Abstract
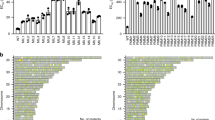
The concept of disease-specific chemotherapy was developed a century ago. Dyes and arsenical compounds that displayed selectivity against trypanosomes were central to this work1,2, and the drugs that emerged remain in use for treating human African trypanosomiasis (HAT)3. The importance of understanding the mechanisms underlying selective drug action and resistance for the development of improved HAT therapies has been recognized, but these mechanisms have remained largely unknown. Here we use all five current HAT drugs for genome-scale RNA interference target sequencing (RIT-seq) screens in Trypanosoma brucei, revealing the transporters, organelles, enzymes and metabolic pathways that function to facilitate antitrypanosomal drug action. RIT-seq profiling identifies both known drug importers4,5 and the only known pro-drug activator6, and links more than fifty additional genes to drug action. A bloodstream stage-specific invariant surface glycoprotein (ISG75) family mediates suramin uptake, and the AP1 adaptin complex, lysosomal proteases and major lysosomal transmembrane protein, as well as spermidine and N-acetylglucosamine biosynthesis, all contribute to suramin action. Further screens link ubiquinone availability to nitro-drug action, plasma membrane P-type H+-ATPases to pentamidine action, and trypanothione and several putative kinases to melarsoprol action. We also demonstrate a major role for aquaglyceroporins in pentamidine and melarsoprol cross-resistance. These advances in our understanding of mechanisms of antitrypanosomal drug efficacy and resistance will aid the rational design of new therapies and help to combat drug resistance, and provide unprecedented molecular insight into the mode of action of antitrypanosomal drugs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
Sequence data from this study have been submitted to the European Nucleotide Archive at http://www.ebi.ac.uk/ena under accession number ERA071064.
References
Ehrlich, P. Address in pathology, on chemiotherapy: delivered before the seventeenth international congress of medicine. BMJ 2, 353–359 (1913)
Williamson, J. in The African Trypanosomiases (ed. Mulligan, H. W.) (Allen and Unwin, 1970)
Fairlamb, A. H. Chemotherapy of human African trypanosomiasis: current and future prospects. Trends Parasitol. 19, 488–494 (2003)
Mäser, P., Sutterlin, C., Kralli, A. & Kaminsky, R. A nucleoside transporter from Trypanosoma brucei involved in drug resistance. Science 285, 242–244 (1999)
Vincent, I. M. et al. A molecular mechanism for eflornithine resistance in African trypanosomes. PLoS Pathog. 6, e1001204 (2010)
Wilkinson, S. R., Taylor, M. C., Horn, D., Kelly, J. M. & Cheeseman, I. A mechanism for cross-resistance to nifurtimox and benznidazole in trypanosomes. Proc. Natl Acad. Sci. USA 105, 5022–5027 (2008)
Fèvre, E. M., Wissmann, B. V., Welburn, S. C. & Lutumba, P. The burden of human African trypanosomiasis. PLoS Negl. Trop. Dis. 2, e333 (2008)
Pépin, J. & Milord, F. The treatment of human African trypanosomiasis. Adv. Parasitol. 33, 1–47 (1994)
de Koning, H. P. Ever-increasing complexities of diamidine and arsenical crossresistance in African trypanosomes. Trends Parasitol. 24, 345–349 (2008)
Alsford, S. et al. High-throughput phenotyping using parallel sequencing of RNA interference targets in the African trypanosome. Genome Res. 21, 915–924 (2011)
Carter, N. S. & Fairlamb, A. H. Arsenical-resistant trypanosomes lack an unusual adenosine transporter. Nature 361, 173–176 (1993)
Matovu, E. et al. Mechanisms of arsenical and diamidine uptake and resistance in Trypanosoma brucei. Eukaryot. Cell 2, 1003–1008 (2003)
Schumann Burkard, G., Jutzi, P. & Roditi, I. Genome-wide RNAi screens in bloodstream form trypanosomes identify drug transporters. Mol. Biochem. Parasitol. 175, 91–94 (2011)
Baker, N., Alsford, S. & Horn, D. Genome-wide RNAi screens in African trypanosomes identify the nifurtimox activator NTR and the eflornithine transporter AAT6. Mol. Biochem. Parasitol. 176, 55–57 (2011)
Steverding, D. The development of drugs for treatment of sleeping sickness: a historical review. Parasit. Vectors 3, 15 (2010)
Overath, P., Chaudhri, M., Steverding, D. & Ziegelbauer, K. Invariant surface proteins in bloodstream forms of Trypanosoma brucei. Parasitol. Today 10, 53–58 (1994)
