Abstract
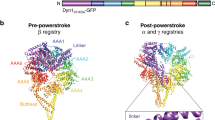
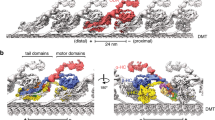
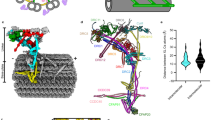
Dyneins are microtubule-based AAA+ motor complexes that power ciliary beating, cell division, cell migration and intracellular transport. Here we report the most complete structure obtained so far, to our knowledge, of the 380-kDa motor domain of Dictyostelium discoideum cytoplasmic dynein at 2.8 Å resolution; the data are reliable enough to discuss the structure and mechanism at the level of individual amino acid residues. Features that can be clearly visualized at this resolution include the coordination of ADP in each of four distinct nucleotide-binding sites in the ring-shaped AAA+ ATPase unit, a newly identified interaction interface between the ring and mechanical linker, and junctional structures between the ring and microtubule-binding stalk, all of which should be critical for the mechanism of dynein motility. We also identify a long-range allosteric communication pathway between the primary ATPase and the microtubule-binding sites. Our work provides a framework for understanding the mechanism of dynein-based motility.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Höök, P. & Vallee, R. B. The dynein family at a glance. J. Cell Sci. 119, 4369–4371 (2006)
Karki, S. & Holzbaur, E. L. Cytoplasmic dynein and dynactin in cell division and intracellular transport. Curr. Opin. Cell Biol. 11, 45–53 (1999)
Vallee, R. B., Williams, J. C., Varma, D. & Barnhart, L. E. Dynein: An ancient motor protein involved in multiple modes of transport. J. Neurobiol. 58, 189–200 (2004)
Scholey, J. M. Intraflagellar transport motors in cilia: moving along the cell’s antenna. J. Cell Biol. 180, 23–29 (2008)
Gibbons, I. R. Cilia and flagella of eukaryotes. J. Cell Biol. 91, 107–124 (1981)
DiBella, L. M. & King, S. M. Dynein motors of the Chlamydomonas flagellum. Int. Rev. Cytol. 210, 227–268 (2001)
Neuwald, A. F., Aravind, L., Spouge, J. L. & Koonin, E. V. AAA+: A class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res. 9, 27–43 (1999)
Tucker, P. A. & Sallai, L. The AAA+ superfamily—a myriad of motions. Curr. Opin. Struct. Biol. 17, 641–652 (2007)
Hanson, P. I. & Whiteheart, S. W. AAA+ proteins: have engine, will work. Nature Rev. Mol. Cell Biol. 6, 519–529 (2005)
Burgess, S. A., Walker, M. L., Sakakibara, H., Knight, P. J. & Oiwa, K. Dynein structure and power stroke. Nature 421, 715–718 (2003)
Samsó, M. & Koonce, M. P. 25 Angstrom resolution structure of a cytoplasmic dynein motor reveals a seven-member planar ring. J. Mol. Biol. 340, 1059–1072 (2004)
Roberts, A. J. et al. AAA+ ring and linker swing mechanism in the dynein motor. Cell 136, 485–495 (2009)
Gee, M. A., Heuser, J. E. & Vallee, R. B. An extended microtubule-binding structure within the dynein motor domain. Nature 390, 636–639 (1997)
Koonce, M. P. Identification of a microtubule-binding domain in a cytoplasmic dynein heavy chain. J. Biol. Chem. 272, 19714–19718 (1997)
Kon, T., Mogami, T., Ohkura, R., Nishiura, M. & Sutoh, K. ATP hydrolysis cycle-dependent tail motions in cytoplasmic dynein. Nature Struct. Mol. Biol. 12, 513–519 (2005)
Carter, A. P., Cho, C., Jin, L. & Vale, R. D. Crystal structure of the dynein motor domain. Science 331, 1159–1165 (2011)
Kon, T., Sutoh, K. & Kurisu, G. X-ray structure of a functional full-length dynein motor domain. Nature Struct. Mol. Biol. 18, 638–642 (2011)
Koonce, M. P. & Samso, M. Overexpression of cytoplasmic dynein’s globular head causes a collapse of the interphase microtubule network in Dictyostelium . Mol. Biol. Cell 7, 935–948 (1996)
Kon, T., Shima, T. & Sutoh, K. Protein engineering approaches to study the dynein mechanism using a dictyostelium expression system. Methods Cell Biol. 92, 65–82 (2009)
Kon, T., Nishiura, M., Ohkura, R., Toyoshima, Y. Y. & Sutoh, K. Distinct functions of nucleotide-binding/hydrolysis sites in the four AAA modules of cytoplasmic dynein. Biochemistry 43, 11266–11274 (2004)
Iyer, L. M., Leipe, D. D., Koonin, E. V. & Aravind, L. Evolutionary history and higher order classification of AAA+ ATPases. J. Struct. Biol. 146, 11–31 (2004)
Erzberger, J. P. & Berger, J. M. Evolutionary relationships and structural mechanisms of AAA+ proteins. Annu. Rev. Biophys. Biomol. Struct. 35, 93–114 (2006)
Mocz, G. & Gibbons, I. R. Phase partition analysis of nucleotide binding to axonemal dynein. Biochemistry 35, 9204–9211 (1996)
Paschal, B. M. & Vallee, R. B. Retrograde transport by the microtubule-associated protein MAP 1C. Nature 330, 181–183 (1987)
Shpetner, H. S., Paschal, B. M. & Vallee, R. B. Characterization of the microtubule-activated ATPase of brain cytoplasmic dynein (MAP 1C). J. Cell Biol. 107, 1001–1009 (1988)
Ogura, T., Whiteheart, S. W. & Wilkinson, A. J. Conserved arginine residues implicated in ATP hydrolysis, nucleotide-sensing, and inter-subunit interactions in AAA and AAA+ ATPases. J. Struct. Biol. 146, 106–112 (2004)
Gai, D., Zhao, R., Li, D., Finkielstein, C. V. & Chen, X. S. Mechanisms of conformational change for a replicative hexameric helicase of SV40 large tumor antigen. Cell 119, 47–60 (2004)
