Abstract
Variable regions 1 and 2 (V1/V2) of human immunodeficiency virus-1 (HIV-1) gp120 envelope glycoprotein are critical for viral evasion of antibody neutralization, and are themselves protected by extraordinary sequence diversity and N-linked glycosylation. Human antibodies such as PG9 nonetheless engage V1/V2 and neutralize 80% of HIV-1 isolates. Here we report the structure of V1/V2 in complex with PG9. V1/V2 forms a four-stranded β-sheet domain, in which sequence diversity and glycosylation are largely segregated to strand-connecting loops. PG9 recognition involves electrostatic, sequence-independent and glycan interactions: the latter account for over half the interactive surface but are of sufficiently weak affinity to avoid autoreactivity. The structures of V1/V2-directed antibodies CH04 and PGT145 indicate that they share a common mode of glycan penetration by extended anionic loops. In addition to structurally defining V1/V2, the results thus identify a paradigm of antibody recognition for highly glycosylated antigens, which—with PG9—involves a site of vulnerability comprising just two glycans and a strand.
Similar content being viewed by others
Main
As the sole viral target of neutralizing antibodies, the HIV-1 viral spike has evolved to evade antibody-mediated neutralization (reviewed in ref. 1). V1/V2 of the gp120 component of the viral spike is critical to this evasion. Localized by electron microscopy to a membrane-distal ‘cap’2,3,4,5, which holds the spike in a neutralization-resistant conformation, V1/V2 is not essential for entry: its removal, however, renders the virus profoundly sensitive to antibody-mediated neutralization6,7,8,9.
The ∼50–90 residues that comprise V1/V2 contain two of the most variable portions of the virus, and roughly 1 in 10 residues of V1/V2 are N-glycosylated. Despite the diversity and glycosylation of V1/V2, a number of broadly neutralizing human antibodies have been identified that target this region, including the somatically related antibodies PG9 and PG16, which neutralize 70–80% of circulating HIV-1 isolates10, antibodies CH01–CH04, which neutralize 40–50%11, and antibodies PGT141–145, which neutralize 40–80%12. These antibodies all share specificity for an N-linked glycan at residue 160 in V1/V2 (HXB2 numbering) and show a preferential binding to the assembled viral spike over monomeric gp120 as well as a sensitivity to changes in V1/V2 and some V3 residues. Sera with these characteristics have been identified in a number of HIV-1 donor cohorts, and these quaternary-structure-preferring V1/V2-directed antibodies are among the most common broadly neutralizing responses in infected donors13,14.
Despite extensive effort, V1/V2 had resisted atomic-level characterization. Here we report crystal structures of the V1/V2 domain of HIV-1 gp120 from clade C strains CAP45 and ZM109 in complexes with the antigen-binding fragment (Fab) of PG9 at 2.19- and 1.80-Å resolution, respectively. We elucidate how the V1/V2 fold accommodates sequence variation and glycosylation, provide an atomic-level description of the PG9 epitope, and analyse other members of this V1/V2-directed class of broadly neutralizing antibodies to identify conserved features that enable recognition of this key glycopeptide target.
Structure determination
Variational crystallization15 of HIV-1 gp120 with V1/V2 was attempted following strategies that were successful for structural determination of other portions of HIV-1 gp120 (refs 15–17); this failed to produce V1/V2-containing crystals suitable for structural analysis (Supplementary Table 1). Because V1/V2 emanates from similar hairpins in core structures of HIV-1 (refs 18–21) and SIV22 (Supplementary Fig. 1), we hypothesized that a protein scaffold that provided an appropriate hairpin might suitably incorporate and express an ectopic V1/V2 region. We identified six proteins with potentially suitable acceptor β-hairpins that ranged in size from 135 to 741 amino acids. Only the smallest of these could be expressed in transfected 293F cells when scaffolded with V1/V2 (Supplementary Table 2), but it behaved poorly in solution. We identified 11 smaller proteins of 36–87 amino acids in size and designed chimaeric proteins encoding V1/V2 from the YU2 strain of HIV-1 (Supplementary Fig. 2 and Supplementary Table 3). The expressed chimaeric glycoproteins from these smaller scaffolds were mostly soluble, permitting us to characterize them antigenically against a panel of six YU2-specific V1/V2 antibodies (Supplementary Tables 4 and 5). Three of the smaller scaffolded YU2 V1/V2 chimaeras showed reactivity with all six YU2-specific antibodies, and two (Protein Data Bank (PDB) accessions 1FD6 (ref. 23) and 1JO8 (ref. 24)) were also recognized by the α4β7 integrin25, suggesting that they retained biological integrity (Supplementary Table 5 and Supplementary Fig. 3). We next identified strains of gp120 that retained PG9 recognition in the gp120 monomer context, including clade B strain TRJO and clade C strains 16055, CAP45, ZM53 and ZM109 (Supplementary Table 6). We placed V1/V2 sequences (residues 126–196) from these strains into the 1FD6 and 1JO8 scaffolds, and assessed PG9 binding. Notably, affinities of PG9 for 1FD6-ZM109 and 1JO8-ZM109 were only 50-fold and threefold lower than wild-type ZM109 gp120, respectively (Supplementary Fig. 4). Scaffold-V1/V2 heterogeneity was apparent after expression in GnTI−/− cells26 as was sulphation heterogeneity on antibody PG9 (ref. 27) (Supplementary Fig. 5). We therefore used an on-column selection procedure coupled to on-column protease cleavage of Fab to obtain homogeneous complexes of scaffold-V1/V2 with PG9 (Supplementary Fig. 6).
Two 1FD6-V1/V2 scaffolds were crystallized in complex with PG9. One scaffold contained the V1/V2 region from the gp120 of the CAP45 strain of HIV-1 with five sites of potential N-linked glycosylation. Crystals of this CAP45 construct with the Fab of PG9 diffracted to 2.19 Å, and the structure was refined to an Rcryst of 18.2% (Rfree = 23.4%) (Fig. 1 and Supplementary Table 7). A second scaffold included the V1/V2 region from the gp120 of the ZM109 strain of HIV-1 with N-linked glycans at positions 160 and 173, and asparagine to alanine substitutions at four other potential N-linked sites. Crystals of this ZM109 construct with the Fab of PG9 diffracted to 1.80 Å, and the structure was refined to an Rcryst of 17.8% (Rfree = 20.5%) (Supplementary Fig. 7 and Supplementary Table 7).
V1/V2 from the CAP45 strain of HIV-1 is shown, in magenta ribbons, in complex with Fab of antibody PG9. The PG9 heavy and light chains are shown as yellow and blue ribbons, respectively, with CDRs in different shades. Although the rest of HIV-1 gp120 has been replaced by the 1FD6 scaffold (shown in white ribbons), the positions of V1/V2, PG9 and the scaffold are consistent with the proposal that the viral spike, and hence the viral membrane, is positioned towards the top of the page. The extended CDR H3 of PG9 is able to penetrate the glycan shield that covers the V1/V2 cap on the spike and to reach conserved elements of polypeptide, while residues in heavy- and light-chain-combining regions recognize N-linked glycans. The disordered region of the V2 loop is represented by a dashed line. Perpendicular views of V1/V2 are shown in Figs 2 and 6, and the structure of PG9 in complex with V1/V2 from HIV-1 strain ZM109 is shown in Supplementary Fig. 7.
