Abstract
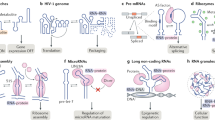
Changes to the conformation of coding and non-coding RNAs form the basis of elements of genetic regulation and provide an important source of complexity, which drives many of the fundamental processes of life. Although the structure of RNA is highly flexible, the underlying dynamics of RNA are robust and are limited to transitions between the few conformations that preserve favourable base-pairing and stacking interactions. The mechanisms by which cellular processes harness the intrinsic dynamic behaviour of RNA and use it within functionally productive pathways are complex. The versatile functions and ease by which it is integrated into a wide variety of genetic circuits and biochemical pathways suggests there is a general and fundamental role for RNA dynamics in cellular processes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kendrew, J. C. et al. A three-dimensional model of the myoglobin molecule obtained by X-ray analysis. Nature 181, 662–666 (1958).
Rould, M. A., Perona, J. J., Söll, D. & Steitz, T. A. Structure of E. coli glutaminyl-tRNA synthetase complexed with tRNA(Gln) and ATP at 2.8 Å resolution. Science 246, 1135–1142 (1989).
Pley, H. W., Flaherty, K. M. & McKay, D. B. Three-dimensional structure of a hammerhead ribozyme. Nature 372, 68–74 (1994).
Scott, W. G., Finch, J. T. & Klug, A. The crystal structure of an all-RNA hammerhead ribozyme: a proposed mechanism for RNA catalytic cleavage. Cell 81, 991–1002 (1995).
Wang, S., Karbstein, K., Peracchi, A., Beigelman, L. & Herschlag, D. Identification of the hammerhead ribozyme metal ion binding site responsible for rescue of the deleterious effect of a cleavage site phosphorothioate. Biochemistry 38, 14363–14378 (1999).
Boehr, D. D., Nussinov, R. & Wright, P. E. The role of dynamic conformational ensembles in biomolecular recognition. Nature Chem. Biol. 5, 789–796 (2009).
Al-Hashimi, H. M. & Walter, N. G. RNA dynamics: it is about time. Curr. Opin. Struct. Biol. 18, 321–329 (2008).
Frauenfelder, H., Sligar, S. G. & Wolynes, P. G. The energy landscapes and motions of proteins. Science 254, 1598–1603 (1991).
Cruz, J. A. & Westhof, E. The dynamic landscapes of RNA architecture. Cell 136, 604–609 (2009).
Bailor, M. H., Mustoe, A. M., Brooks, C. L. 3rd & Al-Hashimi, H. M. Topological constraints: using RNA secondary structure to model 3D conformation, folding pathways, and dynamic adaptation. Curr. Opin. Struct. Biol. 21, 296–305 (2011).
Schultes, E. A., Spasic, A., Mohanty, U. & Bartel, D. P. Compact and ordered collapse of randomly generated RNA sequences. Nature Struct. Mol. Biol. 12, 1130–1136 (2005).
Schultes, E. A., Hraber, P. T. & LaBean, T. H. Estimating the contributions of selection and self-organization in RNA secondary structure. J. Mol. Evol. 49, 76–83 (1999).
Fürtig, B., Wenter, P., Pitsch, S. & Schwalbe, H. Probing mechanism and transition state of RNA refolding. ACS Chem. Biol. 5, 753–765 (2010).
Bailor, M. H., Sun, X. & Al-Hashimi, H. M. Topology links RNA secondary structure with global conformation, dynamics, and adaptation. Science 327, 202–206 (2010). This article reports the simple topological constraints that are governed by steric and stereochemical forces severely restrict the allowed orientation of helices across two-way junctions.
Mustoe, A. M., Bailor, M. H., Teixeira, R. M., Brooks, C. L. 3rd & Al-Hashimi, H. M. New insights into the fundamental role of topological constraints as a determinant of two-way junction conformation. Nucleic Acids Res. 40, 892–904 (2012).
Chu, V. B. et al. Do conformational biases of simple helical junctions influence RNA folding stability and specificity? RNA 15, 2195–2205 (2009).
Venditti, V., Clos, L. 2nd, Niccolai, N. & Butcher, S. E. Minimum-energy path for a U6 RNA conformational change involving protonation, base-pair rearrangement and base flipping. J. Mol. Biol. 391, 894–905 (2009).
Fourmy, D., Yoshizawa, S. & Puglisi, J. D. Paromomycin binding induces a local conformational change in the A-site of 16S rRNA. J. Mol. Biol. 277, 333–345 (1998).
