human RSPO2_F1: TTATTTGCATAGAGGCCGTTGCTTT
Human RSPO2_R1: GCTCCAATGACCAACTTCACATCCTT
Accession codes
Data deposits
Sequencing and genotype data has been deposited at the European Genome-Phenome Archive (http://www.ebi.ac.uk/ega/), which is hosted by the EBI, under accession number EGAS00001000288.
References
Chin, L., Andersen, J. N. & Futreal, P. A. Cancer genomics: from discovery science to personalized medicine. Nature Med. 17, 297–303 (2011)
Siegel, R., Ward, E., Brawley, O. & Jemal, A. Cancer statistics, 2011: the impact of eliminating socioeconomic and racial disparities on premature cancer deaths. CA Cancer J. Clin. 61, 212–236 (2011)
Fearon, E. R. Molecular genetics of colorectal cancer. Annu. Rev. Pathol. 6, 479–507 (2011)
Wood, L. D. et al. The genomic landscapes of human breast and colorectal cancers. Science 318, 1108–1113 (2007)
Timmermann, B. et al. Somatic mutation profiles of MSI and MSS colorectal cancer identified by whole exome next generation sequencing and bioinformatics analysis. PLoS ONE 5, e15661 (2010)
Bass, A. J. et al. Genomic sequencing of colorectal adenocarcinomas identifies a recurrent VTI1A–TCF7L2 fusion. Nature Genet. 43, 964–968 (2011)
Forbes, S. A. et al. COSMIC (the Catalogue of Somatic Mutations in Cancer): a resource to investigate acquired mutations in human cancer. Nucleic Acids Res. 38, D652–D657 (2010)
Sjoblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006)
Ng, P. C. & Henikoff, S. Accounting for human polymorphisms predicted to affect protein function. Genome Res. 12, 436–446 (2002)
Ramensky, V., Bork, P. & Sunyaev, S. Human non-synonymous SNPs: server and survey. Nucleic Acids Res. 30, 3894–3900 (2002)
Yue, P. et al. Inferring the functional effects of mutation through clusters of mutations in homologous proteins. Hum. Mutat. 31, 264–271 (2010)
Kan, Z. et al. Diverse somatic mutation patterns and pathway alterations in human cancers. Nature 466, 869–873 (2010)
March, H. N. et al. Insertional mutagenesis identifies multiple networks of cooperating genes driving intestinal tumorigenesis. Nature Genet. 43, 1202–1209 (2011)
Starr, T. K. et al. A transposon-based genetic screen in mice identifies genes altered in colorectal cancer. Science 323, 1747–1750 (2009)
Mohr, F., Dohner, K., Buske, C. & Rawat, V. P. TET genes: new players in DNA demethylation and important determinants for stemness. Exp. Hematol. 39, 272–281 (2011)
Kaneda, H. et al. FOXQ1 is overexpressed in colorectal cancer and enhances tumorigenicity and tumor growth. Cancer Res. 70, 2053–2063 (2010)
Mermel, C. H. et al. GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol. 12, R41 (2011)
Olshen, A. B., Venkatraman, E. S., Lucito, R. & Wigler, M. Circular binary segmentation for the analysis of array-based DNA copy number data. Biostatistics 5, 557–572 (2004)
Pichiorri, F. et al. Fhit tumor suppressor: guardian of the preneoplastic genome. Future Oncol. 4, 815–824 (2008)
Ozsolak, F. & Milos, P. M. RNA sequencing: advances, challenges and opportunities. Nature Rev. Genet. 12, 87–98 (2011)
Yoon, J. K. & Lee, J. S. Cellular signaling and biological functions of R-spondins. Cell. Signal. 24, 369–377 (2012)
Carmon, K. S., Gong, X., Lin, Q., Thomas, A. & Liu, Q. R-spondins function as ligands of the orphan receptors LGR4 and LGR5 to regulate Wnt/β-catenin signaling. Proc. Natl Acad. Sci. USA 108, 11452–11457 (2011)
de Lau, W. et al. Lgr5 homologues associate with Wnt receptors and mediate R-spondin signalling. Nature 476, 293–297 (2011)
Glinka, A. et al. LGR4 and LGR5 are R-spondin receptors mediating Wnt/β-catenin and Wnt/PCP signalling. EMBO Rep. 12, 1055–1061 (2011)
Zhang, Y. et al. Inhibition of Wnt signaling by Dishevelled PDZ peptides. Nature Chem. Biol. 5, 217–219 (2009)
Morgan, M. et al. ShortRead: a bioconductor package for input, quality assessment and exploration of high-throughput sequence data. Bioinformatics 25, 2607–2608 (2009)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nature Genet. 43, 491–498 (2011)
Sherry, S. T. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001)