Suno, R. et al. Structure of the whole cytosolic region of ATP-dependent protease FtsH. Mol. Cell 22, 575–585 (2006)
Enemark, E. J. & Joshua-Tor, L. On helicases and other motor proteins. Curr. Opin. Struct. Biol. 18, 243–257 (2008)
Gibbons, I. R. et al. The affinity of the dynein microtubule-binding domain is modulated by the conformation of its coiled-coil stalk. J. Biol. Chem. 280, 23960–23965 (2005)
Carter, A. P. et al. Structure and functional role of dynein’s microtubule-binding domain. Science 322, 1691–1695 (2008)
Kon, T. et al. Helix sliding in the stalk coiled coil of dynein couples ATPase and microtubule binding. Nature Struct. Mol. Biol. 16, 325–333 (2009)
Numata, N., Shima, T., Ohkura, R., Kon, T. & Sutoh, K. C-sequence of the Dictyostelium cytoplasmic dynein participates in processivity modulation. FEBS Lett. 585, 1185–1190 (2011)
Vonrhein, C., Blanc, E., Roversi, P. & Bricogne, G. Automated structure solution with autoSHARP. Methods Mol. Biol. 364, 215–230 (2007)
Brünger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Schröder, G. F., Levitt, M. & Brunger, A. T. Super-resolution biomolecular crystallography with low-resolution data. Nature 464, 1218–1222 (2010)
Imamula, K., Kon, T., Ohkura, R. & Sutoh, K. The coordination of cyclic microtubule association/dissociation and tail swing of cytoplasmic dynein. Proc. Natl Acad. Sci. USA 104, 16134–16139 (2007)
White, H. D., Belknap, B. & Webb, M. R. Kinetics of nucleoside triphosphate cleavage and phosphate release steps by associated rabbit skeletal actomyosin, measured using a novel fluorescent probe for phosphate. Biochemistry 36, 11828–11836 (1997)
Blaauw, M., Linskens, M. H. & van Haastert, P. J. Efficient control of gene expression by a tetracycline-dependent transactivator in single Dictyostelium discoideum cells. Gene 252, 71–82 (2000)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D 59, 2023–2030 (2003)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Arnold, K., Bordoli, L., Kopp, J. & Schwede, T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 22, 195–201 (2006)
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007)
McDonald, I. K. & Thornton, J. M. Satisfying hydrogen bonding potential in proteins. J. Mol. Biol. 238, 777–793 (1994)
Wallace, A. C., Laskowski, R. A. & Thornton, J. M. LIGPLOT: a program to generate schematic diagrams of protein-ligand interactions. Protein Eng. 8, 127–134 (1995)
Kabsch, W. & Sander, C. Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22, 2577–2637 (1983)
Walshaw, J. & Woolfson, D. N. Socket: a program for identifying and analysing coiled-coil motifs within protein structures. J. Mol. Biol. 307, 1427–1450 (2001)
Strelkov, S. V. & Burkhard, P. Analysis of α-helical coiled coils with the program TWISTER reveals a structural mechanism for stutter compensation. J. Struct. Biol. 137, 54–64 (2002)
Schrödinger, L. L. C. The PyMOL Molecular Graphics System, Version 1.3r1 (2010)
Acknowledgements
We thank E. Yamashita, Y. Umena, M. Suzuki and A. Nakagawa of SPring-8 BL-44XU for their support during X-ray data collection; and T. Kikuchi and R. Ohkura for their technical support. We are grateful to C. Toyoshima for discussion of X-ray data collection; K. Kinosita Jr and T. Tsukihara for their support and encouragement. This work was supported by Grants-in-Aid for Scientific Research (17770126, 20687011 and 23370073 (T.K.), 16083205 and 17107003 (K.S.), 17053006, 18054008 and 20051006 (G.K.)) from the Ministry of Education, Culture Sports, Science, and Technology of Japan and a grant from the Human Frontier Science Program (T.K.).
Author information
Authors and Affiliations
Contributions
T.K., K.S. and G.K. designed the study. T.K. purified, crystallized and collected X-ray data; T.O. and G.K. processed and refined X-ray data; T.K, R.S.-K, K.I. and T.S. performed functional analyses; T.K., K.S. and G.K. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1–13, Supplementary Methods, Supplementary Tables 1–2, and Supplementary References. (PDF 6364 kb)
Rights and permissions
About this article
Cite this article
Kon, T., Oyama, T., Shimo-Kon, R. et al. The 2.8 Å crystal structure of the dynein motor domain. Nature 484, 345–350 (2012). https://doi.org/10.1038/nature10955
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10955
This article is cited by
-
Lis1 slows force-induced detachment of cytoplasmic dynein from microtubules
Nature Chemical Biology (2024)
-
Distinct dynein complexes defined by DYNLRB1 and DYNLRB2 regulate mitotic and male meiotic spindle bipolarity
Nature Communications (2023)
-
Design of allosteric sites into rotary motor V1-ATPase by restoring lost function of pseudo-active sites
Nature Chemistry (2023)
-
Regulatory mechanisms of the dynein-2 motility by post-translational modification revealed by MD simulation
Scientific Reports (2023)
-
Exome sequencing of families from Ghana reveals known and candidate hearing impairment genes
Communications Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.