Abstract
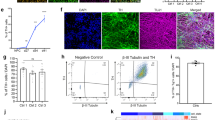
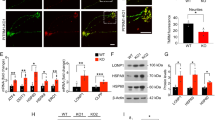
Machado–Joseph disease (MJD; also called spinocerebellar ataxia type 3) is a dominantly inherited late-onset neurodegenerative disorder caused by expansion of polyglutamine (polyQ)-encoding CAG repeats in the MJD1 gene (also known as ATXN3). Proteolytic liberation of highly aggregation-prone polyQ fragments from the protective sequence of the MJD1 gene product ataxin 3 (ATXN3) has been proposed to trigger the formation of ATXN3-containing aggregates, the neuropathological hallmark of MJD1,2,3,4,5. ATXN3 fragments are detected in brain tissue of MJD patients and transgenic mice expressing mutant human ATXN3(Q71)6, and their amount increases with disease severity, supporting a relationship between ATXN3 processing and disease progression. The formation of early aggregation intermediates is thought to have a critical role in disease initiation7,8, but the precise pathogenic mechanism operating in MJD has remained elusive9. Here we show that l-glutamate-induced excitation of patient-specific induced pluripotent stem cell (iPSC)-derived neurons initiates Ca2+-dependent proteolysis of ATXN3 followed by the formation of SDS-insoluble aggregates. This phenotype could be abolished by calpain inhibition, confirming a key role of this protease in ATXN3 aggregation. Aggregate formation was further dependent on functional Na+ and K+ channels as well as ionotropic and voltage-gated Ca2+ channels, and was not observed in iPSCs, fibroblasts or glia, thereby providing an explanation for the neuron-specific phenotype of this disease. Our data illustrate that iPSCs enable the study of aberrant protein processing associated with late-onset neurodegenerative disorders in patient-specific neurons.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Berke, S. J., Schmied, F. A., Brunt, E. R., Ellerby, L. M. & Paulson, H. L. Caspase-mediated proteolysis of the polyglutamine disease protein ataxin-3. J. Neurochem. 89, 908–918 (2004)
Jung, J., Xu, K., Lessing, D. & Bonini, N. M. Preventing ataxin-3 protein cleavage mitigates degeneration in a Drosophila model of SCA3. Hum. Mol. Genet. 18, 4843–4852 (2009)
Tarlac, V. & Storey, E. Role of proteolysis in polyglutamine disorders. J. Neurosci. Res. 74, 406–416 (2003)
Wellington, C. L. et al. Caspase cleavage of gene products associated with triplet expansion disorders generates truncated fragments containing the polyglutamine tract. J. Biol. Chem. 273, 9158–9167 (1998)
Rubinsztein, D. C., Wyttenbach, A. & Rankin, J. Intracellular inclusions, pathological markers in diseases caused by expanded polyglutamine tracts? J. Med. Genet. 36, 265–270 (1999)
Goti, D. et al. A mutant ataxin-3 putative-cleavage fragment in brains of Machado–Joseph disease patients and transgenic mice is cytotoxic above a critical concentration. J. Neurosci. 24, 10266–10279 (2004)
Chen, S., Ferrone, F. A. & Wetzel, R. Huntington’s disease age-of-onset linked to polyglutamine aggregation nucleation. Proc. Natl Acad. Sci. USA 99, 11884–11889 (2002)
Schaffar, G. et al. Cellular toxicity of polyglutamine expansion proteins: mechanism of transcription factor deactivation. Mol. Cell 15, 95–105 (2004)
Gatchel, J. R. & Zoghbi, H. Y. Diseases of unstable repeat expansion: mechanisms and common principles. Nature Rev. Genet. 6, 743–755 (2005)
Williams, A. J. & Paulson, H. L. Polyglutamine neurodegeneration: protein misfolding revisited. Trends Neurosci. 31, 521–528 (2008)
Berridge, M. J. Neuronal calcium signaling. Neuron 21, 13–26 (1998)
Koch, P., Opitz, T., Steinbeck, J. A., Ladewig, J. & Brüstle, O. A rosette-type, self-renewing human ES cell-derived neural stem cell with potential for in vitro instruction and synaptic integration. Proc. Natl Acad. Sci. USA 106, 3225–3230 (2009)
Thakur, A. K. & Wetzel, R. Mutational analysis of the structural organization of polyglutamine aggregates. Proc. Natl Acad. Sci. USA 99, 17014–17019 (2002)
Ikeda, H. et al. Expanded polyglutamine in the Machado–Joseph disease protein induces cell death in vitro and in vivo. Nature Genet. 13, 196–202 (1996)
Scherzinger, E. et al. Huntingtin-encoded polyglutamine expansions form amyloid-like protein aggregates in vitro and in vivo. Cell 90, 549–558 (1997)
