Abstract
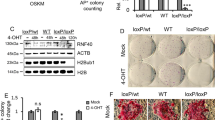
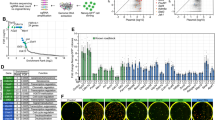
Induced pluripotent stem cells (iPSCs) can be derived from somatic cells by ectopic expression of different transcription factors, classically Oct4 (also known as Pou5f1), Sox2, Klf4 and Myc (abbreviated as OSKM)1. This process is accompanied by genome-wide epigenetic changes2,3,4,5, but how these chromatin modifications are biochemically determined requires further investigation. Here we show in mice and humans that the histone H3 methylated Lys 27 (H3K27) demethylase Utx6,7,8,9 (also known as Kdm6a) regulates the efficient induction, rather than maintenance, of pluripotency. Murine embryonic stem cells lacking Utx can execute lineage commitment and contribute to adult chimaeric animals; however, somatic cells lacking Utx fail to robustly reprogram back to the ground state of pluripotency. Utx directly partners with OSK reprogramming factors and uses its histone demethylase catalytic activity to facilitate iPSC formation. Genomic analysis indicates that Utx depletion results in aberrant dynamics of H3K27me3 repressive chromatin demethylation in somatic cells undergoing reprogramming. The latter directly hampers the derepression of potent pluripotency promoting gene modules (including Sall1, Sall4 and Utf1), which can cooperatively substitute for exogenous OSK supplementation in iPSC formation. Remarkably, Utx safeguards the timely execution of H3K27me3 demethylation observed in embryonic day 10.5–11 primordial germ cells (PGCs)10, and Utx-deficient PGCs show cell-autonomous aberrant epigenetic reprogramming dynamics during their embryonic maturation in vivo. Subsequently, this disrupts PGC development by embryonic day 12.5, and leads to diminished germline transmission in mouse chimaeras generated from Utx-knockout pluripotent cells. Thus, we identify Utx as a novel mediator with distinct functions during the re-establishment of pluripotency and germ cell development. Furthermore, our findings highlight the principle that molecular regulators mediating loss of repressive chromatin during in vivo germ cell reprogramming can be co-opted during in vitro reprogramming towards ground state pluripotency.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
30 July 2012
The second supplementary information file (Supplementary Tables 1-5) was incorrect and has been updated.
References
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Koche, R. P. et al. Reprogramming factor expression initiates widespread targeted chromatin remodeling. Cell Stem Cell 8, 96–105 (2011)
Hemberger, M., Dean, W. & Reik, W. Epigenetic dynamics of stem cells and cell lineage commitment: digging Waddington’s canal. Nature Rev. Mol. Cell Biol. 10, 526–537 (2009)
Singhal, N. et al. Chromatin-remodeling components of the BAF complex facilitate reprogramming. Cell 141, 943–955 (2010)
Onder, T. T. et al. Chromatin-modifying enzymes as modulators of reprogramming. Nature 483, 598–602 (2012)
Agger, K. et al. UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449, 731–734 (2007)
Hong, S. et al. Identification of JmjC domain-containing UTX and JMJD3 as histone H3 lysine 27 demethylases. Proc. Natl Acad. Sci. USA 104, 18439–18444 (2007)
Sen, G. L., Webster, D. E., Barragan, D. I., Chang, H. Y. & Khavari, P. A. Control of differentiation in a self-renewing mammalian tissue by the histone demethylase JMJD3. Genes Dev. 22, 1865–1870 (2008)
Lan, F. et al. A histone H3 lysine 27 demethylase regulates animal posterior development. Nature 449, 689–694 (2007)
Hajkova, P. et al. Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature 452, 877–881 (2008)
Tesar, P. J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007)
Brons, I. G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007)
Guo, G., Huang, Y., Humphreys, P., Wang, X. & Smith, A. A PiggyBac-based recessive screening method to identify pluripotency regulators. PLoS ONE 6, e18189 (2011)
Lee, S., Lee, J. W. & Lee, S. K. UTX, a histone H3-lysine 27 demethylase, acts as a critical switch to activate the cardiac developmental program. Dev. Cell 22, 25–37 (2011)
Hanna, J. et al. Direct cell reprogramming is a stochastic process amenable to acceleration. Nature 462, 595–601 (2009)
Montgomery, N. D. et al. The murine polycomb group protein Eed is required for global histone H3 lysine-27 methylation. Curr. Biol. 15, 942–947 (2005)
Stadtfeld, M., Maherali, N., Breault, D. T. & Hochedlinger, K. Defining molecular cornerstones during fibroblast to iPS cell reprogramming in mouse. Cell Stem Cell 2, 230–240 (2008)
Silva, J. et al. Nanog is the gateway to the pluripotent ground state. Cell 138, 722–737 (2009)
