Abstract
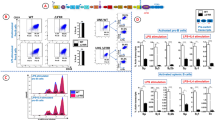
Immunoglobulin heavy chain (IgH) variable region exons are assembled from VH, D and JH gene segments in developing B lymphocytes. Within the 2.7-megabase mouse Igh locus, V(D)J recombination is regulated to ensure specific and diverse antibody repertoires. Here we report in mice a key Igh V(D)J recombination regulatory region, termed intergenic control region 1 (IGCR1), which lies between the VH and D clusters. Functionally, IGCR1 uses CTCF looping/insulator factor-binding elements and, correspondingly, mediates Igh loops containing distant enhancers. IGCR1 promotes normal B-cell development and balances antibody repertoires by inhibiting transcription and rearrangement of DH-proximal VH gene segments and promoting rearrangement of distal VH segments. IGCR1 maintains ordered and lineage-specific VH(D)JH recombination by suppressing VH joining to D segments not joined to JH segments, and VH to DJH joins in thymocytes, respectively. IGCR1 is also required for feedback regulation and allelic exclusion of proximal VH-to-DJH recombination. Our studies elucidate a long-sought Igh V(D)J recombination control region and indicate a new role for the generally expressed CTCF protein.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schatz, D. G. Antigen receptor genes and the evolution of a recombinase. Semin. Immunol. 16, 245–256 (2004)
Zhang, Y. et al. The role of mechanistic factors in promoting chromosomal translocations found in lymphoid and other cancers. Adv. Immunol. 106, 93–133 (2010)
Yancopoulos, G. D. et al. Preferential utilization of the most JH-proximal VH gene segments in pre-B-cell lines. Nature 311, 727–733 (1984)
Perlot, T. & Alt, F. W. Cis-regulatory elements and epigenetic changes control genomic rearrangements of the IgH locus. Adv. Immunol. 99, 1–32 (2008)
Alt, F. W. et al. Ordered rearrangement of immunoglobulin heavy chain variable region segments. EMBO J. 3, 1209–1219 (1984)
Jung, D., Giallourakis, C., Mostoslavsky, R. & Alt, F. W. Mechanism and control of V(D)J recombination at the immunoglobulin heavy chain locus. Annu. Rev. Immunol. 24, 541–570 (2006)
Bates, J. G., Cado, D., Nolla, H. & Schlissel, M. S. Chromosomal position of a V H gene segment determines its activation and inactivation as a substrate for V(D)J recombination. J. Exp. Med. 204, 3247–3256 (2007)
Fuxa, M. et al. Pax5 induces V-to-DJ rearrangements and locus contraction of the immunoglobulin heavy-chain gene. Genes Dev. 18, 411–422 (2004)
Reth, M. G., Ammirati, P., Jackson, S. & Alt, F. W. Regulated progression of a cultured pre-B-cell line to the B-cell stage. Nature 317, 353–355 (1985)
Melchers, F. et al. Repertoire selection by pre-B-cell receptors and B-cell receptors, and genetic control of B-cell development from immature to mature B cells. Immunol. Rev. 175, 33–46 (2000)
Malynn, B. A., Yancopoulos, G. D., Barth, J. E., Bona, C. A. & Alt, F. W. Biased expression of JH-proximal VH genes occurs in the newly generated repertoire of neonatal and adult mice. J. Exp. Med. 171, 843–859 (1990)
Decker, D. J., Boyle, N. E. & Klinman, N. R. Predominance of nonproductive rearrangements of VH81X gene segments evidences a dependence of B cell clonal maturation on the structure of nascent H chains. J. Immunol. 147, 1406–1411 (1991)
ten Boekel, E., Melchers, F. & Rolink, A. G. Changes in the VH gene repertoire of developing precursor B lymphocytes in mouse bone marrow mediated by the pre-B cell receptor. Immunity 7, 357–368 (1997)
Yancopoulos, G. D. & Alt, F. W. Developmentally controlled and tissue-specific expression of unrearranged VH gene segments. Cell 40, 271–281 (1985)
Yancopoulos, G. D., Blackwell, T. K., Suh, H., Hood, L. & Alt, F. W. Introduced T cell receptor variable region gene segments recombine in pre-B cells: evidence that B and T cells use a common recombinase. Cell 44, 251–259 (1986)
Corcoran, A. E. The epigenetic role of non-coding RNA transcription and nuclear organization in immunoglobulin repertoire generation. Semin. Immunol. 22, 353–361 (2010)