Structure of V1/V2
The V1/V2 structure, in the context of scaffold and PG9, folds as four anti-parallel β-strands (labelled A, B, C, D) arranged in (−1, −1, +3) topology28 (Fig. 2a–d and Supplementary Table 8). Important structural elements, such as a hydrophobic core, connecting loops and disulphide bonds, cross between each of the four strands, indicating that, structurally, the V1/V2 domain should be considered a single topological entity.
a–i, The four anti-parallel strands that define V1/V2 fold as a single domain, in a topology known as ‘Greek key’ that is observed in many proteins. a, Schematic of V1/V2 topology. V1/V2 resides between strands β2 and β3 of core gp120, and its structure completes the crystallographic determination of all portions of HIV-1 gp120. Strands are depicted as arrows and disulphide bonds as yellow lines. b, c, Ribbon diagram of V1/V2 residues 126–196 from HIV-1 strains CAP45 (magenta) and ZM109 (green). Conserved disulphide bonds are represented as ball and stick, and the beginning and terminating residues of each strand are labelled. d, Superposition of the structures shown in b and c. e, Amino acid conservation of V1/V2. The backbone is shown as a tube of variable thickness, coloured as a rainbow from cold (blue) to hot (red), corresponding to conserved (thin) and to variable (thick), respectively, based on an alignment of 166 HIV-1 sequences. Aliphatic and aromatic side chains are shown as sticks with semi-transparent molecular surface, coloured by conservation as in i. f, Electrostatic surface potentials of CAP45 V1/V2 coloured blue to red, corresponding to positive and negative surface potentials (+4 to −4 kT/e), respectively. g, Molecular surfaces corresponding to main-chain atoms including Cβ are coloured orange, with other surfaces coloured white. h, Superposition of ZM109 and CAP45 models containing V1 and V2 loops and associated glycans. For each glycosylated asparagine, only the first N-acetylglucosamine attached to the asparagine is shown and represented as sticks. Modelled amino acids and glycans that are disordered in the crystal structures are shown in grey. i, Sequence alignment of nine HIV-1 strains that are potently neutralized by PG9. Glycosylated asparagine residues are boxed and in bold. Identical residues have a dark green background with white characters, whereas conserved residues have white backgrounds with dark green characters. Above the alignment, β-strands are shown as arrows, coloured magenta and green for CAP45 and ZM109, respectively. Residues (including attached glycans) that make hydrogen bonds to PG9 are denoted with symbols above the alignment (side-chain hydrogen bonds are indicated by open circles with dashes, main-chain hydrogen bonds are indicated by closed circles, or main-chain and side-chain hydrogen bonds by closed circles with dashes).
Overall, the four-stranded V1/V2 sheet presents an elegant solution for maintaining a common fold while accommodating V1/V2 diversity and glycosylation. The strands contain mostly conserved residues and are welded in place by inter-strand disulphide bonds (between strand A and neighbouring strands B and D) and extensive hydrogen bonding (between strands A and D and between strands B and C). The two faces of the sheet—concave and convex—have very different characters. The concave face of the sheet is glycan-free and hydrophobic (Fig. 2e), with a cluster of aliphatic and aromatic side chains surrounding the disulphide bond that links strands A and B. This conserved hydrophobic cluster continues onto strand D at the sheet edge, to form a half-exposed hydrophobic core for this domain. The convex face of the sheet is cationic (Fig. 2f), with the main-chain atoms of the conserved strands of the sheet forming stripes on the V1/V2 surface (Fig. 2g) and the N-linked glycan 160 situated at its centre (Fig. 2h). In contrast, two strand-connecting loops—emanating from the same end of the sheet—are highly glycosylated and variable in sequence (Fig. 2i). We can now redefine the ‘V1 loop’ as the residues connecting strands A and B, and the ‘V2 loop’ as those residues between strands C and D (Fig. 2h, i). Of these, the V1 loop is most variable, ranging in length from ∼10–30 residues. The V2 loop is less variable and contains at its start the tripeptide motif recognized by integrin α4β7, the gut homing receptor for HIV-1 (ref. 25).
PG9–V1/V2 interactions
The most prominent interaction between antibody PG9 and V1/V2 occurs with N-linked glycan (Fig. 3, Supplementary Fig. 8 and Supplementary Tables 9 and 10). PG9 grasps the entire ZM109 160 glycan (Fig. 3a). Its protruding third complementarity-determining region of the heavy chain (CDR H3) reaches through the glycan shield to contact the protein-proximal N-acetylglucosamine, burying 200 Å2 of total surface area, with Asp 100 and Arg 100B of PG9 making four hydrogen bonds (Fig. 3b, c) (Kabat numbering29 is used in description of antibody sequences; thus Arg 100B is two residues after Asp 100). Additional hydrogen bonds are made by the base of the CDR H3 (by Asn 100P and by the double-mannose-interacting His 100R) to terminal mannose residues, with Ser 32 and Asp 50 of the light chain contributing three additional hydrogen bonds (Fig. 3b). In sum, a total of 11 hydrogen bonds and over 1,150 Å2 of surface area are buried in the PG9–glycan-160 interface (489 Å2 on PG9 and 670 Å2 on glycan 160), with PG9 contacting 5 of the 7 saccharide moieties of the Man5GlcNAc2 glycan (Fig. 3c). Similar extensive interactions are observed with residue 160 of CAP45 (Supplementary Fig. 8a–c). The preference of PG9 for a Man5GlcNAc2 glycan at residue 160 is now clear: a larger glycan would clash with the antibody light chain and a shorter glycan would not stretch between tip and base of the PG9 CDR H3.
a–f, Glycan, electrostatic and sequence-independent interactions of antibody PG9 facilitate recognition of V1/V2 from the ZM109 strain of HIV-1 gp120. a, PG9 is shown as a grey molecular surface, and strands B and C of V1/V2 are shown as green ribbons. Mannose and N-acetylglucosamine residues are shown in stick representation, as are the side chains of Asn 160 and 173. Electron density (2Fo − Fc) is contoured at 1σ and shown as a blue mesh. b, Ribbon representations of strands B and C of ZM109 V1/V2 (green), PG9 heavy chain (yellow) and PG9 light chain (blue). V1/V2 glycans and PG9 residues that hydrogen bond are shown as sticks. Nitrogen atoms are coloured blue, oxygen atoms are coloured red, and dotted lines represent hydrogen bonds. c, Schematic of the Man5GlcNAc2 moiety attached to Asn 160. GlcNAcs are shown as blue squares, and mannoses as green circles. Hydrogen bonds to PG9 are listed to the right of the symbols, as is the total surface area buried at the interface between PG9 and each sugar. H-bonds, hydrogen bonds. d, Schematic of the PG9–main-chain interaction with V1/V2. Disulphide bonds in V1/V2 are shown as yellow sticks. e, f, Ribbon representation of V1/V2 (green) and PG9 CDR H3 (yellow). Hydrogen bonds are represented by dotted lines. Main-chain interactions are shown in e, and side chain interactions in f (with the two images related by a 90° rotation about a vertical axis). Details of the PG9 interaction with V1/V2 from the CAP45 strain of HIV-1 are shown in Supplementary Fig. 8.
Interactions also occur between PG9 and the N-linked glycan at residue 156 (CAP45) or residue 173 (ZM109). With CAP45, much of the 156 glycan is ordered, stabilizing six of the seven sugars, including four of the five mannose residues (Supplementary Fig. 8). Hydrogen bonds are observed between the 156 glycan and the side chains of Asn 73 and Tyr 100K of the PG9 heavy chain, and 766 Å2 of total buried surface area (337 Å2 on PG9 and 429 Å2 of glycan). Glycan 156 is not preserved in the ZM109 sequence, in which residue 156 is a histidine (Fig. 2i); an additional site of N-linked glycosylation, however, occurs in ZM109 at residue 173, in the middle of strand C. In the ZM109 structure, glycan 173 is in virtually the same spatial location as glycan 156 in the CAP45 structure (Fig. 2h). PG9 binds to the protein-proximal N-acetylglucosamine, with Tyr 100K making a hydrogen bond and a total of 189 Å2 surface area buried (Fig. 3b). Notably, substitutions that result in changes in V1/V2 glycans indicate that glycan at 160 is critical for PG9 recognition (Supplementary Table 11), and 156/173 is important (although PG9 recognizes strains of HIV-1 lacking a 156/173 glycan; Supplementary Fig. 9). Many of the changes in the heavy and light chains that allow for glycan recognition occur during affinity maturation (Supplementary Tables 12 and 13), providing a possible explanation for the observed increase in PG9 (and PG16) breadth and affinity during affinity maturation30.
In addition to glycan recognition, a strand in the CDR H3 of PG9 forms intermolecular parallel β-sheet-like hydrogen bonds to strand C of V1/V2 (Fig. 3d, e). Strand C is the most variable of the V1/V2 strands, and this sequence-independent means of recognition probably allows for increased recognition breadth. Specific electrostatic interactions are also made between cationic residues of strand C and acidic residues on PG9. Notably, several of these occur with sulphated tyrosines on CDR H3. Because parallel β-strand hydrogen bonding would tend to align main-chain atoms of CDR H3 and strand C, the charged tips of Lys and Arg residues would protrude beyond the standard acidic Asp and Glu side chains, whereas tyrosine sulphates provide a closer match to the side-chain length of basic Lys/Arg residues.
