Abstract
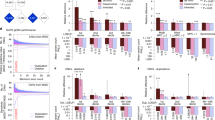
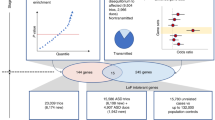
Multiple studies have confirmed the contribution of rare de novo copy number variations to the risk for autism spectrum disorders1,2,3. But whereas de novo single nucleotide variants have been identified in affected individuals4, their contribution to risk has yet to be clarified. Specifically, the frequency and distribution of these mutations have not been well characterized in matched unaffected controls, and such data are vital to the interpretation of de novo coding mutations observed in probands. Here we show, using whole-exome sequencing of 928 individuals, including 200 phenotypically discordant sibling pairs, that highly disruptive (nonsense and splice-site) de novo mutations in brain-expressed genes are associated with autism spectrum disorders and carry large effects. On the basis of mutation rates in unaffected individuals, we demonstrate that multiple independent de novo single nucleotide variants in the same gene among unrelated probands reliably identifies risk alleles, providing a clear path forward for gene discovery. Among a total of 279 identified de novo coding mutations, there is a single instance in probands, and none in siblings, in which two independent nonsense variants disrupt the same gene, SCN2A (sodium channel, voltage-gated, type II, α subunit), a result that is highly unlikely by chance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Accession codes
Primary accessions
Sequence Read Archive
Data deposits
Sequence data from this study is available through the NCBI Sequence Read Archive (accession number SRP010920.1).
References
Sebat, J. et al. Strong association of de novo copy number mutations with autism. Science 316, 445–449 (2007)
Pinto, D. et al. Functional impact of global rare copy number variation in autism spectrum disorders. Nature 466, 368–372 (2010)
Sanders, S. J. et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron 70, 863–885 (2011)
O'Roak, B. J. et al. Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nature Genet. 43, 585–589 (2011)
Fischbach, G. D. & Lord, C. The Simons Simplex Collection: a resource for identification of autism genetic risk factors. Neuron 68, 192–195 (2010)
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009)
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009)
Meisler, M. H., O’Brien, J. E. & Sharkey, L. M. Sodium channel gene family: epilepsy mutations, gene interactions and modifier effects. J. Physiol. (Lond.) 588, 1841–1848 (2010)
Kamiya, K. et al. A nonsense mutation of the sodium channel gene SCN2A in a patient with intractable epilepsy and mental decline. J. Neurosci. 24, 2690–2698 (2004)
Ogiwara, I. et al. De novo mutations of voltage-gated sodium channel alphaII gene SCN2A in intractable epilepsies. Neurology 73, 1046–1053 (2009)
Weiss, L. A. et al. Sodium channels SCN1A, SCN2A and SCN3A in familial autism. Mol. Psychiatry 8, 186–194 (2003)
Xu, B. et al. Exome sequencing supports a de novo mutational paradigm for schizophrenia. Nature Genet. 43, 864–868 (2011)
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nature Methods 7, 248–249 (2010)
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nature Protocols 4, 1073–1081 (2009)
Cooper, G. M. et al. Single-nucleotide evolutionary constraint scores highlight disease-causing mutations. Nature Methods 7, 250–251 (2010)
Cooper, G. M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005)
Grantham, R. Amino acid difference formula to help explain protein evolution. Science 185, 862–864 (1974)
Abul-Husn, N. S. et al. Systems approach to explore components and interactions in the presynapse. Proteomics 9, 3303–3315 (2009)
Bayés, A. et al. Characterization of the proteome, diseases and evolution of the human postsynaptic density. Nature Neurosci. 14, 19–21 (2011)
Collins, M. O. et al. Molecular characterization and comparison of the components and multiprotein complexes in the postsynaptic proteome. J. Neurochem. 97 (suppl. 1). 16–23 (2006)
Girard, S. L. et al. Increased exonic de novo mutation rate in individuals with schizophrenia. Nature Genet. 43, 860–863 (2011)
