Abstract
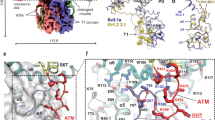
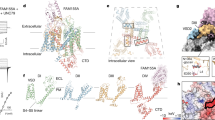
The KCNH family of ion channels, comprising ether-à-go-go (EAG), EAG-related gene (ERG), and EAG-like (ELK) K+-channel subfamilies, is crucial for repolarization of the cardiac action potential1, regulation of neuronal excitability2 and proliferation of tumour cells3. The carboxy-terminal region of KCNH channels contains a cyclic-nucleotide-binding homology domain (CNBHD) and C-linker that couples the CNBHD to the pore4. The C-linker/CNBHD is essential for proper function and trafficking of ion channels in the KCNH family5,6,7,8,9. However, despite the importance of the C-linker/CNBHD for the function of KCNH channels, the structural basis of ion-channel regulation by the C-linker/CNBHD is unknown. Here we report the crystal structure of the C-linker/CNBHD of zebrafish ELK channels at 2.2-Å resolution. Although the overall structure of the C-linker/CNBHD of ELK channels is similar to the cyclic-nucleotide-binding domain (CNBD) structure of the related hyperpolarization-activated cyclic-nucleotide-modulated (HCN) channels10, there are marked differences. Unlike the CNBD of HCN, the CNBHD of ELK displays a negatively charged electrostatic profile that explains the lack of binding and regulation of KCNH channels by cyclic nucleotides4,11. Instead of cyclic nucleotide, the binding pocket is occupied by a short β-strand. Mutations of the β-strand shift the voltage dependence of activation to more depolarized voltages, implicating the β-strand as an intrinsic ligand for the CNBHD of ELK channels. In both ELK and HCN channels the C-linker is the site of virtually all of the intersubunit interactions in the C-terminal region. However, in the zebrafish ELK structure there is a reorientation in the C-linker so that the subunits form dimers instead of tetramers, as observed in HCN channels. These results provide a structural framework for understanding the regulation of ion channels in the KCNH family by the C-linker/CNBHD and may guide the design of specific drugs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Sanguinetti, M. C. & Tristani-Firouzi, M. hERG potassium channels and cardiac arrhythmia. Nature 440, 463–469 (2006)
Zhang, X. et al. Deletion of the potassium channel Kv12.2 causes hippocampal hyperexcitability and epilepsy. Nature Neurosci. 13, 1056–1058 (2010)
Camacho, J. Ether à go-go potassium channels and cancer. Cancer Lett. 233, 1–9 (2006)
Ganetzky, B., Robertson, G. A., Wilson, G. F., Trudeau, M. C. & Titus, S. A. The Eag family of K+ channels in Drosophila and mammals. Ann. NY Acad. Sci. 868, 356–369 (1999)
Al-Owais, M., Bracey, K. & Wray, D. Role of intracellular domains in the function of the herg potassium channel. Eur. Biophys. J. 38, 569–576 (2009)
Stevens, L., Ju, M. & Wray, D. Roles of surface residues of intracellular domains of heag potassium channels. Eur. Biophys. J. 38, 523–532 (2009)
Gustina, A. S. & Trudeau, M. C. hERG potassium channel gating is mediated by N- and C-terminal region interactions. J. Gen. Physiol. 137, 315–325 (2011)
Muskett, F. W. et al. Mechanistic insight into human ether-à-go-go-related gene (hERG) K+ channel deactivation gating from the solution structure of the EAG domain. J. Biol. Chem. 286, 6184–6191 (2011)
Zhou, Z., Gong, Q., Epstein, M. L. & January, C. T. HERG channel dysfunction in human long QT syndrome. Intracellular transport and functional defects. J. Biol. Chem. 273, 21061–21066 (1998)
Zagotta, W. N. et al. Structural basis for modulation and agonist specificity of HCN pacemaker channels. Nature 425, 200–205 (2003)
Brelidze, T. I., Carlson, A. E. & Zagotta, W. N. Absence of direct cyclic nucleotide modulation of mEAG1 and hERG1 channels revealed with fluorescence and electrophysiological methods. J. Biol. Chem. 284, 27989–27997 (2009)
Morais Cabral, J. H. et al. Crystal structure and functional analysis of the HERG potassium channel N terminus: a eukaryotic PAS domain. Cell 95, 649–655 (1998)
Schonherr, R. & Heinemann, S. H. Molecular determinants for activation and inactivation of HERG, a human inward rectifier potassium channel. J. Physiol. (Lond.) 493, 635–642 (1996)
Wang, J., Trudeau, M. C., Zappia, A. M. & Robertson, G. A. Regulation of deactivation by an amino terminal domain in human ether-à-go-go-related gene potassium channels. J. Gen. Physiol. 112, 637–647 (1998)
Muskett, F. W. et al. Mechanistic insight into hERG K+ channel deactivation gating from the solution structure of the EAG domain. J. Biol. Chem. 286, 6184–6191 (2011)
Li, Q. et al. NMR solution structure of the N-terminal domain of hERG and its interaction with the S4–S5 linker. Biochem. Biophys. Res. Commun. 403, 126–132 (2010)
Ng, C. A. et al. The N-terminal tail of hERG contains an amphipathic α-helix that regulates channel deactivation. PLoS ONE 6, e16191 (2011)
Craven, K. B. & Zagotta, W. N. CNG and HCN channels: two peas, one pod. Annu. Rev. Physiol. 68, 375–401 (2006)