Overall, the structure of PG9 is consistent with published mutational data10,14 (Supplementary Table 14). Some residues such as Phe 176 are critical because they form part of the hydrophobic core on the concave face of the V1/V2 sheet. Others form direct contacts: for example, the tyrosine sulphate at residue 100H of PG9 interacts with residue 168 when it is an Arg (strain ZM109) or Lys (strain CAP45), but would be repelled by a Glu (as in strain JR-FL); JR-FL is resistant to neutralization by PG9, but becomes sensitive if Glu 168 is changed to Lys10.
Quaternary preferences of PG9 and PG16
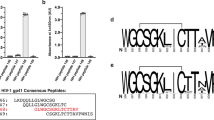
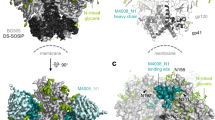
PG9 and the somatically related PG16 typically recognize the assembled viral spike with higher affinity than monomeric gp120 (ref. 10). For PG9, the average monomeric gp120 affinity, as assessed by ELISA or surface plasmon resonance, was at least tenfold weaker than viral spike affinity, as assessed by neutralization; with PG16, the difference was at least 100-fold (Fig. 4a, Supplementary Tables 6 and 15–17). Such differences are probably greater as the concentration required for neutralization (half-maximum inhibitory concentration, IC50) is often higher than the affinity (half-maximum effective concentration, EC50, or equilibrium dissociation constant, Kd). To investigate differences between monomeric and oligomeric contexts, we acquired negatively stained electron microscopy images of PG9 in complex with monomeric gp120 (Fig. 4b, Supplementary Figs 10 and 11). To define the orientation of monomeric gp120, we used the CD4-binding-site-directed antibody T13, for which we defined the crystal structure of gp120-bound T13 Fab at 6-Å resolution (Supplementary Figs 12, 13 and Supplementary Table 18). This structure along with the V1/V2–PG9 structure allowed for the definition of six classes of relative gp120–PG9 orientations, indicating that the position of V1/V2 varies in the monomeric gp120 context. In contrast, prior cryoelectron microscopy results indicate that the position of V1/V2 in the unliganded Env trimer spike is fixed2,3,4,5.
a–c, Quaternary-structure-preferring antibodies show different affinities for oligomeric, monomeric and scaffolded V1/V2. Both structural and arginine-scanning mapping, however, suggest that the epitopes of PG9 and PG16 are mostly present in scaffolded V1/V2. a, Affinities of PG9 (filled symbols) and PG16 (open symbols) are shown for the functional viral spike (gp120/gp41)3 (circles), monomeric gp120 (triangles), and scaffold-V1/V2 (squares), based on neutralization (black), ELISA (blue) and surface plasmon resonance (red). b, Negative stained class averages are shown for ternary complexes of wild-type gp120 (HIV-1 strain 16055) in complex with antibody PG9 and the CD4-binding-site antibody T13. Six different classifications were observed, and are superimposed in the left panel and labelled PG9-1 to PG9-6. Individual fitting for classes PG9-1, PG9-3 and PG9-5 are shown after rigid-body alignment of Fab PG9–scaffold-V1/V2, Fab T13 and core gp120 (in the conformation bound by the CD4-binding site antibody F105; ref. 20). c, Comparison of crystallographically defined PG9 paratope with neutralization-defined PG16 paratope. Scaffold-V1/V2 interactive surface of PG9 in ZM109 (left) and CAP45 (middle) contexts is shown along with the PG16 paratope (right) as defined by arginine-scanning mutagenesis (residues mutated to arginine are coloured according to the fold-increase in IC50 for the mutant relative to wild type: yellow for <50, orange for <100 and red for >100; orange-highlighted residue is Trp 64 in the CDR H2). Perpendicular views of the paratope, rotated by 90° about a horizontal axis, are shown in top and bottom rows.
Additionally, we chose to map the antibody paratope by assessing neutralization with arginine mutants. We selected the PG16 paratope for characterization, as its recognition seemed to be both more quaternary-structure preferring (Fig. 4a) and more V3 dependent10 than that of PG9. We parsed its combining site into 21 surface segments plus one in the framework as a control. Each of these was altered by the introduction of a single arginine mutation, expressed as an immunoglobulin, and assessed for neutralization on a panel of diverse HIV-1 isolates (Supplementary Fig. 14). The resultant ‘arginine-scanning’ mutagenesis revealed a close match to the observed V1/V2 interface for PG9 (Fig. 4c). We also measured the binding of PG9 and PG16 to monomeric gp120 in wild-type and V3-deleted contexts, and observed similar affinities, indicating that—in the context of monomeric gp120—V3 does not have a substantial role in PG9 or PG16 recognition (Supplementary Fig. 15). Lastly, accumulating data indicate that V1/V2 in the viral spike both shields and interacts with V3 (refs 6–9).
Collectively, these results suggest that the V1/V2–PG9 interaction observed in the scaffolded V1/V2–PG9 crystal structures encompasses much of the PG9/PG16 epitope, and that the structural integrity of this epitope is sensitive to appropriate assembly of the viral spike. We note parenthetically that the ability of the PG9/PG16-recognized epitope to be preferentially present in the assembled viral spike provides a useful strategy to hide this potential site of vulnerability. That is, the site may be preferentially present on the assembled viral spike, but not on shed or other monomeric forms of gp120, which are thought to be the predominant form of Env in infected individuals, and we note in this regard that many V1/V2-directed antibodies are substantially more quaternary-structure preferring than PG9 (refs 10–12, 31). The quaternary-specific nature of the epitope may thus reflect a functional adaptation of HIV-1.
A conserved CDR H3 motif for V1/V2 recognition
Sequences of other V1/V2-directed broadly neutralizing antibodies indicate the presence of long CDR H3s, but little other sequence conservation (Fig. 5a). We have not yet been able to determine structures of other class members in complex with V1/V2, but nonetheless sought to provide insight into their conserved features of recognition by analysing unbound Fab structures.
a, b, A protruding anionic CDR H3 is preserved in members of this broadly neutralizing class of antibodies. a, CDR H3 sequence alignment. Cohort, donor information and sequences in the CDR H3 are shown for V1/V2-directed antibodies (note that Kabat29 numbering and formatting is shown for PG9, to which other antibodies are aligned). Positively charged residues are boxed in blue and negatively charged residues in red. Residues that make hydrogen bonds to CAP45 residues (magenta) or glycans (purple) are denoted with symbols above the alignment (side-chain hydrogen bonds are indicated by open circles with dashes, main-chain hydrogen bonds by closed circles, or main-chain and side-chain hydrogen bonds by closed circles with dashes). Similar contacts are shown for ZM109 residues (green) or glycans (grey). Sulphated tyrosines are circled or squared if the post-translational modification has been confirmed crystallographically or by mass spectrometry, respectively. The sequence for the V1/V2-directed strain-specific antibody, 2909, is also included. b, Protruding CDR H3, displayed as ribbon diagrams with sulphated tyrosines shown in spheres and paired with electrostatic surface potentials coloured blue to red, corresponding to positive and negative surface potentials, respectively. All CDR H3s are aligned so that the light chain would be on the left and the heavy chain on the right (as in Supplementary Fig. 17). Average electrostatic potentials are shown for specified CDR H3 and V1V2 surfaces.
The structure of unbound PG9 Fab (3.3-Å resolution, four molecules per asymmetric unit; Supplementary Fig. 16 and Supplementary Table 19) revealed considerable CDR H3 flexibility, similar to that observed previously with PG16 (ref. 30). For CH01–CH04 antibodies11, crystallization was attempted for antigen-binding fragments and for six heavy/light-chain somatic chimaeras (Supplementary Table 20). Structures were determined for CH04 Fab and also for the CH04H/CH02L Fab, the latter in two different crystal forms (Supplementary Fig. 17 and Supplementary Table 19). These structures revealed an anionic CDR H3 for CH04, which extended above the rest of the combining site in a manner similar to the CDR H3s of PG9 and PG16 (Fig. 5b). With CH04, however, the extended hairpin was twisted ∼90°, to an orientation that bisected heavy and light chains. The spacing between the protruding CDR H3 and the rest of the combining region was reduced by 8 Å relative to that of PG9, and no CDR H3 tyrosine sulphation was observed.
