Abstract
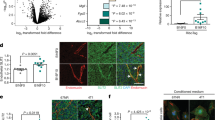
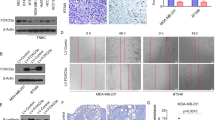
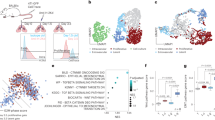
Metastatic progression of cancer is a complex and clinically daunting process1,2,3,4. We previously identified a set of human microRNAs (miRNAs) that robustly suppress breast cancer metastasis to lung and bone5,6 and which display expression levels that predict human metastasis. Although these findings revealed miRNAs as suppressors of cell-autonomous metastatic phenotypes, the roles of non-coding RNAs in non-cell-autonomous cancer progression processes remain unknown. Here we reveal that endogenous miR-126, an miRNA silenced in a variety of common human cancers7,8, non-cell-autonomously regulates endothelial cell recruitment to metastatic breast cancer cells, in vitro and in vivo. It suppresses metastatic endothelial recruitment, metastatic angiogenesis and metastatic colonization through coordinate targeting of IGFBP2, PITPNC1 and MERTK—novel pro-angiogenic genes and biomarkers of human metastasis. Insulin-like growth factor binding protein 2 (IGFBP2) secreted by metastatic cells recruits endothelia by modulating IGF1-mediated activation of the IGF type-I receptor on endothelial cells; whereas c-Mer tyrosine kinase (MERTK) receptor cleaved from metastatic cells promotes endothelial recruitment by competitively antagonizing the binding of its ligand GAS6 to endothelial MERTK receptors. Co-injection of endothelial cells with breast cancer cells non-cell-autonomously rescues their miR-126-induced metastatic defect, revealing a novel and important role for endothelial interactions in metastatic initiation. Through loss-of-function and epistasis experiments, we delineate an miRNA regulatory network’s individual components as novel and cell-extrinsic regulators of endothelial recruitment, angiogenesis and metastatic colonization. We also identify the IGFBP2/IGF1/IGF1R and GAS6/MERTK signalling pathways as regulators of cancer-mediated endothelial recruitment. Our work further reveals endothelial recruitment and endothelial interactions in the tumour microenvironment to be critical features of metastatic breast cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chiang, A. C. & Massague, J. Molecular basis of metastasis. N. Engl. J. Med. 359, 2814–2823 (2008)
Talmadge, J. E. & Fidler, I. J. AACR centennial series: the biology of cancer metastasis: historical perspective. Cancer Res. 70, 5649–5669 (2010)
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011)
Valastyan, S. & Weinberg, R. A. MicroRNAs: crucial multi-tasking components in the complex circuitry of tumor metastasis. Cell Cycle 8, 3506–3512 (2009)
Tavazoie, S. F. et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451, 147–152 (2008)
Png, K. J. et al. MicroRNA-335 inhibits tumor reinitiation and is silenced through genetic and epigenetic mechanisms in human breast cancer. Genes Dev. 25, 226–231 (2011)
Guo, C. et al. The noncoding RNA, miR-126, suppresses the growth of neoplastic cells by targeting phosphatidylinositol 3-kinase signaling and is frequently lost in colon cancers. Genes Chromosom. Cancer 47, 939–946 (2008)
Feng, R. et al. miR-126 functions as a tumour suppressor in human gastric cancer. Cancer Lett. 298, 50–63 (2010)
Gupta, G. P. et al. Mediators of vascular remodelling co-opted for sequential steps in lung metastasis. Nature 446, 765–770 (2007)
Jones, J. I. & Clemmons, D. R. Insulin-like growth factors and their binding proteins: biological actions. Endocr. Rev. 16, 3–34 (1995)
Balogh, I. et al. Analysis of Gas6 in human platelets and plasma. Arterioscler. Thromb. Vasc. Biol. 25, 1280–1286 (2005)
Sather, S. et al. A soluble form of the Mer receptor tyrosine kinase inhibits macrophage clearance of apoptotic cells and platelet aggregation. Blood 109, 1026–1033 (2007)
Minn, A. J. et al. Genes that mediate breast cancer metastasis to lung. Nature 436, 518–524 (2005)
Kang, Y. et al. A multigenic program mediating breast cancer metastasis to bone. Cancer Cell 3, 537–549 (2003)
Yin, J. J. et al. TGF-beta signaling blockade inhibits PTHrP secretion by breast cancer cells and bone metastases development. J. Clin. Invest. 103, 197–206 (1999)
Arnold, S. A. et al. Lack of host SPARC enhances vascular function and tumor spread in an orthotopic murine model of pancreatic carcinoma. Dis Model Mech 3, 57–72 (2010)
Chin, K. et al. Genomic and transcriptional aberrations linked to breast cancer pathophysiologies. Cancer Cell 10, 529–541 (2006)
Wang, Y. et al. Gene-expression profiles to predict distant metastasis of lymph-node-negative primary breast cancer. Lancet 365, 671–679 (2005)
Acknowledgements
We thank the members of the Tavazoie laboratory, Saeed Tavazoie, M. Tavazoie, S. Kurdistani, C. Alarcon and E. Mandel for comments on previous versions of this manuscript. We thank V. Gueorguiev and the Memorial Sloan-Kettering Cancer Center’s Molecular Cytology Core Facility for help. We thank the Memorial Sloan-Kettering Cancer Center’s High-Throughput Screening Core Facility for providing the shRNAs. We thank H. Lee for cloning assistance. K.J.P. is an A*STAR National Science Scholar. S.F.T. is a Department of Defense Era of Hope Scholar and the Leon Hess Head of the Elizabeth and Vincent Mayer Laboratory at Rockefeller University.
Author information
Authors and Affiliations
Contributions
S.F.T. conceived the project and supervised all research. K.J.P., N.H, and S.F.T. wrote the manuscript. K.J.P., N.H. and S.F.T. designed the experiments. K.J.P., N.H. and M.Y. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-24 with legends, Supplementary Tables 1-9, a Supplementary Discussion and additional references. (PDF 4998 kb)
Rights and permissions
About this article
Cite this article
Png, K., Halberg, N., Yoshida, M. et al. A microRNA regulon that mediates endothelial recruitment and metastasis by cancer cells. Nature 481, 190–194 (2012). https://doi.org/10.1038/nature10661
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10661
This article is cited by
-
The various role of microRNAs in breast cancer angiogenesis, with a special focus on novel miRNA-based delivery strategies
Cancer Cell International (2023)
-
MiR-200/183 family-mediated module biomarker for gastric cancer progression: an AI-assisted bioinformatics method with experimental functional survey
Journal of Translational Medicine (2023)
-
Angiogenic signaling pathways and anti-angiogenic therapy for cancer
Signal Transduction and Targeted Therapy (2023)
-
Predictive value of an exosomal microRNA-based signature for tumor immunity in cervical cancer patients treated with chemoradiotherapy
Medical Molecular Morphology (2023)
-
Understanding crosstalk of organ tropism, tumor microenvironment and noncoding RNAs in breast cancer metastasis
Molecular Biology Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.