Peck, R. F. et al. The LAMP-like protein p67 plays an essential role in the lysosome of African trypanosomes. Mol. Microbiol. 68, 933–946 (2008)
Leung, K. F., Riley, F. S., Carrington, M. & Field, M. C. Ubiquitylation and developmental regulation of invariant surface protein expression in trypanosomes. Eukaryot. Cell 10, 916–931 (2011)
Koumandou, V. L. et al. Evolutionary reconstruction of the retromer complex and its function in Trypanosoma brucei. J. Cell Sci. 124, 1496–1509 (2011)
Caffrey, C. R. et al. Active site mapping, biochemical properties and subcellular localization of rhodesain, the major cysteine protease of Trypanosoma brucei rhodesiense. Mol. Biochem. Parasitol. 118, 61–73 (2001)
Fairlamb, A. H. & Bowman, I. B. Uptake of the trypanocidal drug suramin by bloodstream forms of Trypanosoma brucei and its effect on respiration and growth rate in vivo. Mol. Biochem. Parasitol. 1, 315–333 (1980)
Vansterkenburg, E. L. et al. The uptake of the trypanocidal drug suramin in combination with low-density lipoproteins by Trypanosoma brucei and its possible mode of action. Acta Trop. 54, 237–250 (1993)
Scott, A. G., Tait, A. & Turner, C. M. Characterisation of cloned lines of Trypanosoma brucei expressing stable resistance to MelCy and suramin. Acta Trop. 60, 251–262 (1996)
Natesan, S. K., Peacock, L., Matthews, K., Gibson, W. & Field, M. C. Activation of endocytosis as an adaptation to the mammalian host by trypanosomes. Eukaryot. Cell 6, 2029–2037 (2007)
Uzcategui, N. L. et al. Cloning, heterologous expression, and characterization of three aquaglyceroporins from Trypanosoma brucei. J. Biol. Chem. 279, 42669–42676 (2004)
Lanteri, C. A., Tidwell, R. R. & Meshnick, S. R. The mitochondrion is a site of trypanocidal action of the aromatic diamidine DB75 in bloodstream forms of Trypanosoma brucei. Antimicrob. Agents Chemother. 52, 875–882 (2008)
Luo, S., Fang, J. & Docampo, R. Molecular characterization of Trypanosoma brucei P-type H+-ATPases. J. Biol. Chem. 281, 21963–21973 (2006)
Fairlamb, A. H., Henderson, G. B. & Cerami, A. Trypanothione is the primary target for arsenical drugs against African trypanosomes. Proc. Natl Acad. Sci. USA 86, 2607–2611 (1989)
Morris, J. C., Wang, Z., Drew, M. E. & Englund, P. T. Glycolysis modulates trypanosome glycoprotein expression as revealed by an RNAi library. EMBO J. 21, 4429–4438 (2002)
Alsford, S., Kawahara, T., Glover, L. & Horn, D. Tagging a T. brucei RRNA locus improves stable transfection efficiency and circumvents inducible expression position effects. Mol. Biochem. Parasitol. 144, 142–148 (2005)
Berriman, M. et al. The genome of the African trypanosome Trypanosoma brucei. Science 309, 416–422 (2005)
Ning, Z., Cox, A. J. & Mullikin, J. C. SSAHA: a fast search method for large DNA databases. Genome Res. 11, 1725–1729 (2001)
Carver, T. et al. Artemis and ACT: viewing, annotating and comparing sequences stored in a relational database. Bioinformatics 24, 2672–2676 (2008)
Buschini, A. et al. Genotoxicity revaluation of three commercial nitroheterocyclic drugs: nifurtimox, benznidazole, and metronidazole. J. Parasitol. Res. 2009, 463575 (2009)
Redmond, S., Vadivelu, J. & Field, M. C. RNAit: an automated web-based tool for the selection of RNAi targets in Trypanosoma brucei. Mol. Biochem. Parasitol. 128, 115–118 (2003)
Alsford, S. & Horn, D. Single-locus targeting constructs for reliable regulated RNAi and transgene expression in Trypanosoma brucei. Mol. Biochem. Parasitol. 161, 76–79 (2008)
Mackey, Z. B., O'Brien, T. C., Greenbaum, D. C., Blank, R. B. & McKerrow, J. H. A cathepsin B-like protease is required for host protein degradation in Trypanosoma brucei. J. Biol. Chem. 279, 48426–48433 (2004)
Singh, P. K., Tack, B. F., McCray, P. B., Jr & Welsh, M. J. Synergistic and additive killing by antimicrobial factors found in human airway surface liquid. Am. J. Physiol. Lung Cell. Mol. Physiol. 279, L799–L805 (2000)
Räz, B., Iten, M., Grether-Buhler, Y., Kaminsky, R. & Brun, R. The Alamar Blue assay to determine drug sensitivity of African trypanosomes (T. b. rhodesiense and T. b. gambiense) in vitro.. Acta Trop. 68, 139–147 (1997)
Ausubel, F. M., Brent, R., Kingston, R. E., Moore, D. D., Seidman, J. G., Smith, J. A., Struhl, K., eds. Current Protocols in Molecular Biology (John Wiley and Sons, 1998).