Haacke, A. et al. Proteolytic cleavage of polyglutamine-expanded ataxin-3 is critical for aggregation and sequestration of non-expanded ataxin-3. Hum. Mol. Genet. 15, 555–568 (2006)
Haacke, A., Hartl, F. U. & Breuer, P. Calpain inhibition is sufficient to suppress aggregation of polyglutamine-expanded ataxin-3. J. Biol. Chem. 282, 18851–18856 (2007)
Uchihara, T. et al. Non-expanded polyglutamine proteins in intranuclear inclusions of hereditary ataxias: triple-labeling immunofluorescence study. Acta Neuropathol. 102, 149–152 (2001)
Hartl, F. U., Bracher, A. & Hayer-Hartl, M. Molecular chaperones in protein folding and proteostasis. Nature 475, 324–332 (2011)
Yoo, S. Y. et al. SCA7 knockin mice model human SCA7 and reveal gradual accumulation of mutant ataxin-7 in neurons and abnormalities in short-term plasticity. Neuron 37, 383–401 (2003)
Watase, K. et al. A long CAG repeat in the mouse Sca1 locus replicates SCA1 features and reveals the impact of protein solubility on selective neurodegeneration. Neuron 34, 905–919 (2002)
Li, M., Chevalier-Larsen, E. S., Merry, D. E. & Diamond, M. I. Soluble androgen receptor oligomers underlie pathology in a mouse model of spinobulbar muscular atrophy. J. Biol. Chem. 282, 3157–3164 (2007)
Williams, A. J., Knutson, T. M., Colomer Gould, V. F. & Paulson, H. L. In vivo suppression of polyglutamine neurotoxicity by C-terminus of Hsp70-interacting protein (CHIP) supports an aggregation model of pathogenesis. Neurobiol. Dis. 33, 342–353 (2009)
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007)
Hazeki, N., Tukamoto, T., Goto, J. & Kanazawa, I. Formic acid dissolves aggregates of an N-terminal huntingtin fragment containing an expanded polyglutamine tract: applying to quantification of protein components of the aggregates. Biochem. Biophys. Res. Commun. 277, 386–393 (2000)
Acknowledgements
We thank S. Auel, D. Kühne, M. Segschneider and V. Dobberahn for technical support, A. Leinhaas for conducting the teratoma assays and H. L. Paulson for providing the polyclonal ATXN3 antibody used for immunocytochemistry. The human embryonic stem cell lines H9 and I3 (used as control for quantitative PCR) and I6 (originally used to derive the lt-NES cells) were provided by J. Itskovitz-Eldor (Technion, Israel Institute of Technology, Haifa, Israel). The work was supported by the German Federal Ministry for Education and Research (BMBF; grants 01GNO813, 01GS0860), the European Union (LSHG-CT-2006-018739, ESTOOLS; HEALTH-F5-2010-266753, SCR&Tox), the Deutsche Forschungsgemeinschaft (WU184/6-1, EV143/1-1), BONFOR and the Hertie Foundation.
Author information
Authors and Affiliations
Contributions
P.K. and P.B.: conception and study design, directed differentiation of iPSCs, cellular/molecular assays for disease modelling, data assembly, analysis and interpretation, writing of manuscript; M.P. and J.J.: iPSC clone derivation, maintenance and validation, directed differentiation of iPSCs; J.K., D.P., J.D., J.L., J.M., P.H.: data collection, analysis and interpretation; B.O.E., U.W., T.K.: conception, data analysis and interpretation; T.T.: provision of material; O.B.: conception, data analysis and interpretation, writing of manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-16 with legends. (PDF 12324 kb)
Rights and permissions
About this article
Cite this article
Koch, P., Breuer, P., Peitz, M. et al. Excitation-induced ataxin-3 aggregation in neurons from patients with Machado–Joseph disease. Nature 480, 543–546 (2011). https://doi.org/10.1038/nature10671
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10671
This article is cited by
-
Establishment and characterization of human pluripotent stem cells-derived brain organoids to model cerebellar diseases
Scientific Reports (2022)
-
Calpains as novel players in the molecular pathogenesis of spinocerebellar ataxia type 17
Cellular and Molecular Life Sciences (2022)
-
Human Induced Pluripotent Stem Cell-Based Modelling of Spinocerebellar Ataxias
Stem Cell Reviews and Reports (2022)
-
Therapeutic roles of natural remedies in combating hereditary ataxia: A systematic review
Chinese Medicine (2021)
-
CRISPR/Cas9 mediated gene correction ameliorates abnormal phenotypes in spinocerebellar ataxia type 3 patient-derived induced pluripotent stem cells
Translational Psychiatry (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.