Abstract
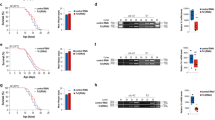
Chromatin modifiers regulate lifespan in several organisms, raising the question of whether changes in chromatin states in the parental generation could be incompletely reprogrammed in the next generation and thereby affect the lifespan of descendants. The histone H3 lysine 4 trimethylation (H3K4me3) complex, composed of ASH-2, WDR-5 and the histone methyltransferase SET-2, regulates Caenorhabditis elegans lifespan. Here we show that deficiencies in the H3K4me3 chromatin modifiers ASH-2, WDR-5 or SET-2 in the parental generation extend the lifespan of descendants up until the third generation. The transgenerational inheritance of lifespan extension by members of the ASH-2 complex is dependent on the H3K4me3 demethylase RBR-2, and requires the presence of a functioning germline in the descendants. Transgenerational inheritance of lifespan is specific for the H3K4me3 methylation complex and is associated with epigenetic changes in gene expression. Thus, manipulation of specific chromatin modifiers only in parents can induce an epigenetic memory of longevity in descendants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Cubas, P., Vincent, C. & Coen, E. An epigenetic mutation responsible for natural variation in floral symmetry. Nature 401, 157–161 (1999)
Brink, R. A. A genetic change associated with the R locus in maize which is directed and potentially reversible. Genetics 41, 872–889 (1956)
Woodhouse, M. R., Freeling, M. & Lisch, D. Initiation, establishment, and maintenance of heritable MuDR transposon silencing in maize are mediated by distinct factors. PLoS Biol. 4, e339 (2006)
Katz, D. J., Edwards, T. M., Reinke, V. & Kelly, W. G. A C. elegans LSD1 demethylase contributes to germline immortality by reprogramming epigenetic memory. Cell 137, 308–320 (2009)
Cavalli, G. & Paro, R. The Drosophila Fab-7 chromosomal element conveys epigenetic inheritance during mitosis and meiosis. Cell 93, 505–518 (1998)
Cavalli, G. & Paro, R. Epigenetic inheritance of active chromatin after removal of the main transactivator. Science 286, 955–958 (1999)
Seong, K. H., Li, D., Shimizu, H., Nakamura, R. & Ishii, S. Inheritance of stress-induced, ATF-2-dependent epigenetic change. Cell 145, 1049–1061 (2011)
Morgan, H. D., Sutherland, H. G., Martin, D. I. & Whitelaw, E. Epigenetic inheritance at the agouti locus in the mouse. Nature Genet. 23, 314–318 (1999)
Blewitt, M. E., Vickaryous, N. K., Paldi, A., Koseki, H. & Whitelaw, E. Dynamic reprogramming of DNA methylation at an epigenetically sensitive allele in mice. PLoS Genet. 2, e49 (2006)
Rassoulzadegan, M. et al. RNA-mediated non-mendelian inheritance of an epigenetic change in the mouse. Nature 441, 469–474 (2006)
Dang, W. et al. Histone H4 lysine 16 acetylation regulates cellular lifespan. Nature 459, 802–807 (2009)
Greer, E. L. et al. Members of the H3K4 trimethylation complex regulate lifespan in a germline-dependent manner in C. elegans . Nature 466, 383–387 (2010)
Siebold, A. P. et al. Polycomb Repressive Complex 2 and Trithorax modulate Drosophila longevity and stress resistance. Proc. Natl Acad. Sci. USA 107, 169–174 (2010)
McColl, G. et al. Pharmacogenetic analysis of lithium-induced delayed aging in Caenorhabditis elegans . J. Biol. Chem. 283, 350–357 (2008)
Chen, S. et al. The conserved NAD(H)-dependent corepressor CTBP-1 regulates Caenorhabditis elegans life span. Proc. Natl Acad. Sci. USA 106, 1496–1501 (2009)
Takahashi, Y. et al. Asymmetric arginine dimethylation determines life span in C. elegans by regulating forkhead transcription factor DAF-16. Cell Metab. 13, 505–516 (2011)
Maures, T. J., Greer, E. L., Hauswirth, A. G. & Brunet, A. The H3K27 demethylase UTX-1 regulates C. elegans lifespan in a germline-independent, insulin-dependent manner. Aging Cell 10.1111/j.1474-9726.2011.00738.x (11 August 2011)
Jin, C. et al. Histone demethylase UTX-1 regulates C. elegans life span by targeting the insulin/IGF-1 signaling pathway. Cell Metab. 14, 161–172 (2011)