Le, S. Y., Zhang, K. & Maizel, J. V. Jr. RNA molecules with structure dependent functions are uniquely folded. Nucleic Acids Res. 30, 3574–3582 (2002).
Stelzer, A. C., Kratz, J. D., Zhang, Q. & Al-Hashimi, H. M. RNA dynamics by design: biasing ensembles towards the ligand-bound state. Angew. Chem. Int. Ed. Engl. 49, 5731–5733 (2010).
Shankar, N. et al. NMR reveals the absence of hydrogen bonding in adjacent UU and AG mismatches in an isolated internal loop from ribosomal RNA. Biochemistry 46, 12665–12678 (2007).
Frank, J. & Gonzalez, R. L., Jr. Structure and dynamics of a processive Brownian motor: the translating ribosome. Annu. Rev. Biochem. 79, 381–412 (2010).
Haller, A., Souliere, M. F. & Micura, R. The dynamic nature of RNA as key to understanding riboswitch mechanisms. Acc. Chem. Res. 44, 1339–1348 (2011).
Paukstelis, P. J., Chen, J. H., Chase, E., Lambowitz, A. M. & Golden, B. L. Structure of a tyrosyl-tRNA synthetase splicing factor bound to a group I intron RNA. Nature 451, 94–97 (2008).
Puglisi, J. D., Tan, R., Calnan, B. J., Frankel, A. D. & Williamson, J. R. Conformation of the TAR RNA-arginine complex by NMR spectroscopy. Science 257, 76–80 (1992).
Orr, J. W., Hagerman, P. J. & Williamson, J. R. Protein and Mg2+-induced conformational changes in the S15 binding site of 16S ribosomal RNA. J. Mol. Biol. 275, 453–464 (1998).
Turner, B., Melcher, S. E., Wilson, T. J., Norman, D. G. & Lilley, D. M. Induced fit of RNA on binding the L7Ae protein to the kink-turn motif. RNA 11, 1192–1200 (2005).
Falb, M., Amata, I., Gabel, F., Simon, B. & Carlomagno, T. Structure of the K-turn U4 RNA: a combined NMR and SANS study. Nucleic Acids Res. 38, 6274–6285 (2010).
Kim, H. D. et al. Mg2+-dependent conformational change of RNA studied by fluorescence correlation and FRET on immobilized single molecules. Proc. Natl Acad. Sci. USA 99, 4284–4289 (2002).
Zacharias, M. & Hagerman, P. J. The influence of symmetric internal loops on the flexibility of RNA. J. Mol. Biol. 257, 276–289 (1996).
Zhang, Q., Stelzer, A. C., Fisher, C. K. & Al-Hashimi, H. M. Visualizing spatially correlated dynamics that directs RNA conformational transitions. Nature 450, 1263–1267 (2007).
Shajani, Z., Drobny, G. & Varani, G. Binding of U1A protein changes RNA dynamics as observed by 13C NMR relaxation studies. Biochemistry 46, 5875–5883 (2007).
Bokinsky, G. et al. Two distinct binding modes of a protein cofactor with its target RNA. J. Mol. Biol. 361, 771–784 (2006).
Bardaro, M. F. Jr., Shajani, Z., Patora-Komisarska, K., Robinson, J. A. & Varani, G. How binding of small molecule and peptide ligands to HIV-1 TAR alters the RNA motional landscape. Nucleic Acids Res. 37, 1529–1540 (2009).
Herschlag, D., Khosla, M., Tsuchihashi, Z. & Karpel, R. L. An RNA chaperone activity of non-specific RNA binding proteins in hammerhead ribozyme catalysis. EMBO J. 13, 2913–2924 (1994).
Pyle, A. M. & Green, J. B. RNA folding. Curr. Opin. Struct. Biol. 5, 303–310 (1995).
Treiber, D.K. & Williamson, J.R. Beyond kinetic traps in RNA folding. Curr. Opin. Struct. Biol. 11, 309–314 (2001).
Hirling, H., Scheffner, M., Restle, T. & Stahl, H. RNA helicase activity associated with the human p68 protein. Nature 339, 562–564 (1989).
Yang, Q. & Jankowsky, E. ATP- and ADP-dependent modulation of RNA unwinding and strand annealing activities by the DEAD-box protein DED1. Biochemistry 44, 13591–13601 (2005).
Will, C. L. & Lührmann, R. Spliceosome structure and function. Cold Spring Harb. Perspect. Biol. 3, a003707 (2011).
Kosowski, T. R., Keys, H. R., Quan, T. K. & Ruby, S. W. DExD/H-box Prp5 protein is in the spliceosome during most of the splicing cycle. RNA 15, 1345–1362 (2009).