With PGT141–145 antibodies12, Fab crystals of unbound PGT145 diffracted to 2.3 Å and revealed an extended, tyrosine-sulphated CDR H3 loop, which, like those of PG9, PG16 and CH04, reached substantially beyond the rest of the CDR loops. In contrast, the β-hairpin of CDR H3 extended vertically (parallel to the long axis of the Fab) (Fig. 5b, Supplementary Fig. 18 and Supplementary Table 19) and was rigidified by extensive tyrosine stacking (along with the standard strand–strand hydrogen bonding). Its negatively charged tip (including two sulphated tyrosines) was followed by a Gly-containing potential ‘hinge’ and resembled an extended version of the CDR H3 of antibody 2909 (refs 32, 33), a highly quaternary-structure-sensitive antibody31,34 that recognizes an immunotype variant of the V1/V2 target site in which a Lys is substituted for the N-linked glycan at position 160 (ref. 35).
Thus, despite having been derived from three different individuals, antibodies from this class of V1/V2-directed broadly neutralizing antibodies all showed anionic protruding CDR H3s (Fig. 5b), most of which were tyrosine sulphated. All also displayed β-hairpins and, although these varied substantially in orientation relative to the rest of the combining site, all appeared capable of penetrating an N-linked glycan shield to reach a cationic protein surface.
A V1/V2 site of HIV-1 vulnerability
With both CAP45 and ZM109 strains of gp120, the V1/V2 site recognized by PG9 consists primarily of two glycans and a strand (Fig. 6a). Minor interaction with strand B and with the B–C connecting loop (3% and 3–5% of the total interactive surface, respectively) complete the epitope, with the entire PG9-recognized surface of V1/V2 contained within the B–C hairpin (Supplementary Table 21). The minimal nature of this epitope suggests that it might be easier to engineer and to present to the immune system than other, more complex, epitopes. The epitopes for antibodies b12 and VRC01, for example, are comprised of seven and six independent protein segments19,36, respectively. The presence in the PG9 epitope of N-linked glycosylation, which is added by host cell machinery, does provide a potentially complicating factor for humoral recognition.
a, b, Glycan, electrostatic and sequence-independent interactions allow PG9 to recognize a glycopeptide site on V1/V2. a, Site characteristics in CAP45 strain of HIV-1. Glycans 160 and 156 (173 with ZM109) are highlighted in green, and strands B and C are highlighted in magenta, with the rest of V1/V2 in semi-transparent white. The interactive surface of V1/V2 with PG9 is shown, coloured according to the local electrostatic potential as in Fig. 5b. The contribution of each structural element to that surface is provided as a percentage of the total. Although the V1/V2 scaffolds used here do not allow a comprehensive analysis of the overall antibody response to this region of gp120, in addition to assisting with structural definition of effective V1/V2-directed neutralization, the V1/V2 scaffolds may have utility in attempts to direct the V1/V2-elicited response away from the hypervariable loops to the conserved strands—especially the site of vulnerability highlighted here. b, Saturation transfer difference (STD) NMR for Man5GlcNAc2-Asn binding to PG9. Top, STD spectrum of 1.5 mM Man5GlcNAc2-Asn in the presence of 15 µM Fab PG9 (lower spectrum) is paired with the corresponding reference spectrum (upper spectrum). Bottom left, Langmuir binding curve as a function of glycan concentration, used to obtain the Kd (ASTD signals correspond to N-acetyl protons, which are shown in the boxed area of the top panel). Bottom right, stacked STD NMR spectra as a function of Man5GlcNAc2-Asn concentration.
To assess glycan affinities, we used saturation transfer difference NMR. In the crystal structure, recognition by PG9 occurs with protein-proximal N-acetylglucosamine and terminal mannose saccharides. With 1.5 mM (N-acetylglucosamine)2, interaction with PG9 was not observed (Supplementary Fig. 19), whereas with 1.5 mM oligomannose-5, weak interactions were observed (Supplementary Fig. 20). We further carried out a titration series with Asn-(N-acetylglucosamine)2(mannose)5 and determined its affinity for PG9 to be 1.6 ± 0.9 mM (Fig. 6b). The weak affinity for glycan (surprising in the face of such a large contact surface and multiple hydrogen bonds) provides a potential explanation for the reported lack of PG9 autoreactivity despite its N-glycan dependence10 (specificity for oligomannose-5 probably also reduces PG9 autoreactivity, as this glycan is infrequently displayed on the surface of mammalian cells).
Strand C is the most cationic of the V1/V2 strands. This conserved cationic character—present in the target-cell-facing V1/V2 cap of the viral spike—may relate to the observed anionic interactions of the viral spike, both with dextran sulphate37,38 and other polyanions39,40 or with heparan sulphate on the cell surface41. In terms of the ionic interactions of PG9 itself, we observed sulphation to increase affinity and neutralization potency by ∼10-fold27 (Supplementary Fig. 5). Ionic PG9 interactions may thus mimic the functional polyanion–V1/V2 interactions that HIV-1 uses for cell surface attachment during the initial stages of virus cell entry.
Strand C is also the most variable of the V1/V2 strands. Its location, at the edge of the sheet, however, provides an opportunity for sequence-independent recognition, through its exposed main-chain atoms. Whereas the four hydrogen bonds made by the main chain of PG9 probably contribute only a small portion of the overall binding energy, the surface of V1/V2 involved in main-chain interactions with PG9 totals 348 and 350 Å2 in the CAP45 and ZM109 complexes, respectively, potentially providing a substantial contribution to the overall binding energy (Supplementary Table 21). This type of β-sheet interaction, for example, is the primary interaction between the CDR H3 of antibody 447-52D and the V3 of gp120 (ref. 42) in a 3-and-almost-4-stranded β-sheet.
Vaccine implications
The different types of PG9 interaction, involving glycan, electrostatic and sequence-independent interactions, all seem to be necessary for PG9 function. Such multicomponent recognition may also provide a mechanism that enables the immune system to overcome evasion associated with individual components of the interaction. Thus, for example, glycan-only affinity might lead to autoreactivity, and surface areas of electrostatic and sequence-independent interactions might be individually too small to generate sufficient affinity for tight interactions. Together, however, the glycan, electrostatic and sequence-independent interactions achieve the substantial level of affinity required for potent neutralization.
In longitudinal studies, antibody responses that require glycan, either at residue 160, as described here, or at residue 332, constitute the most commonly elicited initial broadly neutralizing responses43, an observation also seen with elite neutralizers13. In such studies, transmitted/founder viruses in some cases did not show canonical glycosylation (for example, at positions 160 or 332), but acquired these under immune selection44. Thus it seems that N-linked glycosylation at particular residues is selected as a means of immune evasion, but that these same glycans—now part of a homogeneous glycan array—can be recognized by very broadly neutralizing antibodies. Recent structural results indicate that a number of 332-glycan-dependent antibodies also use protruding CDR H3s and, in at least one case, the antibody (PGT128) recognizes an epitope composed of two glycans and a strand45. Collectively these results suggest that an antibody with a penetrating CDR H3 that recognizes a conserved glycan and neighbouring polypeptide is a paradigm for humoral recognition of heavily glycosylated antigens.
Lastly, we note that recent immune analyses of the RV144 vaccine trial suggest that the presence of V1/V2-reactive antibodies associates inversely with HIV-1-infection risk46,47. What role these antibodies have in protection, however, is unknown. It is nonetheless fascinating that the V1/V2 domain—which functions in evading antibody-mediated neutralization—is itself a site of effective neutralization by a number of broadly neutralizing antibodies (for example, by PG9, CH04 and PGT145) and that antibodies against this domain seem to be an indicator of a successful vaccine outcome.
Methods Summary
Proteins that could accommodate backbone grafting of the V1/V2 stub from HIV-1 gp120 were identified using the Multigraft Match algorithm48 implemented in Rosetta. Potential V1/V2 scaffolds were examined manually and, if necessary, optimizations were made to accommodate full-length V1/V2 loops (residues 126–196) or to alter scaffold properties (for example, mutating the intrinsic immunoglobulin affinity of 1FD6; ref. 49). For each V1/V2 scaffold, protein-A-purified PG9, altered to remove light-chain glycosylation and to introduce an HRV3C cleavage site in the hinge, was bound to Protein A Plus agarose, and the V1/V2 scaffold added. After washing away unbound scaffold, HRV3C protease was added to elute the PG9 Fab–V1/V2 scaffold complex. Complexes of PG9 Fab bound to 1FD6-CAP45 or 1FD6-ZM109 crystallized in similar conditions50 (8–17% (w/v) PEG 3350, 5–10% (v/v) 2-methyl-2,4-pentanediol, 0.2 M lithium sulphate, 0.1 M imidazole pH 6.5). Crystals were cryoprotected with 15% (v/v) 2R,3R-butanediol, diffraction data were collected to 2.19 and 1.80 Å for PG9–1FD6-CAP45 and PG9–1FD6-ZM109, respectively, and structures solved by molecular replacement.