Feeney, A. Epigenetic regulation of V(D)J recombination. Semin. Immunol. 22, 311–312 (2010)
Subrahmanyam, R. & Sen, R. RAGs’ eye view of the immunoglobulin heavy chain gene locus. Semin. Immunol. 22, 337–345 (2010)
Jhunjhunwala, S., van Zelm, M. C., Peak, M. M. & Murre, C. Chromatin architecture and the generation of antigen receptor diversity. Cell 138, 435–448 (2009)
Abarrategui, I. & Krangel, M. S. Germline transcription: a key regulator of accessibility and recombination. Adv. Exp. Med. Biol. 650, 93–102 (2009)
Bergman, Y. & Cedar, H. Epigenetic control of recombination in the immune system. Semin. Immunol. 22, 323–329 (2010)
Sayegh, C. E., Jhunjhunwala, S., Riblet, R. & Murre, C. Visualization of looping involving the immunoglobulin heavy-chain locus in developing B cells. Genes Dev. 19, 322–327 (2005)
Jhunjhunwala, S. et al. The 3D structure of the immunoglobulin heavy-chain locus: implications for long-range genomic interactions. Cell 133, 265–279 (2008)
Roldán, E. et al. Locus ‘decontraction’ and centromeric recruitment contribute to allelic exclusion of the immunoglobulin heavy-chain gene. Nature Immunol. 6, 31–41 (2004)
Pinaud, E. et al. The IgH locus 3′ regulatory region: pulling the strings from behind. Adv. Immunol. 110, 27–70 (2011)
Sakai, E., Bottaro, A., Davidson, L., Sleckman, B. P. & Alt, F. W. Recombination and transcription of the endogenous Ig heavy chain locus is effected by the Ig heavy chain intronic enhancer core region in the absence of the matrix attachment regions. Proc. Natl Acad. Sci. USA 96, 1526–1531 (1999)
Perlot, T., Alt, F. W., Bassing, C. H., Suh, H. & Pinaud, E. Elucidation of IgH intronic enhancer functions via germ-line deletion. Proc. Natl Acad. Sci. USA 102, 14362–14367 (2005)
Afshar, R., Pierce, S., Bolland, D. J., Corcoran, A. & Oltz, E. M. Regulation of IgH gene assembly: role of the intronic enhancer and 5′DQ52 region in targeting DHJH recombination. J. Immunol. 176, 2439–2447 (2006)
Featherstone, K., Wood, A. L., Bowen, A. J. & Corcoran, A. E. The mouse immunoglobulin heavy chain V-D intergenic sequence contains insulators that may regulate ordered V(D)J recombination. J. Biol. Chem. 285, 9327–9338 (2010)
Giallourakis, C. C. et al. Elements between the IgH variable (V) and diversity (D) clusters influence antisense transcription and lineage-specific V(D)J recombination. Proc. Natl Acad. Sci. USA 107, 22207–22212 (2010)
Lin, Y. C. et al. A global network of transcription factors, involving E2A, EBF1 and Foxo1, that orchestrates B cell fate. Nature Immunol. 11, 635–643 (2010)
Ebert, A. et al. The distal VH gene cluster of the Igh locus contains distinct regulatory elements with pax5 transcription factor-dependent activity in pro-B cells. Immunity 34, 175–187 (2011)
Degner, S. C., Wong, T. P., Jankevicius, G. & Feeney, A. J. Cutting edge: developmental stage-specific recruitment of cohesin to CTCF sites throughout immunoglobulin loci during B lymphocyte development. J. Immunol. 182, 44–48 (2009)
Williams, A. & Flavell, R. A. The role of CTCF in regulating nuclear organization. J. Exp. Med. 205, 747–750 (2008)
Phillips, J. E. & Corces, V. G. CTCF: master weaver of the genome. Cell 137, 1194–1211 (2009)
Bulger, M. & Groudine, M. Enhancers: the abundance and function of regulatory sequences beyond promoters. Dev. Biol. 339, 250–257 (2010)
Degner, S. C. et al. CCCTC-binding factor (CTCF) and cohesin influence the genomic architecture of the Igh locus and antisense transcription in pro-B cells. Proc. Natl Acad. Sci. USA 108, 9566–9571 (2011)
Wuerffel, R. et al. S-S synapsis during class switch recombination is promoted by distantly located transcriptional elements and activation-induced deaminase. Immunity 27, 711–722 (2007)
Hirano, S. L. et al. Identity of IGHV-7183.1 (V81x) coding and recombination signal sequences among wild-derived mice. Immunogenetics 53, 54–58 (2001)
Hardy, R. R. & Hayakawa, K. B cell development pathways. Annu. Rev. Immunol. 19, 595–621 (2001)
Hesslein, D. G. et al. Pax5 is required for recombination of transcribed, acetylated, 5′ IgH V gene segments. Genes Dev. 17, 37–42 (2003)