Maeder, C., Kutach, A. K. & Guthrie, C. ATP-dependent unwinding of U4/U6 snRNAs by the Brr2 helicase requires the C terminus of Prp8. Nature Struct. Mol. Biol. 16, 42–48 (2009).
Schwer, B. A conformational rearrangement in the spliceosome sets the stage for Prp22-dependent mRNA release. Mol. Cell 30, 743–754 (2008).
Bhaskaran, H. & Russell, R. Kinetic redistribution of native and misfolded RNAs by a DEAD-box chaperone. Nature 449, 1014–1018 (2007).
Winkler, W., Nahvi, A. & Breaker, R. R. Thiamine derivatives bind messenger RNAs directly to regulate bacterial gene expression. Nature 419, 952–956 (2002). This article reports the discovery of an RNA switch in the 5′ untranslated region of bacterial mRNA that regulates gene expression in response to ligands without assistance from proteins.
Cromie, M. J., Shi, Y., Latifi, T. & Groisman, E. A. An RNA sensor for intracellular Mg2+. Cell 125, 71–84 (2006).
Nechooshtan, G., Elgrably-Weiss, M., Sheaffer, A., Westhof, E. & Altuvia, S. A pH-responsive riboregulator. Genes Dev. 23, 2650–2662 (2009).
Greenleaf, W. J., Frieda, K. L., Foster, D. A. Woodside, M. T. & Block, S. M. Direct observation of hierarchical folding in single riboswitch aptamers. Science 319, 630–633 (2008).
Mandal, M. et al. A glycine-dependent riboswitch that uses cooperative binding to control gene expression. Science 306, 275–279 (2004).
Sudarsan, N. et al. Tandem riboswitch architectures exhibit complex gene control functions. Science 314, 300–304 (2006).
Lee, E. R., Baker, J. L., Weinberg, Z., Sudarsan, N. & Breaker, R. R. An allosteric self-splicing ribozyme triggered by a bacterial second messenger. Science 329, 845–848 (2010).
Ferre-D'Amare, A. R., Zhou, K. & Doudna, J. A. Crystal structure of a hepatitis delta virus ribozyme. Nature 395, 567–574 (1998).
Ke, A., Zhou, K., Ding, F., Cate, J. H. D. & Doudna, J. A. A conformational switch controls hepatitis delta virus ribozyme catalysis. Nature 429, 201–205 (2004). This article reports a significant local conformational change in the active site of the HDV ribozyme is observed post-cleavage and is associated with ejection of the substrate and a catalytically critical divalent metal ion.
Harris, D. A., Rueda, D. & Walter, N. G. Local conformational changes in the catalytic core of the trans-acting hepatitis delta virus ribozyme accompany catalysis. Biochemistry 41, 12051–12061 (2002).
Lamanna, A. C. & Karbstein, K. An RNA conformational switch regulates pre-18S rRNA cleavage. J. Mol. Biol. 405, 3–17 (2011).
Nocker, A. et al. A mRNA-based thermosensor controls expression of rhizobial heat shock genes. Nucleic Acids Res. 29, 4800–4807 (2001).
Johansson, J. et al. An RNA thermosensor controls expression of virulence genes in Listeria monocytogenes. Cell 110, 551–561 (2002).
Watts, J. M. et al. Architecture and secondary structure of an entire HIV-1 RNA genome. Nature 460, 711–716 (2009).
Grundy, F. J., Winkler, W. C. & Henkin, T. M. tRNA-mediated transcription antitermination in vitro: codon-anticodon pairing independent of the ribosome. Proc. Natl Acad. Sci. USA 99, 11121–11126 (2002).
Babitzke, P. & Yanofsky, C. Reconstitution of Bacillus subtilis trp attenuation in vitro with TRAP, the trp RNA-binding attenuation protein. Proc. Natl Acad. Sci. USA 90, 133–137 (1993).
Diaz-Toledano, R., Ariza-Mateos, A., Birk, A., Martinez-Garcia, B. & Gomez, J. In vitro characterization of a miR-122-sensitive double-helical switch element in the 5´ region of hepatitis C virus RNA. Nucleic Acids Res. 37, 5498–5510 (2009).
Ray, P. S. et al. A stress-responsive RNA switch regulates VEGFA expression. Nature 457, 915–919 (2009). This article reports that the 3′ untranslated region of human VEGFA mRNA undergoes a binary conformational switch in response to inflammatory and hypoxic protein stress signals to regulate VEGFA expression.