Steward, M. M. et al. Molecular regulation of H3K4 trimethylation by ASH2L, a shared subunit of MLL complexes. Nature Struct. Mol. Biol. 13, 852–854 (2006)
Simonet, T., Dulermo, R., Schott, S. & Palladino, F. Antagonistic functions of SET-2/SET1 and HPL/HP1 proteins in C. elegans development. Dev. Biol. 312, 367–383 (2007)
Xiao, Y. et al. Caenorhabditis elegans chromatin-associated proteins SET-2 and ASH-2 are differentially required for histone H3 Lys 4 methylation in embryos and adult germ cells. Proc. Natl Acad. Sci. USA 108, 8305–8310 (2011)
Li, T. & Kelly, W. G. A role for Set1/MLL-related components in epigenetic regulation of the Caenorhabditis elegans germ line. PLoS Genet. 7, e1001349 (2011)
Dou, Y. et al. Regulation of MLL1 H3K4 methyltransferase activity by its core components. Nature Struct. Mol. Biol. 13, 713–719 (2006)
Haag, E. S., Wang, S. & Kimble, J. Rapid coevolution of the nematode sex-determining genes fem-3 and tra-2 . Curr. Biol. 12, 2035–2041 (2002)
Kawasaki, I. et al. PGL-1, a predicted RNA-binding component of germ granules, is essential for fertility in C. elegans . Cell 94, 635–645 (1998)
Fisher, K., Southall, S. M., Wilson, J. R. & Poulin, G. B. Methylation and demethylation activities of a C. elegans MLL-like complex attenuate RAS signalling. Dev. Biol. 341, 142–153 (2010)
Curran, S. P. & Ruvkun, G. Lifespan regulation by evolutionarily conserved genes essential for viability. PLoS Genet. 3, e56 (2007)
Hamilton, B. et al. A systematic RNAi screen for longevity genes in C. elegans . Genes Dev. 19, 1544–1555 (2005)
Kim, Y. & Sun, H. Functional genomic approach to identify novel genes involved in the regulation of oxidative stress resistance and animal lifespan. Aging Cell 6, 489–503 (2007)
Lee, S. S. et al. A systematic RNAi screen identifies a critical role for mitochondria in C. elegans longevity. Nature Genet. 33, 40–48 (2003)
Dillin, A. et al. Rates of behavior and aging specified by mitochondrial function during development. Science 298, 2398–2401 (2002)
Murphy, C. T. et al. Genes that act downstream of DAF-16 to influence the lifespan of Caenorhabditis elegans . Nature 424, 277–283 (2003)
Kenyon, C., Chang, J., Gensch, E., Rudner, A. & Tabtiang, R. A C. elegans mutant that lives twice as long as wild type. Nature 366, 461–464 (1993)
Liu, T. et al. Broad chromosomal domains of histone modification patterns in C. elegans . Genome Res. 21, 227–236 (2011)
Gerstein, M. B. et al. Integrative analysis of the Caenorhabditis elegans genome by the modENCODE project. Science 330, 1775–1787 (2010)
Spencer, W. C. et al. A spatial and temporal map of C. elegans gene expression. Genome Res. 21, 325–341 (2011)
Wu, S. C. & Zhang, Y. Active DNA demethylation: many roads lead to Rome. Nature Rev. Mol. Cell Biol. 11, 607–620 (2010)
Martin, C. & Zhang, Y. Mechanisms of epigenetic inheritance. Curr. Opin. Cell Biol. 19, 266–272 (2007)
Muramoto, T., Muller, I., Thomas, G., Melvin, A. & Chubb, J. R. Methylation of H3K4 Is required for inheritance of active transcriptional states. Curr. Biol. 20, 397–406 (2010)
Greer, E. L. et al. An AMPK-FOXO pathway mediates longevity induced by a novel method of dietary restriction in C. elegans . Curr. Biol. 17, 1646–1656 (2007)
Pferdehirt, R. R., Kruesi, W. S. & Meyer, B. J. An MLL/COMPASS subunit functions in the C. elegans dosage compensation complex to target X chromosomes for transcriptional regulation of gene expression. Genes Dev. 25, 499–515 (2011)
Tusher, V. G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl Acad. Sci. USA 98, 5116–5121 (2001)
Suzuki, R. & Shimodaira, H. Pvclust: an R package for assessing the uncertainty in hierarchical clustering. Bioinformatics 22, 1540–1542 (2006)
Pearson, K. On lines and planes of closest fit to systems of points in space. Phil. Mag. 6, 559–572 (1901)
Buck, M. J., Nobel, A. B. & Lieb, J. D. ChIPOTle: a user-friendly tool for the analysis of ChIP-chip data. Genome Biol. 6, R97 (2005)
Acknowledgements