Rossin, E. J. et al. Proteins encoded in genomic regions associated with immune-mediated disease physically interact and suggest underlying biology. PLoS Genet. 7, e1001273 (2011)
O’Roak, B. J. et al. Sporadic autism exomes reveal a highly interconnected protein network of de novo mutations. Naturehttp://dx.doi.org/10.1038/nature10989 (this issue)
Kang, H. J. et al. Spatio-temporal transcriptome of the human brain. Nature 478, 483–489 (2011)
Acknowledgements
We are grateful to all of the families participating in the Simons Foundation Autism Research Initiative (SFARI) Simplex Collection (SSC). This work was supported by a grant from the Simons Foundation. R.P.L. is an Investigator of the Howard Hughes Medical Institute. We thank the SSC principal investigators A. L. Beaudet, R. Bernier, J. Constantino, E. H. Cook Jr, E. Fombonne, D. Geschwind, D. E. Grice, A. Klin, D. H. Ledbetter, C. Lord, C. L. Martin, D. M. Martin, R. Maxim, J. Miles, O. Ousley, B. Peterson, J. Piggot, C. Saulnier, M. W. State, W. Stone, J. S. Sutcliffe, C. A. Walsh and E. Wijsman and the coordinators and staff at the SSC sites for the recruitment and comprehensive assessment of simplex families; the SFARI staff, in particular M. Benedetti, for facilitating access to the SSC; Prometheus Research for phenotypic data management and Prometheus Research and the Rutgers University Cell and DNA repository for accessing biomaterials; the Yale Center of Genomic Analysis, in particular M. Mahajan, S. Umlauf, I. Tikhonova and A. Lopez, for generating sequencing data; T. Brooks-Boone, N. Wright-Davis and M. Wojciechowski for their help in administering the project at Yale; I. Hart for support; G. D. Fischbach, A. Packer, J. Spiro, M. Benedetti and M. Carlson for their suggestions throughout; and B. Neale and M. Daly for discussions regarding de novo variation. We also acknowledge T. Lehner and the Autism Sequencing Consortium for providing an opportunity for pre-publication data exchange among the participating groups.
Author information
Authors and Affiliations
Contributions
S.J.S., M.T.M., R.P.L., M.G., D.H.G. and M.W.S. designed the study; M.T.M., A.R.G., J.M., M.R., A.G.E.-S., N.M.D., S.M., M.W., G.O., Y.S., P.E., R.M. and J.O. designed and performed high-throughput sequencing experiments and variant confirmations; S.J.S., M.C., K.B., R.B. and N.C. designed the exome-analysis bioinformatics pipeline; S.J.S., A.J.W., N.N.P., J.L.S., N.T., K.A.M., N.Š., K.R., D.H.G., B.D. and M.W.S. analysed the data; S.J.S., A.J.W., K.R., B.D. and M.W.S. wrote the paper; J.M., M.R., A.J.W., A.R.G., A.G.E.-S. and N.M.D. contributed equally to the study. All authors discussed the results and contributed to editing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-12, Supplementary Methods, Supplementary Tables 1-7, Supplementary Equations, legends for Supplementary Data files 1 and 2 and additional references – See Table of contents for more details. (PDF 4427 kb)
Supplementary Data 1
This file contains quality metrics and sample IDs - see Supplementary information file for full legend. (XLS 673 kb)
Supplementary Data 2
This file contains a list of de novo variants - see Supplementary information file for full legend. (XLS 185 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Sanders, S., Murtha, M., Gupta, A. et al. De novo mutations revealed by whole-exome sequencing are strongly associated with autism. Nature 485, 237–241 (2012). https://doi.org/10.1038/nature10945
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10945
This article is cited by
-
Detection of autism spectrum disorder-related pathogenic trio variants by a novel structure-based approach
Molecular Autism (2024)
-
Neuronal knockdown of Cullin3 as a Drosophila model of autism spectrum disorder
Scientific Reports (2024)
-
Severity of Autism Spectrum Disorder Symptoms Associated with de novo Variants and Pregnancy-Induced Hypertension
Journal of Autism and Developmental Disorders (2024)
-
Endophenotype trait domains for advancing gene discovery in autism spectrum disorder
Journal of Neurodevelopmental Disorders (2023)
-
DGH-GO: dissecting the genetic heterogeneity of complex diseases using gene ontology
BMC Bioinformatics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.