Online Methods
Design of large V1/V2 scaffolds
Large V1/V2 scaffolds were identified by a search of a culled database of high-resolution crystal structures from the PDB, using the Multigraft Match algorithm implemented in Rosetta Multigraft48. Briefly, the stub of the V1/V2 region from gp120 (PDB code 1RZJ) was treated as an epitope, and an exhaustive search was conducted for scaffolds that could accommodate backbone grafting of the V1/V2 stub while maintaining backbone continuity and avoiding steric clash. Multiple combinations of endpoints on the V1/V2 stub were tested, including the following pairs of endpoints in 1RZJ: (124, 196), (125, 196), (126, 196), (124, 197), (125, 197), (126, 197), (124, 198), (125, 198), (126, 198). Matches were initially accepted with a loop closure r.m.s.d. of < 2.0 Å and a steric clash between the V1/V2 stub and the scaffold of less than 1.0 Rosetta units with all atoms present and having allowed for side-chain repacking. Only three scaffolds with >500 residues were identified with very low r.m.s.d. loop closure (<0.5 Å) for the V1/V2 stub. To obtain additional scaffolds, a list of high-resolution structures of large chains was constructed (346 chains included) and the V1/V2 stub was grafted at manually selected sites on all unique proteins in that list, using explicit flexible backbone loop closure in RosettaRemodel51. If RosettaRemodel could produce a grafted V1/V2 stub with a fully closed chain while maintaining hydrogen bonding in the remodelled region and without creating significant pockets in the structure, the output model was accepted as a scaffold candidate. The final scaffold sequences included the full-length YU2 V1/V2 sequence in place of the stub.
Design of small V1/V2 scaffolds
A database of small protein structures was created, with ligands removed and non-standard amino acids replaced by appropriate analogues. Candidate scaffolds were identified using the Multigraft Match algorithm as described above48. From the thousands of matches that passed these filters, the lowest r.m.s.d. match for each PDB code was examined manually to identify scaffolds with good packing, adequate tertiary structure supporting the V1/V2 stub, a minimum of buried unsatisfied polar residues, and adequate space to accommodate the large, glycosylated V1/V2 loops. In some cases, scaffolds were re-designed to improve these features using human-guided computational (fixed backbone) design. Once the scaffold design and grafting of the V1/V2 stub was completed, it was considered possible to insert any desired full-length V1/V2 sequence. This study initially used the YU2 V1/V2 sequence. A total of 11 scaffolds were designed in this manner, based on the following PDB entries: 1CHLA, 1FD6A, 1G6MA, 1IP9A, 1IW4A, 1JLZA, 1QPMA, 1XBDA, 1XQQA, 1YWJA and 1BRZ. An alanine mutation was made in the 1FD6 scaffold to remove intrinsic immunoglobulin affinity49. Two additional scaffolds were selected manually from crystal structures of small, stable proteins but were designed similarly using Multigraft Match; these scaffolds were based on PDB entries 1E6G and 1JO8.
Expression and purification of V1/V2 scaffolds
Mammalian codon-optimized genes encoding V1/V2 scaffolds were synthesized with an artificial N-terminal secretion signal and a C-terminal HRV3C recognition site followed by an 8×-His tag and a StreptagII. V1/V2 sequences were from HIV-1 strains TRJO, CAP45, ZM53, ZM109 or 16055. The genes were cloned into the XbaI/BamHI sites of the mammalian expression vector pVRC8400, and transiently transfected into HEK 293S GnTI−/− cells26, which were used due to a requirement for a Man5GlcNAc2 at position 160 by PG9 and other broadly neutralizing V1/V2-directed antibodies. Scaffolds were purified from the media using Ni2+-NTA resin (Qiagen), and the eluted proteins were digested with HRV3C (Novagen) before passage over a 16/60 S200 size exclusion column. Monodisperse fractions were pooled and passed over Ni2+-NTA resin to remove any uncleaved scaffold or residual HRV3C protease. The scaffolds were flash frozen in liquid nitrogen and stored at −80 °C. Glycosylation mutants were expressed and purified in a similar manner.
Expression and purification of PG9 N23Q HRV3C
A mammalian codon-optimized gene encoding the PG9 heavy chain with an HRV3C recognition site (GLEVLFQGP) inserted after Lys 235 was synthesized and cloned into pVRC8400. Similarly, the PG9 light chain was synthesized and cloned into the pVRC8400 vector, and an N23Q mutation was introduced to remove the sole glycosylation site on PG9. The modified PG9 heavy and light chain plasmids were transiently co-transfected into HEK 293F cells, and IgG was purified from the supernatant after 5 days using Protein A agarose (Pierce).
Formation and purification of PG9–V1/V2 scaffold complexes
Approximately 3 mg of purified PG9 N23Q HRV3C IgG was bound to 750 μl Protein A Plus agarose (Pierce) in a disposable 10 ml column. To this resin was added 6 mg of purified V1/V2 scaffold (∼20-fold molar excess over PG9 IgG). After washing away unbound scaffold with PBS, the column was capped and 40 μl of HRV3C protease at 2 U μl−1 was added to the resin along with 1 ml of PBS. After 1 h at room temperature, the resin was drained, the eluate collected and passed over a 16/60 S200 column. Fractions corresponding to the PG9–V1/V2 complex were pooled and concentrated to ∼5 mg ml−1.
PG9–V1/V2 complex crystallization and data collection
A complex of PG9 bound to 1FD6-ZM109 with four N-linked asparagines mutated to alanine (Asn 160 and Asn 173 were retained) was screened against 576 crystallization conditions using a Cartesian Honeybee crystallization robot. Initial crystals were grown by the vapour diffusion method in sitting drops at 20 °C by mixing 0.2 μl of protein complex with 0.2 μl of reservoir solution50 (17% (w/v) PEG 3350, 10% (v/v) 2-methyl-2,4-pentanediol, 0.2 M lithium sulphate, 0.1 M imidazole pH 6.5). Crystals suitable for diffraction were manually reproduced in hanging drops by mixing equal volumes of protein complex with reservoir solution50 (8% (w/v) PEG 3350, 5% (v/v) 2-methyl-2,4-pentanediol, 90 mM lithium sulphate, 45 mM imidazole pH 6.5). Single crystals were flash frozen in liquid nitrogen in 12% (w/v) PEG 3350, 0.2 M lithium sulphate, 0.1 M imidazole pH 6.5 and 15% (v/v) 2R,3R-butanediol. Data to 1.80 Å were collected at a wavelength of 1.00 Å at the SER-CAT beamline ID-22 (Advanced Photon Source, Argonne National Laboratory).
A complex of PG9 and 1FD6-CAP45 at 2.2 mg ml−1 was also screened against 576 crystallization conditions. Initial crystals were grown in the same reservoir solution as for PG9-1FD6-ZM109. Crystals were manually reproduced in hanging drops by mixing equal volumes of protein complex with reservoir solution50 (13% (w/v) PEG 3350, 11% (v/v) 2-methyl-2,4-pentanediol, 0.2 M lithium sulphate, 0.1 M imidazole pH 6.5). Single crystals were bathed in a cryoprotectant of 20% (w/v) PEG 3350, 0.2 M lithium sulphate, 0.1 M imidazole pH 6.5, and 15% (v/v) 2R,3R-butanediol followed by immersion in Paratone-N and flash frozen in liquid nitrogen. Data to 2.19 Å were collected at a wavelength of 1.00 Å at the SER-CAT beamline BM-22.
PG9–V1/V2 complex structure determination, model building and refinement
Diffraction data were processed with the HKL2000 suite52 and a molecular replacement solution for the 1FD6-ZM109 data set consisting of two unbound PG9 Fab molecules per asymmetric unit was obtained using PHASER53. Model building was carried out using COOT54, and refinement was performed with PHENIX55. Electron density for the Man5GlcNAc2 attached to Asn 160 and the two disulphide bonds were used as landmarks to build the V1/V2 strands. Final data collection and refinement statistics are presented in Supplementary Table 7. The Ramachandran plot as determined by MOLPROBITY56 shows 98.0% of all residues in favoured regions and 100% of all residues in allowed regions.
The PG9–1FD6-ZM109 structure was used as the search model for the PG9-1FD6-CAP45 data set. A molecular replacement solution consisting of two complexes per asymmetric unit was obtained using PHASER53, and COOT54 and PHENIX55 were used for model building and refinement, respectively. The Ramachandran plot for this complex as determined by MOLPROBITY56 shows 97.3% of all residues in favoured regions and 100% of all residues in allowed regions.