Su, I. H. et al. Ezh2 controls B cell development through histone H3 methylation and Igh rearrangement. Nature Immunol. 4, 124–131 (2003)
Liu, H. et al. Yin Yang 1 is a critical regulator of B-cell development. Genes Dev. 21, 1179–1189 (2007)
Ji, Y. et al. The in vivo pattern of binding of RAG1 and RAG2 to antigen receptor loci. Cell 141, 419–431 (2010)
Degner-Leisso, S. C. & Feeney, A. J. Epigenetic and 3-dimensional regulation of V(D)J rearrangement of immunoglobulin genes. Semin. Immunol. 22, 346–352 (2010)
MacPherson, M. J. & Sadowski, P. D. The CTCF insulator protein forms an unusual DNA structure. BMC Mol. Biol. 11, 101 (2010)
Kyrchanova, O., Chetverina, D., Maksimenko, O., Kullyev, A. & Georgiev, P. Orientation-dependent interaction between Drosophila insulators is a property of this class of regulatory elements. Nucleic Acids Res. 36, 7019–7028 (2008)
Dudley, D. D. et al. Impaired V(D)J recombination and lymphocyte development in core RAG1-expressing mice. J. Exp. Med. 198, 1439–1450 (2003)
Hagège, H. et al. Quantitative analysis of chromosome conformation capture assays (3C-qPCR). Nature Protocols 2, 1722–1733 (2007)
Chen, J. et al. Mutations of the intronic IgH enhancer and its flanking sequences differentially affect accessibility of the JH locus. EMBO J. 12, 4635–4645 (1993)
Barreto, V. & Cumano, A. Frequency and characterization of phenotypic Ig heavy chain allelically included IgM-expressing B cells in mice. J. Immunol. 164, 893–899 (2000)
Sonoda, E. et al. B cell development under the condition of allelic inclusion. Immunity 6, 225–233 (1997)
McManus, S. et al. The transcription factor Pax5 regulates its target genes by recruiting chromatin-modifying proteins in committed B cells. EMBO J. 30, 2388–2404 (2011)
Acknowledgements
We thank Y. Fujiwara and P.-Y. Huang for generating chimaeric mice. This work was supported by NIH grants RO1 AI20047 (to F.W.A.), RO1 HL48702 and AI40227 (to M.S.S.), CA054198-20 (to C.M.) and K08 AI070839 (to C.C.G.). M.B. was supported by the Austrian GEN-AU initiative and Boehringer Ingelheim. C.G. is supported by an Irvington Institute Postdoctoral Fellowship from the Cancer Research Institute. C.V. is supported by a Marie Curie Fellowship. C.B. is supported by an EMBO fellowship. F.W.A. is an Investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.G., H.S.Y., A.F., C.C.G. and F.W.A. conceived, designed and/or peformed most experiments, interpreted most results, and wrote the manuscript. With respect to other authors, S.J. performed experiments for Figs 2, 3, 4 and Supplementary Fig. 11 and contributed to interpretation of the data; A.E. and M.B. contributed the work in Fig. 2c and Supplementary 7a–c. C.V., J.G.B. and M.S.S. contributed the work in Supplementary Fig. 10; and C.B. and C.M. performed FISH experiments on IGCR1−/− cells that helped frame aspects of the discussion and models; all of these authors contributed to polishing the manuscript. All other authors provided technical assistance with various experiments or data analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-14 with legends, Supplementary Tables 1-6, a Supplementary Discussion and additional references. (PDF 8762 kb)
Rights and permissions
About this article
Cite this article
Guo, C., Yoon, H., Franklin, A. et al. CTCF-binding elements mediate control of V(D)J recombination. Nature 477, 424–430 (2011). https://doi.org/10.1038/nature10495
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10495
This article is cited by
-
Boundary stacking interactions enable cross-TAD enhancer–promoter communication during limb development
Nature Genetics (2024)
-
An Igh distal enhancer modulates antigen receptor diversity by determining locus conformation
Nature Communications (2023)
-
Genetic variation in the immunoglobulin heavy chain locus shapes the human antibody repertoire
Nature Communications (2023)
-
Genome control by SMC complexes
Nature Reviews Molecular Cell Biology (2023)
-
The role of chromatin loop extrusion in antibody diversification
Nature Reviews Immunology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.