Cheah, M. T., Wachter, A., Sudarsan, N. & Breaker, R. R. Control of alternative RNA splicing and gene expression by eukaryotic riboswitches. Nature 447, 497–500 (2007). This article reports a secondary structural change in a eukaryotic thiamine pyrophosphate riboswitch regulates gene expression through the control of alternative splicing.
Kedde, M. et al. A Pumilio-induced RNA structure switch in p27-3′ untranslated region controls miR-221 and miR-222 accessibility. Nature Cell Biol. 12, 1014–1020 (2010).
Casey, J. L. Control of ADAR1 editing of hepatitis delta virus RNAs. Curr. Top. Microbiol. Immunol. 353, 123–143 (2012).
Abbink, T. E., Ooms, M., Haasnoot, P. C. & Berkhout, B. The HIV-1 leader RNA conformational switch regulates RNA dimerization but does not regulate mRNA translation. Biochemistry 44, 9058–9066 (2005).
Miyazaki, Y. et al. An RNA structural switch regulates diploid genome packaging by Moloney murine leukemia virus. J. Mol. Biol. 396, 141–152 (2010). This article reports that dimerization of the 5′ untranslated region of the Moloney murine leukaemia virus results in a secondary structural change that promotes genome packaging.
Giege, R. Toward a more complete view of tRNA biology. Nature Struct. Mol. Biol. 15, 1007–1014 (2008).
Mulder, A. M. et al. Visualizing ribosome biogenesis: parallel assembly pathways for the 30S subunit. Science 330, 673–677 (2010).
Adilakshmi, T., Bellur, D. L. & Woodson, S. A. Concurrent nucleation of 16S folding and induced fit in 30S ribosome assembly. Nature 455, 1268–1272 (2008).
Menichelli, E., Isel, C., Oubridge, C. & Nagai, K. Protein-induced conformational changes of RNA during the assembly of human signal recognition particle. J. Mol. Biol. 367, 187–203 (2007).
Stone, M. D. et al. Stepwise protein-mediated RNA folding directs assembly of telomerase ribonucleoprotein. Nature 446, 458–461 (2007).
Held, W. A., Ballou, B., Mizushima, S. & Nomura, M. Assembly mapping of 30S ribosomal proteins from Escherichia coli. Further studies. J. Biol. Chem. 249, 3103–3111 (1974).
Agalarov, S. C., Prasad, G. S., Funke, P. M., Stout, C. D. & Williamson, J. R. Structure of the S15,S6,S18-rRNA complex: assembly of the 30S ribosome central domain. Science 288, 107–112 (2000).
Duncan, C. D. & Weeks, K. M. Nonhierarchical ribonucleoprotein assembly suggests a strain-propagation model for protein-facilitated RNA folding. Biochemistry 49, 5418–5425 (2010).
Wilson, T. J., Nahas, M., Ha, T. & Lilley, D. M. Folding and catalysis of the hairpin ribozyme. Biochem. Soc. Trans. 33, 461–465 (2005).
Zhang, Q., Kim, N. K., Peterson, R. D., Wang, Z. & Feigon, J. Structurally conserved five nucleotide bulge determines the overall topology of the core domain of human telomerase RNA. Proc. Natl Acad. Sci. USA 107, 18761–18768 (2010).
Solomatin, S. V., Greenfeld, M., Chu, S. & Herschlag, D. Multiple native states reveal persistent ruggedness of an RNA folding landscape. Nature 463, 681–684 (2010). This article reports the observation of slowly interconverting catalytically active states in a ribozyme, thereby establishing the coexistence of multiple native states.
Greenfeld, M., Solomatin, S. V. & Herschlag, D. Removal of covalent heterogeneity reveals simple folding behavior for P4–P6 RNA. J. Biol. Chem. 286, 19872–19879 (2011).
Frank, J. & Agrawal, R. K. A ratchet-like inter-subunit reorganization of the ribosome during translocation. Nature 406, 318–322 (2000).
Valle, M. et al. Locking and unlocking of ribosomal motions. Cell 114, 123–134 (2003).
Zhang, W., Dunkle, J. A. & Cate, J. H. Structures of the ribosome in intermediate states of ratcheting. Science 325, 1014–1017 (2009).
Ratje, A. H. et al. Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites. Nature 468, 713–716 (2010).
Fischer, N., Konevega, A. L. Wintermeyer, W., Rodnina, M. V. & Stark, H. Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy. Nature 466, 329–333 (2010). This article demonstrates the cryo-electron microscopy observation of thermally driven tRNA retrotranslocation on the ribosome.
Shoji, S., Walker, S. E. & Fredrick, K. Reverse translocation of tRNA in the ribosome. Mol. Cell 24, 931–942 (2006).