Surface plasmon resonance
The binding kinetics of different V1/V2 scaffolds to antibodies PG9 and PG16 were determined on a Biacore T-200 (GE Healthcare) at 25 °C with buffer HBS-EP+ (10 mM HEPES, pH 7.4, 150 mM NaCl, 3 mM EDTA, and 0.05% surfactant P-20). For comparison, PG9 and PG16 binding to full-length HIV-1 gp120 proteins was performed in parallel. The effects of the gp120 V3 loop on antibody binding were also assessed with V3 loop-deleted gp120 proteins. In total, five full-length gp120 proteins (strains ZM109, 16055, AD244, CAP45, and TRJO), two V3 loop-deleted gp120 proteins (16055ΔV3 and AD244ΔV3) and five V1/V2 scaffolds (1FD6-ZM109, 1JO8-ZM109, 1FD6-16055, 1JO8-CAP45 and 1JO8-TRJO) were immobilized onto CM5 chips to 500 response units (R.U.) with standard amine coupling. PG9 Fab and PG16 Fab were injected over the channels at 2-fold increasing concentrations with a flow rate of 30 μl min−1 for 3 min and allowed to dissociate for another 5 min. Regenerations were performed with one 25 μl injection of 3.0 M MgCl2 at a flow rate of 50 μl min−1 following the dissociation phase. T-200 Biacore Evaluation software was used to subtract appropriate blank references and fit sensorgrams globally using a 1:1 Langmuir model. In some cases, especially the binding to V1/V2 scaffolds, the sensorgrams could not reasonably be fit to a 1:1 Langmuir model due to heterogeneity of the immobilized ligands, and thus a 1:1 model assuming heterogeneous ligands was used. The relative percentage of each component in the heterogeneous ligands was calculated by its contribution to the total Rmax and the kinetic parameters are listed separately in Supplementary figures. Mass transfer effects were assessed by the tc values given by the T-200 Biacore Evaluation software. No significant mass transport effects were detected in all measurements (tc > 1010).
Electron microscopy and image processing
Negative stained grids were prepared by applying 40 μg ml−1 of the purified T13-gp120 16055 (82-492)-PG9 ternary complex to a freshly glow discharged carbon-coated 400 Cu mesh grid and stained with 2% uranyl formate. Grids were viewed using a FEI Tecnai TF20 electron microscope operating at a high tension of 120 kV at the National Resource for Automated Molecular Microscopy. Initial models were generated using the random conical tilt method through the Appion package57,58. Images were acquired at a magnification of ×62,000 with a defocus range of 1.5 to 2.5 μm onto a Gatan 4k × 4k CCD using the Leginon package59. The pixel size of the CCD was calibrated using a two-dimensional catalase crystal with known cell parameters. The initial models were improved using a data set collected at a magnification of ×150,000 at 0°, 15°, 30°, 45° and 55° tilts with a defocus range of 500–700 nm through a multi-model approach developed in-house with the SPIDER package60. The tilts provided additional particle orientations to improve the image reconstructions.
PG9 Fab crystallization and refinement
PG9 Fab with an N23Q mutation in the light chain was obtained by cleaving the recombinant IgG described above with HRV3C protease, followed by gel filtration chromatography. PG9 Fab at a concentration of 13.7 mg ml−1 was screened against 576 crystallization conditions, and initial crystals were obtained using the sitting-drop vapour diffusion method. Crystals were obtained from a reservoir50 containing 25% (w/v) PEG 3350, 15% (v/v) 2-methyl-2,4-pentanediol, 0.2 M lithium sulphate, 0.1 M imidazole pH 6.5. After cryo-protection with 15% 2R,3R-butanediol, crystals were mounted and flash frozen in liquid nitrogen. Data to 3.30 Å were collected at a wavelength of 1.00 Å at the SER-CAT beamline ID-22. Statistics for data collection and data processing in HKL200052 are summarized in Supplementary Table 19. The structure in space group P1 was solved by molecular replacement using the program PHASER53 with the PG16 Fab structure (PDB ID 3LRS)30 as a search model. Model building and refinement were performed using COOT54 and PHENIX55, respectively. Refinement statistics for the PG9 Fab model are reported in Supplementary Table 19.
CH04 and CH04H/CH02L Fab expression, crystallization and refinement
A mammalian codon-optimized gene encoding the CH04 heavy chain with a stop codon inserted after Asp 234 was synthesized and cloned into pVRC8400. Similarly, the CH04 and CH02 light chains were synthesized and cloned into the pVRC8400 vector. The CH04 heavy and light chain plasmids were transiently co-transfected into HEK 293F cells (or CH04 heavy with CH02 light chain), and Fab was purified from the supernatant after 5 days using Kappa agarose column (CaptureSelect Fab κ; BAC). CH04 and CH04H/CH02L Fabs at a concentration of 16 mg ml−1 and 10 mg ml−1, respectively, were screened against 576 crystallization conditions using a Cartesian Honeybee crystallization robot. CH04 Fab crystals were obtained in 20% (w/v) PEG 8000, 3% (v/v) 2-methyl-2,4-pentanediol, 70 mM imidazole pH 6.5. Single crystals were flash frozen in liquid nitrogen in 24% (w/v) PEG 8000, 3.4% (v/v) 2-methyl-2,4-pentanediol, 85 mM imidazole pH 6.5 and 15% (v/v) 2R,3R-butanediol. CH04H/CH02L Fab crystals were obtained in 16% PEG 400, 8% PEG 8000, 0.1 M acetate pH 4.5 (orthorhombic forms) and 15% PEG 3350, 9% 2-methyl-2,4-pentanediol, 0.1 M lithium sulphate, 0.1 M imidazole pH 6.5 (tetragonal forms). Data to 1.90 Å (CH04 Fab) and 2.90 Å (CH04H/CH02L Fab) were collected at a wavelength of 1.00 Å at the SER-CAT beamline ID-22 and BM-22, respectively.
Diffraction data were processed with the HKL2000 suite52 and a molecular replacement solution for the CH04 data set consisting of two CH04 Fab molecules per asymmetric unit was obtained using PHASER53 and PDB ID codes 1DFB (heavy chain)61 and 1QLR (light chain)62 as search models. CH04 Fab was used as the search model for CH04H/CH02L. Model building was carried out using COOT54, and refinement was performed with PHENIX55. Final data collection and refinement statistics are presented in Supplementary Table 19.
PGT145 Fab expression, crystallization and refinement
Expression and purification of PGT145 was performed using a similar protocol to that previously described27. Briefly, the Fab was produced as a secreted protein by co-transfecting the heavy and light chain genes into HEK 293T cells. Three days after transfection, the media was recovered, concentrated and flowed over an anti-human κ light chain affinity matrix (CaptureSelect Fab κ; BAC). The eluted fraction containing the Fab was further purified by cation exchange chromatography followed by size-exclusion chromatography. PGT145 Fab at a concentration of 10 mg ml−1 was crystallized using the sitting-drop vapour diffusion method. Crystals were obtained in a mother liquor containing 0.1 M HEPES, pH 7.5, 2 M ammonium sulphate and 20% PEG 400. After cryo-protection in 20% glycerol, crystals were mounted and flash frozen in liquid nitrogen. PGT145 Fab crystals were exposed to a monochromatic X-ray beam at the Advanced Photon Source Sector 23-ID (Argonne National Laboratory). Statistics for data collection and data processing in HKL200052 are summarized in Supplementary Table 19. The structure in space group P41212 was solved by molecular replacement using the program PHASER53 with the PG16 Fab structure (PDB ID 3MUG)27 as a search model. Refinement of the structure was performed using a combination of CNS63, CCP464 and COOT54. The final statistics of the refined PGT145 Fab model are reported in Supplementary Table 19.
STD experiments by NMR
All NMR experiments were carried out at 298 K on Bruker avance 600 or avance 500 instruments equipped with a triple resonance cryo-probe incorporating gradients in z-axis. 1D STD spectra were acquired by selectively irradiating at −1 p.p.m. and +40 p.p.m. as on- and off-resonance frequencies, respectively, using a train of 50 ms Gaussian-shaped radio frequency pulses separated by 1-ms delays and an optimized power level of 57 dB. During NMR experiments water suppression was achieved by binomial 3-9-19 pulse sequence and protein resonances were suppressed by applying 10 ms T1ρ filter. Samples were prepared in 20 mM sodium phosphate buffer containing 50 mM sodium chloride at pH 6.8. The NMR data were processed and analysed by using TOPSPIN 2.1. The STD amplification factor, ASTD, was obtained according to the equation, ASTD = (I0 – ISAT) I0−1 ([Lt]/[P]), where Lt and P are the total ligand and protein concentrations, respectively65.