Wu, T. D. & Nacu, S. Fast and SNP-tolerant detection of complex variants and splicing in short reads. Bioinformatics 26, 873–881 (2010)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Chen, K. et al. BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nature Methods 6 677–681 (2009) CrossRef
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009)
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010)
Greenman, C. D. et al. PICNIC: an algorithm to predict absolute allelic copy number variation with microarray cancer data. Biostatistics 11, 164–175 (2010)
Venkatraman, E. S. & Olshen, A. B. A faster circular binary segmentation algorithm for the analysis of array CGH data. Bioinformatics 23, 657–663 (2007)
Nacu, S. et al. Deep RNA sequencing analysis of readthrough gene fusions in human prostate adenocarcinoma and reference samples. BMC Med. Genomics 4, 11 (2011)
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005)
Acknowledgements
The authors would like to acknowledge Genentech DNA Sequencing, Oligo, Microarray and gCell laboratories for their help with the project. We thank the Genentech Bioinformatics group for informatics infrastructure support and the Pathology Core Labs for providing histology, immunohistochemistry and tissue-management support. We also thank M. Costa, M. Callow, P. Polakis and the de Sauvage and Seshagiri laboratories for comments and suggestions. Work in the Largaespada laboratory was supported by National Institutes of Health grant R01-CA134759 (to D.A.L.).
Author information
Authors and Affiliations
Contributions
S.S. and F.J.d.S. conceived the study and designed experiments. E.W.S. and S.D. performed the exome, WGS, RNA-seq and copy data analysis. Z.M., C.H. and Y.G. performed the validation of the fusions. Z.M. managed exome capture. S.C., V.J., B.S.J. and E.E.S. validated and tested the fusions for activity. C.B.C., T.K.S. and D.A.L. performed the mouse model analysis. G.J.P.D., J.S. and Z.M. performed tissue management and nucleic acid preparation. J.S. and J.G. collected and performed mutation validation. F.G. and Z.Z. predicted the functional effect of mutations. E.W.S., M.A.H., J.D.D., T.D.W. and R.G. provided bioinformatics support, including an algorithm for variant calling, fusion detection and copy number calling. R.B. and P.M.H. analysed SNP array data. W.W. performed structural biology support and H.K. performed pathology support. S.S., E.W.S., S.D., Z.M. and F.J.d.S. wrote the paper, which was reviewed and edited by the other co-authors.
Corresponding authors
Ethics declarations
Competing interests
A majority of the authors are employees of Genentech Inc., and hold shares in Roche.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-28. (PDF 4416 kb)
Supplementary Data
This file contains Supplementary Tables 1-17. (XLS 13218 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/).
About this article
Cite this article
Seshagiri, S., Stawiski, E., Durinck, S. et al. Recurrent R-spondin fusions in colon cancer. Nature 488, 660–664 (2012). https://doi.org/10.1038/nature11282
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11282
This article is cited by
-
Recommendations for the classification of germline variants in the exonuclease domain of POLE and POLD1
Genome Medicine (2023)
-
MTAP-ANRIL gene fusion promotes melanoma epithelial-mesenchymal transition-like process by activating the JNK and p38 signaling pathways
Scientific Reports (2023)
-
Transcriptional activity mediated by β-CATENIN and TCF/LEF family members is completely dispensable for survival and propagation of multiple human colorectal cancer cell lines
Scientific Reports (2023)
-
Integrated Analysis Identifies Novel Fusion Transcripts in Laterally Spreading Tumors Suggestive of Distinct Etiology Than Colorectal Cancers
Journal of Gastrointestinal Cancer (2023)
-
Incidence and clinical significance of 491 known fusion genes in a large cohort of Japanese patients with colorectal cancer
International Journal of Clinical Oncology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.