Ogle, J. M., Murphy, F. V., Tarry, M. J. & Ramakrishnan, V. Selection of tRNA by the ribosome requires a transition from an open to a closed form. Cell 111, 721–732 (2002).
Valle, M. et al. Incorporation of aminoacyl-tRNA into the ribosome as seen by cryo-electron microscopy. Nature Struct. Biol. 10, 899–906 (2003).
Lee, T. H., Blanchard, S. C., Kim, H. D., Puglisi, J. D. & Chu, S. The role of fluctuations in tRNA selection by the ribosome. Proc. Natl Acad. Sci. USA 104, 13661–13665 (2007).
Schmeing, T. M. et al. The crystal structure of the ribosome bound to EF-Tu and aminoacyl-tRNA. Science 326, 688–694 (2009).
Voorhees, R. M., Schmeing, T. M., Kelley, A. C. & Ramakrishnan, V. The mechanism for activation of GTP hydrolysis on the ribosome. Science 330, 835–838 (2010).
Pape, T., Wintermeyer, W. & Rodnina, M. V. Conformational switch in the decoding region of 16S rRNA during aminoacyl-tRNA selection on the ribosome. Nature Struct. Mol. Biol. 7, 104–107 (2000).
Blanchard, S. C., Gonzalez, R. L., Kim, H. D., Chu, S. & Puglisi, J. D. tRNA selection and kinetic proofreading in translation. Nature Struct. Mol. Biol. 11, 1008–1014 (2004). This important single-molecule FRET study directly observes the dynamics of tRNA initial selection and proofreading by the ribosome.
Fei, J. et al., Allosteric collaboration between elongation factor G and the ribosomal L1 stalk directs tRNA movements during translation. Proc. Natl Acad. Sci. USA 106, 15702–15707 (2009).
Blaha, G., Stanley, R. E. & Steitz, T. A. Formation of the first peptide bond: the structure of EF-P bound to the 70S ribosome. Science 325, 966–970 (2009).
Dunkle, J. A. et al. Structures of the bacterial ribosome in classical and hybrid states of tRNA binding. Science 332, 981–984 (2011).
Laurberg, M. et al. Structural basis for translation termination on the 70S ribosome. Nature 454, 852–857 (2008).
Cornish, P. V., Ermolenko, D. N., Noller, H. F. & Ha, T. Spontaneous intersubunit rotation in single ribosomes. Mol. Cell 30, 578–588 (2008).
Tama, F., Valle, M., Frank, J. & Brooks, C. L 3rd. Dynamic reorganization of the functionally active ribosome explored by normal mode analysis and cryo-electron microscopy. Proc. Natl Acad. Sci. USA 100, 9319–9323 (2003).
Green, N. J., Grundy, F. J. & Henkin, T. M. The T box mechanism: tRNA as a regulatory molecule. FEBS Lett. 584, 318–324 (2010).
Cornish, P. V. et al. Following movement of the L1 stalk between three functional states in single ribosomes. Proc. Natl Acad. Sci. USA 106, 2571–2576 (2009).
Acknowledgements
E.A.D. and J.C. contributed equally to this Review. We thank C. Eichhorn and Q. Zhang for their input and assistance in the preparation of figures, and S. Butcher and S. Serganov for their comments on this Review. A.M.M. is supported by an NSF graduate research fellowship. The authors gratefully acknowledge the Michigan Economic Development Cooperation and the Michigan Technology Tri-Corridor for their support in the purchase of a 600 MHz spectrometer. This work was supported by the US National Institutes of Health (R01 AI066975 and R01 GM089846) and the US National Science Foundation (NSF Career Award CHE-0918817).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reprints and permissions information is available at www.nature.com/reprints.
Supplementary information
Rights and permissions
About this article
Cite this article
Dethoff, E., Chugh, J., Mustoe, A. et al. Functional complexity and regulation through RNA dynamics. Nature 482, 322–330 (2012). https://doi.org/10.1038/nature10885
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10885
This article is cited by
-
Orthogonal inducible control of Cas13 circuits enables programmable RNA regulation in mammalian cells
Nature Communications (2024)
-
A guideline for the distance measurement plans of site-directed spin labels for structural prediction of nucleic acids
Journal of Molecular Modeling (2024)
-
Identification of genetic associations and functional SNPs of bovine KLF6 gene on milk production traits in Chinese holstein
BMC Genomic Data (2023)
-
Probing the dynamic RNA structurome and its functions
Nature Reviews Genetics (2023)
-
CtIP-dependent nascent RNA expression flanking DNA breaks guides the choice of DNA repair pathway
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.