Surface areas and average surface electrostatic potential calculations
Surface area calculations were performed using PISA66 and MS67. The interactive surfaces with PG9 for CAP45 and ZM109 were obtained using PyMOL68 and selecting atoms of V1/V2 within 5.5 Å of PG9 residues. Electrostatic surface potentials for the CDR H3 and interacting surface for CAP45 and ZM109 were obtained using GRASP69. The Poisson–Boltzmann (PB) potential grid map and surface points of each CDR H3 region and CAP45 and ZM109 interacting surfaces were determined using GRASP. The PB potential for each surface point was determined by trilinear interpolation from the values of the eight corners of the cube where the surface point resided in. The average surface PB potential is the linear average of the PB potentials of all surface points.
Figures
Structure figures were prepared using PYMOL68.
Accession codes
Primary accessions
Protein Data Bank
Data deposits
Coordinates and structure factors for PG9 Fab in complexes with V1/V2 from CAP45 and ZM109 strains of HIV-1 have been deposited with the Protein Data Bank under accession codes 3U4E and 3U2S, respectively. Coordinates and structure factors for unbound Fab structures of PG9, CH04, CH04H/CH02L (in two lattices) and PGT145 have been deposited with the Protein Data Bank under accession codes, 3U36, 3TCL, 3U46, 3U4B and 3U1S, respectively.
References
Wyatt, R. & Sodroski, J. The HIV-1 envelope glycoproteins: fusogens, antigens, and immunogens. Science 280, 1884–1888 (1998)
Liu, J., Bartesaghi, A., Borgnia, M. J., Sapiro, G. & Subramaniam, S. Molecular architecture of native HIV-1 gp120 trimers. Nature 455, 109–113 (2008)
Wu, S. R. et al. Single-particle cryoelectron microscopy analysis reveals the HIV-1 spike as a tripod structure. Proc. Natl Acad. Sci. USA 107, 18844–18849 (2010)
White, T. A. et al. Molecular architectures of trimeric SIV and HIV-1 envelope glycoproteins on intact viruses: strain-dependent variation in quaternary structure. PLoS Pathog. 6, e1001249 (2010)
Hu, G., Liu, J., Taylor, K. A. & Roux, K. H. Structural comparison of HIV-1 envelope spikes with and without the V1/V2 loop. J. Virol. 85, 2741–2750 (2011)
Cao, J. et al. Replication and neutralization of human immunodeficiency virus type 1 lacking the V1 and V2 variable loops of the gp120 envelope glycoprotein. J. Virol. 71, 9808–9812 (1997)
Stamatatos, L. & Cheng-Mayer, C. An envelope modification that renders a primary, neutralization-resistant clade B human immunodeficiency virus type 1 isolate highly susceptible to neutralization by sera from other clades. J. Virol. 72, 7840–7845 (1998)
Pinter, A. et al. The V1/V2 domain of gp120 is a global regulator of the sensitivity of primary human immunodeficiency virus type 1 isolates to neutralization by antibodies commonly induced upon infection. J. Virol. 78, 5205–5215 (2004)
Rusert, P. et al. Interaction of the gp120 V1/V2 loop with a neighboring gp120 unit shields the HIV envelope trimer against cross-neutralizing antibodies. J. Exp. Med. 208, 1419–1433 (2011)
Walker, L. M. et al. Broad and potent neutralizing antibodies from an African donor reveal a new HIV-1 vaccine target. Science 326, 285–289 (2009)
Bonsignori, M. et al. Analysis of a clonal lineage of HIV-1 envelope V2/V3 conformational epitope-specific broadly neutralizing antibodies and their inferred unmutated common ancestors. J. Virol. 85, 9998–10009 (2011)
Walker, L. M. et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477, 466–470 (2011)
Walker, L. M. et al. A limited number of antibody specificities mediate broad and potent serum neutralization in selected HIV-1 infected individuals. PLoS Pathog. 6, e1001028 (2010)
Moore, P. L. et al. Potent and broad neutralization of HIV-1 subtype C by plasma antibodies targeting a quaternary epitope including residues in the V2 loop. J. Virol. 85, 3128–3141 (2011)
Kwong, P. D. et al. Probability analysis of variational crystallization and its application to gp120, the exterior envelope glycoprotein of type 1 human immunodeficiency virus (HIV-1). J. Biol. Chem. 274, 4115–4123 (1999)
Huang, C. C. et al. Structure of a V3-containing HIV-1 gp120 core. Science 310, 1025–1028 (2005)
Pancera, M. et al. Structure of HIV-1 gp120 with gp41-interactive region reveals layered envelope architecture and basis of conformational mobility. Proc. Natl Acad. Sci. USA 107, 1166–1171 (2010)
Kwong, P. D. et al. Structure of an HIV gp120 envelope glycoprotein in complex with the CD4 receptor and a neutralizing human antibody. Nature 393, 648–659 (1998)
Zhou, T. et al. Structural definition of a conserved neutralization epitope on HIV-1 gp120. Nature 445, 732–737 (2007)
Chen, L. et al. Structural basis of immune evasion at the site of CD4 attachment on HIV-1 gp120. Science 326, 1123–1127 (2009)
Kwong, P. D. et al. Structures of HIV-1 gp120 envelope glycoproteins from laboratory-adapted and primary isolates. Structure 8, 1329–1339 (2000)
Chen, B. et al. Structure of an unliganded simian immunodeficiency virus gp120 core. Nature 433, 834–841 (2005)
Ross, S. A., Sarisky, C. A., Su, A. & Mayo, S. L. Designed protein G core variants fold to native-like structures: sequence selection by ORBIT tolerates variation in backbone specification. Protein Sci. 10, 450–454 (2001)
Fazi, B. et al. Unusual binding properties of the SH3 domain of the yeast actin-binding protein Abp1: structural and functional analysis. J. Biol. Chem. 277, 5290–5298 (2002)
Arthos, J. et al. HIV-1 envelope protein binds to and signals through integrin α4β7, the gut mucosal homing receptor for peripheral T cells. Nature Immunol. 9, 301–309 (2008)
Reeves, P. J., Callewaert, N., Contreras, R. & Khorana, H. G. Structure and function in rhodopsin: high-level expression of rhodopsin with restricted and homogeneous N-glycosylation by a tetracycline-inducible N-acetylglucosaminyltransferase I-negative HEK293S stable mammalian cell line. Proc. Natl Acad. Sci. USA 99, 13419–13424 (2002)
Pejchal, R. et al. Structure and function of broadly reactive antibody PG16 reveal an H3 subdomain that mediates potent neutralization of HIV-1. Proc. Natl Acad. Sci. USA 107, 11483–11488 (2010)
Richardson, J. S. The anatomy and taxonomy of protein structure. Adv. Protein Chem. 34, 167–339 (1981)
Kabat, E. A., Wu, T. T., Perry, H. M., Gottesman, K. S. & Foeller, C. Sequences of Proteins of Immunological Interest 5th edn (US Department of Health and Human Service, National Institutes of Health, 1991)
Pancera, M. et al. Crystal structure of PG16 and chimeric dissection with somatically related PG9: structure–function analysis of two quaternary-specific antibodies that effectively neutralize HIV-1. J. Virol. 84, 8098–8110 (2010)
Gorny, M. K. et al. Identification of a new quaternary neutralizing epitope on human immunodeficiency virus type 1 virus particles. J. Virol. 79, 5232–5237 (2005)
Changela, A. et al. Crystal structure of human antibody 2909 reveals conserved features of quaternary structure-specific antibodies that potently neutralize HIV-1. J. Virol. 85, 2524–2535 (2011)
Spurrier, B. et al. Structural analysis of human and macaque mAbs 2909 and 2.5B: implications for the configuration of the quaternary neutralizing epitope of HIV-1 gp120. Structure 19, 691–699 (2011)
Honnen, W. J. et al. Type-specific epitopes targeted by monoclonal antibodies with exceptionally potent neutralizing activities for selected strains of human immunodeficiency virus type 1 map to a common region of the V2 domain of gp120 and differ only at single positions from the clade B consensus sequence. J. Virol. 81, 1424–1432 (2007)
Wu, X. et al. Immunotypes of a quaternary site of HIV-1 vulnerability and their recognition by antibodies. J. Virol. 85, 4578–4585 (2011)