Brelidze, T. I., Carlson, A. E., Davies, D. R., Stewart, L. J. & Zagotta, W. N. Identifying regulators for EAG1 channels with a novel electrophysiology and tryptophan fluorescence based screen. PLoS ONE 5, e12523 (2010)
Splawski, I. et al. Spectrum of mutations in long-QT syndrome genes. KVLQT1, HERG, SCN5A, KCNE1, and KCNE2 . Circulation 102, 1178–1185 (2000)
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006)
Becchetti, A. et al. The functional properties of the human ether-à-go-go-like (HELK2) K+ channel. Eur. J. Neurosci. 16, 415–428 (2002)
Engeland, B., Neu, A., Ludwig, J., Roeper, J. & Pongs, O. Cloning and functional expression of rat ether-à-go-go-like K+ channel genes. J. Physiol. (Lond.) 513, 647–654 (1998)
Trudeau, M. C., Titus, S. A., Branchaw, J. L., Ganetzky, B. & Robertson, G. A. Functional analysis of a mouse brain Elk-type K+ channel. J. Neurosci. 19, 2906–2918 (1999)
Zou, A. et al. Distribution and functional properties of human KCNH8 (Elk1) potassium channels. Am. J. Physiol. Cell Physiol. 285, C1356–C1366 (2003)
Rehmann, H., Wittinghofer, A. & Bos, J. L. Capturing cyclic nucleotides in action: snapshots from crystallographic studies. Nature Rev. Mol. Cell Biol. 8, 63–73 (2007)
Altieri, S. L. et al. Structural and energetic analysis of activation by a cyclic nucleotide binding domain. J. Mol. Biol. 381, 655–669 (2008)
Clayton, G. M., Silverman, W. R., Heginbotham, L. & Morais-Cabral, J. H. Structural basis of ligand activation in a cyclic nucleotide regulated potassium channel. Cell 119, 615–627 (2004)
Schunke, S., Stoldt, M., Lecher, J., Kaupp, U. B. & Willbold, D. Structural insights into conformational changes of a cyclic nucleotide-binding domain in solution from Mesorhizobium loti K1 channel. Proc. Natl Acad. Sci. USA 108, 6121–6126 (2011)
Napolitano, C. et al. Genetic testing in the long QT syndrome: development and validation of an efficient approach to genotyping in clinical practice. J. Am. Med. Assoc. 294, 2975–2980 (2005)
Guerrero, S. A., Hecht, H. J., Hofmann, B., Biebl, H. & Singh, M. Production of selenomethionine-labelled proteins using simplified culture conditions and generally applicable host/vector systems. Appl. Microbiol. Biotechnol. 56, 718–723 (2001)
Collaborative Computation Project 4 The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Otwinowski, Z. & Minor, W. Processing X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
DeLano, W. L. The PyMOL molecular graphics system. http://www.pymol.org (DeLano Scientific, 2002)
Huson, D. H. et al. Dendroscope: an interactive viewer for large phylogenetic trees. BMC Bioinformatics 8, 460 (2007)
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl Acad. Sci. USA 98, 10037–10041 (2001)
Acknowledgements
We thank M. Munari, S. Camp, S. Cunnington and G. Sheridan for excellent technical assistance. We thank the beamline staff at the Advanced Light Source (ALS) and especially P. Zwart for help with data analysis. We also thank the members of the Zagotta laboratory for helpful discussions. This work was supported by the Howard Hughes Medical Institute, National Institutes of Health (NIH) grant R01 EY010329 (W.N.Z.) and NIH grant F32 HL095241 (A.E.C.). The Berkeley Center for Structural Biology is supported in part by the NIH, National Institute of General Medical Sciences and the Howard Hughes Medical Institute. The ALS is supported by the Director, Office of Science, Office of Basic Energy Sciences, of the US Department of Energy under contract no. DE-AC02-05CH11231.
Author information
Authors and Affiliations
Contributions
T.I.B. and W.N.Z. conceived the experiments. T.I.B. performed the crystallographic experiments, and B.S. helped with the crystallographic data analysis. A.E.C. and W.N.Z. performed the electrophysiology experiments and data analysis. T.I.B. and W.N.Z. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-10 with legends, Supplementary Table 1, a Supplementary Discussion and additional references. This file was replaced on 18 April 2012 (PDF 6933 kb)
Rights and permissions
About this article
Cite this article
Brelidze, T., Carlson, A., Sankaran, B. et al. Structure of the carboxy-terminal region of a KCNH channel. Nature 481, 530–533 (2012). https://doi.org/10.1038/nature10735
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10735
This article is cited by
-
Diversity of voltage-gated potassium channels and cyclic nucleotide-binding domain-containing channels in eukaryotes
Scientific Reports (2020)
-
Identification of undecylenic acid as EAG channel inhibitor using surface plasmon resonance-based screen of KCNH channels
BMC Pharmacology and Toxicology (2019)
-
Eag1 Voltage-Dependent Potassium Channels: Structure, Electrophysiological Characteristics, and Function in Cancer
The Journal of Membrane Biology (2017)
-
Structure of the Cyclic Nucleotide-Binding Homology Domain of the hERG Channel and Its Insight into Type 2 Long QT Syndrome
Scientific Reports (2016)
-
Human EAG channels are directly modulated by PIP2 as revealed by electrophysiological and optical interference investigations
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.