We are grateful to J. Lieb, A. Rechtsteiner and S. Strome for sharing their ModENCODE data pre-publication and for helpful discussion. We thank K. Shen, M. W. Tan and T. Stiernagle and the Caenorhabditis Genetics Center for gifts of strains and reagents. We thank B. Meyer for her gift of the ASH-2 antibody. We thank A. Fire, S. Kim, J. Sage, S. Iwase, J. Lipsick, E. Pollina, A. Villeneuve and members of the Brunet lab for discussions and critical reading of the manuscript. We thank S. Han for screening different H3K4me3 antibodies for western blots in worm extracts. We thank R. Liefke and H. Tang for help with microarray analysis. This work was supported by NIH R01-AG31198 grant and by a generous gift from the Glenn Foundation for Medical Research to A.B.; E.L.G. was supported by an NSF graduate fellowship, by NIH ARRA-AG31198, by T32-CA009361, by a Helen Hay Whitney Post-Doctoral fellowship, and by a NIH R01-GM058012 (to Y.S.). T.J.M. was supported by NIH F32-AG037254. J.P.L. was supported by NIH T32-MH020016.
Author information
Authors and Affiliations
Contributions
E.L.G. conceived and planned the study with the help of A.B. E.L.G. performed the experiments and wrote the paper with the help of A.B.; E.L.G. performed some of the experiments in the lab of Y.S.; T.J.M. performed immunocytochemistry experiments (Fig. 6b and Supplementary Figs 6c and 7c); D.U. performed Pvclust and PCA microarray analysis (Fig. 6d, e and Supplementary Fig. 10a, b). A.G.H. helped with Figs 3b, c and 6a and Supplementary Figs 6a, b and 7a, b. E.M. performed an independent repeat of the transgenerational wdr-5 RNAi longevity experiments (Supplementary Table 2). J.P.L. helped with Fig. 3c and Supplementary Fig. 7a, b. B.A.B. helped with bioinformatics analysis (Supplementary Table 7). All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains a Supplementary Discussion, additional references, Supplementary Figures 1-10 with legends and Supplementary Tables 1-4. (PDF 12484 kb)
Supplementary Table 5
The table shows Normalized and log-transformed microarray gene expression values of all genes from L3 stage worms collected at the first day of egg-laying. (XLS 8204 kb)
Supplementary Table 6
The table shows normalized and log-transformed microarray gene expression values of all genes from L3 stage worms collected at the second day of egg-laying. (XLS 8204 kb)
Supplementary Table 7
The table shows lists of genes regulated by WDR-5, genes with transgenerational inheritance of expression, and genes with transgenerational inheritance of expression that are expressed in the germline. (XLS 404 kb)
Supplementary Table 8
The table shows GO enrichment for WDR-5 regulated genes. (XLS 17 kb)
Supplementary Table 9
The table shows normalized and log-transformed microarray gene expression values of WDR-5 regulated genes from L3 stage worms collected at the first day of egg- laying. (XLS 3213 kb)
Supplementary Table 10
The table shows normalized and log-transformed microarray gene expression values of WDR-5 regulated genes from L3 stage worms collected at the second day of egg-laying. (XLS 644 kb)
Rights and permissions
About this article
Cite this article
Greer, E., Maures, T., Ucar, D. et al. Transgenerational epigenetic inheritance of longevity in Caenorhabditis elegans. Nature 479, 365–371 (2011). https://doi.org/10.1038/nature10572
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10572
This article is cited by
-
Persistence of parental age effect on somatic mutation rates across generations in Arabidopsis
BMC Plant Biology (2023)
-
The CRTC-1 transcriptional domain is required for COMPASS complex-mediated longevity in C. elegans
Nature Aging (2023)
-
Tubular lysosome induction couples animal starvation to healthy aging
Nature Aging (2023)
-
Inheritance of associative memories and acquired cellular changes in C. elegans
Nature Communications (2023)
-
Endogenous DAF-16 spatiotemporal activity quantitatively predicts lifespan extension induced by dietary restriction
Communications Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.