Leung, K. F., Dacks, J. B. & Field, M. C. Evolution of the multivesicular body ESCRT machinery; retention across the eukaryotic lineage. Traffic 9, 1698–1716 (2008)
Ziegelbauer, K. & Overath, P. Organization of two invariant surface glycoproteins in the surface coat of Trypanosoma brucei. Infect. Immun. 61, 4540–4545 (1993)
Kelley, R. J., Brickman, M. J. & Balber, A. E. Processing and transport of a lysosomal membrane glycoprotein is developmentally regulated in African trypanosomes. Mol. Biochem. Parasitol. 74, 167–178 (1995)
Lingnau, A., Zufferey, R., Lingnau, M. & Russell, D. G. Characterization of tGLP-1, a Golgi and lysosome-associated, transmembrane glycoprotein of African trypanosomes. J. Cell Sci. 112, 3061–3070 (1999)
Allen, C. L., Liao, D., Chung, W. L. & Field, M. C. Dileucine signal-dependent and AP-1-independent targeting of a lysosomal glycoprotein in Trypanosoma brucei. Mol. Biochem. Parasitol. 156, 175–190 (2007)
Acknowledgements
We thank J. Morris, Z. Wang, M. Drew and P. Englund for the RNAi plasmid library, V. Yardley for antitrypanosomal drugs, J. Bangs for anti-p67 and CatL sera, D. Russell for anti-GLP1 sera, A. Varghese for assistance with preliminary Sanger sequencing and J. Kelly, M. Taylor and B. Wren for comments on the draft manuscript. The work was funded by grants from The Wellcome Trust (093010/Z/10/Z at the London School of Hygiene & Tropical Medicine, 085775/Z/08/Z at The Wellcome Trust Sanger Institute and 090007/Z/09/Z at The University of Cambridge). N.B. was supported by a Bloomsbury colleges PhD studentship.
Author information
Authors and Affiliations
Contributions
S.A., N.B., L.G. and K.F.L. carried out the T. brucei manipulation and analyses, S.E., A.S.-F. and D.J.T. carried out the Illumina sequencing and mapping, D.H. coordinated the study and S.A., M.C.F., M.B. and D.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-6 with legends, Supplementary Table 1 and additional references. Growth curves, sequence read-density signatures, EC50 data and comparative genomic information and HAT drug information are included. (PDF 497 kb)
Supplementary Data 1
This file shows two spreadsheets. The first (a) shows all 'primary' and 'secondary' hits from the HAT drug resistance screens with comments and links to databases. The second (b) shows all genes associated with >9 sequence reads in the HAT drug resistance screens. (XLS 362 kb)
Rights and permissions
About this article
Cite this article
Alsford, S., Eckert, S., Baker, N. et al. High-throughput decoding of antitrypanosomal drug efficacy and resistance. Nature 482, 232–236 (2012). https://doi.org/10.1038/nature10771
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10771
This article is cited by
-
First comprehensive untargeted metabolomics study of suramin-treated Trypanosoma brucei: an integrated data analysis workflow from multifactor data modelling to functional analysis
Metabolomics (2024)
-
Identification of a small-molecule inhibitor that selectively blocks DNA-binding by Trypanosoma brucei replication protein A1
Nature Communications (2023)
-
Arsenic in medicine: past, present and future
BioMetals (2023)
-
Trypanosoma brucei: Metabolomics for analysis of cellular metabolism and drug discovery
Metabolomics (2022)
-
Structure of trypanosome coat protein VSGsur and function in suramin resistance
Nature Microbiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.