Hansen, K. H. et al. A model for transmission of the H3K27me3 epigenetic mark. Nature Cell Biol. 10, 1291–1300 (2008)
Sridharan, R. et al. Role of the murine reprogramming factors in the induction of pluripotency. Cell 136, 364–377 (2009)
Hayashi, K., Ohta, H., Kurimoto, K., Aramaki, S. & Saitou, M. Reconstitution of the mouse germ cell specification pathway in culture by pluripotent stem cells. Cell 146, 519–532 (2011)
Leitch, H. G. et al. Embryonic germ cells from mice and rats exhibit properties consistent with a generic pluripotent ground state. Development 137, 2279–2287 (2010)
Irizarry, R. A. et al. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4, 249–264 (2003)
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005)
Mikkelsen, T. S. et al. Genome-wide maps of chromatin state in pluripotent and lineage-committed cells. Nature 448, 553–560 (2007)
Loh, Y. H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nature Genet. 38, 431–440 (2006)
Kim, J., Chu, J., Shen, X., Wang, J. & Orkin, S. H. An extended transcriptional network for pluripotency of embryonic stem cells. Cell 132, 1049–1061 (2008)
Bruce, A. W. et al. Genome-wide analysis of repressor element 1 silencing transcription factor/neuron-restrictive silencing factor (REST/NRSF) target genes. Proc. Natl Acad. Sci. USA 101, 10458–10463 (2004)
Zhao, X. D. et al. Whole-genome mapping of histone H3 Lys4 and 27 trimethylations reveals distinct genomic compartments in human embryonic stem cells. Cell Stem Cell 1, 286–298 (2007)
Mikkelsen, T. S. et al. Dissecting direct reprogramming through integrative genomic analysis. Nature 454, 49–55 (2008)
De Santa, F. et al. The histone H3 lysine-27 demethylase Jmjd3 links inflammation to inhibition of polycomb-mediated gene silencing. Cell 130, 1083–1094 (2007)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Robinson, J. T. et al. Integrative genomics viewer. Nature Biotechnol. 29, 24–26 (2011)
He, H. H. et al. Nucleosome dynamics define transcriptional enhancers. Nature Genet. 42, 343–347 (2010)
Acknowledgements
J.H.H. is supported by a generous gift from Ilana and Pascal Mantoux, The Sir Charles Clore Research Prize, an ERC starting investigator grant (StG-2011-281906), the Israel Science Foundation Regular and Bikura research grants, the ICRF Foundation, Fritz Thyssen Stiftung, the Alon Foundation scholar award, a grant from E. A. and R. Drake, and the Leona M. and Harry B. Helmsley Charitable Trust. N.N. and A.P. are supported by Weizmann Dean fellowship awards. We thank A. Smith, H. Niwa and M. Saitou for reagents. We thank the Weizmann Institute management for providing critical financial and infrastructural support.
Author information
Authors and Affiliations
Contributions
O.G., A.A.M., N.N. and J.H.H. concieved the idea for this project, designed and conducted experiments and wrote the manuscript. L.W. conducted PGC differentiation. A.Z. conducted computational simulations. N.N. conducted bioinformatics analysis. O.G., I.A., D.A.-Z., S.H.-S. and E.C. conducted genomic and chromatin immunoprecipiation experiments. A.A.M., A.P., L.H. and I.M. conducted teratoma and embryo analysis. O.G. and S.V. generated knockout cells. J.H.H., O.G., V.K. and Y.R. conducted reprogramming experiments. M.Z. conducted microinjections. S.G. conducted pre-implantation analysis. M.A. provided critical technical advice.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This fie contains Supplementary Figures 1-21 and legends for Supplementary Tables 1-5 (see separate zipped file for tables). (PDF 4398 kb)
Supplementary Tables
This file contains Supplementary Tables 1-5 (see Supplementary Information file for legends). The original incorrect zipped file was replaced on 30 July 2012. (ZIP 2005 kb)
Rights and permissions
About this article
Cite this article
Mansour, A., Gafni, O., Weinberger, L. et al. The H3K27 demethylase Utx regulates somatic and germ cell epigenetic reprogramming. Nature 488, 409–413 (2012). https://doi.org/10.1038/nature11272
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11272
This article is cited by
-
X chromosome dosage and the genetic impact across human tissues
Genome Medicine (2023)
-
Epigenetic markers in the embryonal germ cell development and spermatogenesis
Basic and Clinical Andrology (2023)
-
B1 SINE-binding ZFP266 impedes mouse iPSC generation through suppression of chromatin opening mediated by reprogramming factors
Nature Communications (2023)
-
Dual-Regulated Mechanism of EZH2 and KDM6A on SALL4 Modulates Tumor Progression via Wnt/β-Catenin Pathway in Gastric Cancer
Digestive Diseases and Sciences (2023)
-
Short telomeres impede germ cell specification by upregulating MAPK and TGFβ signaling
Science China Life Sciences (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.