Zhou, T. et al. Structural basis for broad and potent neutralization of HIV-1 by antibody VRC01. Science 329, 811–817 (2010)
Mitsuya, H. et al. Dextran sulfate suppression of viruses in the HIV family: inhibition of virion binding to CD4+ cells. Science 240, 646–649 (1988)
Schols, D., Pauwels, R., Desmyter, J. & De Clercq, E. Dextran sulfate and other polyanionic anti-HIV compounds specifically interact with the viral gp120 glycoprotein expressed by T-cells persistently infected with HIV-1. Virology 175, 556–561 (1990)
Moulard, M. et al. Selective interactions of polyanions with basic surfaces on human immunodeficiency virus type 1 gp120. J. Virol. 74, 1948–1960 (2000)
Fletcher, P. S., Wallace, G. S., Mesquita, P. M. & Shattock, R. J. Candidate polyanion microbicides inhibit HIV-1 infection and dissemination pathways in human cervical explants. Retrovirology 3, 46 (2006)
Mondor, I., Ugolini, S. & Sattentau, Q. J. Human immunodeficiency virus type 1 attachment to HeLa CD4 cells is CD4 independent and gp120 dependent and requires cell surface heparans. J. Virol. 72, 3623–3634 (1998)
Stanfield, R. L., Gorny, M. K., Williams, C., Zolla-Pazner, S. & Wilson, I. A. Structural rationale for the broad neutralization of HIV-1 by human monoclonal antibody 447-52D. Structure 12, 193–204 (2004)
Gray, E. S. et al. The neutralization breadth of HIV-1 develops incrementally over four years and is associated with CD4 T cell decline and high viral load during acute infection. J. Virol. 85, 4828–4840 (2011)
Moore, P. L. et al. Evolution of HIV-1 transmitted/founder viruses results in the formation of epitopes for later broadly cross-neutralizing antibodies. AIDS Res. Hum. Retroviruses 27, A-29 (2011)
Pejchal, R. et al. A potent and broad neutralizing antibody recognizes and penetrates the HIV glycan shield. Science 10.1126/science.1213256 (13 October 2011)
Karasavvas, N. et al. The Thai Phase III clinical trial (RV144) induces the generation of antibodies that target a conserved region within the V2 loop of gp120. AIDS Res. Hum. Retroviruses 27, A-29 (2011)
Zolla-Pazner, S. et al. V2-reactive antibodies in RV144 vaccinees’ plasma. AIDS Res. Hum. Retroviruses 27, A-21 (2011)
Azoitei, M. L. et al. Computation-guided backbone grafting of a discontinuous motif onto a protein scaffold. Science 334, 373–376 (2011)
Sloan, D. J. & Hellinga, H. W. Dissection of the protein G B1 domain binding site for human IgG Fc fragment. Protein Sci. 8, 1643–1648 (1999)
Majeed, S. et al. Enhancing protein crystallization through precipitant synergy. Structure 11, 1061–1070 (2003)
Huang, P.-S. et al. RosettaRemodel: A generalized framework for flexible backbone protein design. PLoS ONE 6, e24109 (2011)
Otwinowski, Z. & Minor, W. in Methods Enzymol. Vol. 276, 307–326 (Academic Press, 1997)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007)
Lander, G. C. et al. Appion: an integrated, database-driven pipeline to facilitate EM image processing. J. Struct. Biol. 166, 95–102 (2009)
Radermacher, M., Wagenknecht, T., Verschoor, A. & Frank, J. Three-dimensional reconstruction from a single-exposure, random conical tilt series applied to the 50S ribosomal subunit of Escherichia coli. J. Microsc. 146, 113–136 (1987)
Suloway, C. et al. Automated molecular microscopy: the new Leginon system. J. Struct. Biol. 151, 41–60 (2005)
Frank, J., Shimkin, B. & Dowse, H. SPIDER-a modular software system for electron image processin. Ultramicroscopy 6, 343–358 (1981)
He, X. M., Rüker, F., Casale, E. & Carter, D. C. Structure of a human monoclonal antibody Fab fragment against gp41 of human immunodeficiency virus type 1. Proc. Natl Acad. Sci. USA 89, 7154–7158 (1992)
Cauerhff, A. et al. Three-dimensional structure of the Fab from a human IgM cold agglutinin. J. Immunol. 165, 6422–6428 (2000)
Brunger, A. T. et al. Crystallography & NMR system: A new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011)
Mayer, M. & Meyer, B. Group epitope mapping by saturation transfer difference NMR to identify segments of a ligand in direct contact with a protein receptor. J. Am. Chem. Soc. 123, 6108–6117 (2001)
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007)
Connolly, M. L. Analytical molecular surface calculation. J. Appl. Cryst. 16, 548–558 (1983)
The PyMOL Molecular Graphics System. Version 1.4 (Schrödinger, LLC, 2011).
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991)
Acknowledgements
We thank A. Changela and X. Wu for assistance with mapping the PG16 paratope, J. Sodroski for discussions, J. Stuckey for assistance with figures, and members of the Structural Biology Section and Structural Bioinformatics Core, Vaccine Research Center, for discussions and comments on the manuscript. Support for this work was provided by the Intramural Research Program of the National Institutes of Health (NIH), by the International AIDS Vaccine Initiative, by the Ragon Institute, by the Canadian Institute of Health Research and by grants from the NIH. The three-dimensional reconstructions were conducted at the National Resource for Automated Molecular Microscopy (NRAMM), which is supported by the NIH through the National Center for Research Resources' P41 program (RR017573). Use of sector 22 (Southeast Regional Collaborative Access Team) at the Advanced Photon Source was supported by the US Department of Energy, Basic Energy Sciences, Office of Science, under contract number W-31-109-Eng-38.
Author information
Authors and Affiliations
Contributions
J.S.M., M.P., M.S., T.Z., J.Z., J.A., C.A.B., J.R.M., G.J.N., W.R.S., A.B.W., I.A.W. and P.D.K. designed research and analysed the data; J.S.M., M.P., C.C., J.G., J.-P.J., R.K., R.L., R.P., M.S., K.D., S.O’D., N.P., S.S.H., Y.Y., T.Z., J.C.B., G.-Y.C., D.D., I.G., Y.D.K., D.L., M.K.L., S.M., S.D.S., Z.-Y.Y. and B.Z. performed research and are listed in four alphabetical groups: J.S.M. and M.P. performed the majority of research and, with C.C., J.G., J.-P.J., R.K., R.L., R.P. and M.S., determined structures, K.D., S.O’D., N.P., S.S.H., Y.Y., B.Z., T.Z. and J.Z. contributed substantial experiments, J.C.B., G.-Y.C., D.D., I.G., Y.D.K., D.L., M.K.L., S.M., S.D.S. and Z.-Y.Y. contributed supporting research; M.B., J.A.C., S.H.K., N.E.S. and B.F.H. contributed donor 0219 materials; D.R.B., W.C.K. and L.M.W. contributed donor 24 and donor 84 materials and T13 antibody; S.P. and R.W. contributed 16055 gp120; J.O. and L.-X.W. contributed polysaccharides; J.S.M., M.P., G.J.N., I.A.W. and P.D.K. wrote the paper, with all principal investigators providing comments or revisions.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Tables 1-21 and Supplementary Figures 1-20 with legends. (PDF 3375 kb)
Rights and permissions
About this article
Cite this article
McLellan, J., Pancera, M., Carrico, C. et al. Structure of HIV-1 gp120 V1/V2 domain with broadly neutralizing antibody PG9. Nature 480, 336–343 (2011). https://doi.org/10.1038/nature10696
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10696
This article is cited by
-
Probabilities of developing HIV-1 bNAb sequence features in uninfected and chronically infected individuals
Nature Communications (2023)
-
Antibody-directed evolution reveals a mechanism for enhanced neutralization at the HIV-1 fusion peptide site
Nature Communications (2023)
-
Differential V2-directed antibody responses in non-human primates infected with SHIVs or immunized with diverse HIV vaccines
Nature Communications (2022)
-
Fine-mapping the immunodominant antibody epitopes on consensus sequence-based HIV-1 envelope trimer vaccine candidates
npj Vaccines (2022)
-
Using cross-species vaccination approaches to counter emerging infectious diseases
Nature Reviews Immunology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.