Abstract
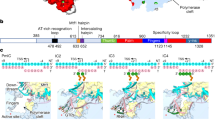
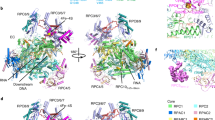
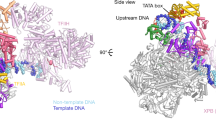
Transcription of the mitochondrial genome is performed by a single-subunit RNA polymerase (mtRNAP) that is distantly related to the RNAP of bacteriophage T7, the pol I family of DNA polymerases, and single-subunit RNAPs from chloroplasts1,2,3,4. Whereas T7 RNAP can initiate transcription by itself, mtRNAP requires the factors TFAM and TFB2M for binding and melting promoter DNA5,6,7. TFAM is an abundant protein that binds and bends promoter DNA 15–40 base pairs upstream of the transcription start site, and stimulates the recruitment of mtRNAP and TFB2M to the promoter8,9. TFB2M assists mtRNAP in promoter melting and reaches the active site of mtRNAP to interact with the first base pair of the RNA–DNA hybrid10. Here we report the X-ray structure of human mtRNAP at 2.5 Å resolution, which reveals a T7-like catalytic carboxy-terminal domain, an amino-terminal domain that remotely resembles the T7 promoter-binding domain, a novel pentatricopeptide repeat domain, and a flexible N-terminal extension. The pentatricopeptide repeat domain sequesters an AT-rich recognition loop, which binds promoter DNA in T7 RNAP, probably explaining the need for TFAM during promoter binding. Consistent with this, substitution of a conserved arginine residue in the AT-rich recognition loop, or release of this loop by deletion of the N-terminal part of mtRNAP, had no effect on transcription. The fingers domain and the intercalating hairpin, which melts DNA in phage RNAPs, are repositioned, explaining the need for TFB2M during promoter melting. Our results provide a new venue for the mechanistic analysis of mitochondrial transcription. They also indicate how an early phage-like mtRNAP lost functions in promoter binding and melting, which were provided by initiation factors in trans during evolution, to enable mitochondrial gene regulation and the adaptation of mitochondrial function to changes in the environment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Masters, B. S., Stohl, L. L. & Clayton, D. A. Yeast mitochondrial RNA polymerase is homologous to those encoded by bacteriophages T3 and T7. Cell 51, 89–99 (1987)
Cheetham, G. M. & Steitz, T. A. Structure of a transcribing T7 RNA polymerase initiation complex. Science 286, 2305–2309 (1999)
Gaspari, M., Larsson, N. G. & Gustafsson, C. M. The transcription machinery in mammalian mitochondria. Biochim. Biophys. Acta 1659, 148–152 (2004)
Kohlstaedt, L. A., Wang, J., Friedman, J. M., Rice, P. A. & Steitz, T. A. Crystal structure at 3.5 A resolution of HIV-1 reverse transcriptase complexed with an inhibitor. Science 256, 1783–1790 (1992)
Litonin, D. et al. Human mitochondrial transcription revisited: only TFAM and TFB2M are required for transcription of the mitochondrial genes in vitro . J. Biol. Chem. 285, 18129–18133 (2010)
Falkenberg, M. et al. Mitochondrial transcription factors B1 and B2 activate transcription of human mtDNA. Nature Genet. 31, 289–294 (2002)
Gaspari, M., Falkenberg, M., Larsson, N. G. & Gustafsson, C. M. The mitochondrial RNA polymerase contributes critically to promoter specificity in mammalian cells. EMBO J. 23, 4606–4614 (2004)
Ekstrand, M. I. et al. Mitochondrial transcription factor A regulates mtDNA copy number in mammals. Hum. Mol. Genet. 13, 935–944 (2004)
Dairaghi, D. J., Shadel, G. S. & Clayton, D. A. Human mitochondrial transcription factor A and promoter spacing integrity are required for transcription initiation. Biochim. Biophys. Acta 1271, 127–134 (1995)
Sologub, M., Litonin, D., Anikin, M., Mustaev, A. & Temiakov, D. TFB2 is a transient component of the catalytic site of the human mitochondrial RNA polymerase. Cell 139, 934–944 (2009)
Durniak, K. J., Bailey, S. & Steitz, T. A. The structure of a transcribing T7 RNA polymerase in transition from initiation to elongation. Science 322, 553–557 (2008)
Yin, Y. W. & Steitz, T. A. The structural mechanism of translocation and helicase activity in T7 RNA polymerase. Cell 116, 393–404 (2004)
Temiakov, D. et al. Structural basis for substrate selection by T7 RNA polymerase. Cell 116, 381–391 (2004)
Tahirov, T. H. et al. Structure of a T7 RNA polymerase elongation complex at 2.9 Å resolution. Nature 420, 43–50 (2002)
Yin, Y. W. & Steitz, T. A. Structural basis for the transition from initiation to elongation transcription in T7 RNA polymerase. Science 298, 1387–1395 (2002)
Rodeheffer, M. S., Boone, B. E., Bryan, A. C. & Shadel, G. S. Nam1p, a protein involved in RNA processing and translation, is coupled to transcription through an interaction with yeast mitochondrial RNA polymerase. J. Biol. Chem. 276, 8616–8622 (2001)
Small, I. D. & Peeters, N. The PPR motif—a TPR-related motif prevalent in plant organellar proteins. Trends Biochem. Sci. 25, 46–47 (2000)
Lightowlers, R. N. & Chrzanowska-Lightowlers, Z. M. PPR (pentatricopeptide repeat) proteins in mammals: important aids to mitochondrial gene expression. Biochem. J. 416, e5–e6 (2008)
Hammani, K. et al. The pentatricopeptide repeat protein OTP87 is essential for RNA editing of nad7 and atp1 transcripts in Arabidopsis mitochondria. J. Biol. Chem. 286, 21361–21371 (2011)
Aphasizheva, I., Maslov, D., Wang, X., Huang, L. & Aphasizhev, R. Pentatricopeptide repeat proteins stimulate mRNA adenylation/uridylation to activate mitochondrial translation in trypanosomes. Mol. Cell 42, 106–117 (2011)
Cheetham, G. M., Jeruzalmi, D. & Steitz, T. A. Structural basis for initiation of transcription from an RNA polymerase–promoter complex. Nature 399, 80–83 (1999)
Gleghorn, M. L., Davydova, E. K., Rothman-Denes, L. B. & Murakami, K. S. Structural basis for DNA-hairpin promoter recognition by the bacteriophage N4 virion RNA polymerase. Mol. Cell 32, 707–717 (2008)
Nayak, D., Guo, Q. & Sousa, R. A promoter recognition mechanism common to yeast mitochondrial and phage T7 RNA polymerases. J. Biol. Chem. 284, 13641–13647 (2009)
Schubot, F. D. et al. Crystal structure of the transcription factor sc-mtTFB offers insights into mitochondrial transcription. Protein Sci. 10, 1980–1988 (2001)
Mangus, D. A., Jang, S. H. & Jaehning, J. A. Release of the yeast mitochondrial RNA polymerase specificity factor from transcription complexes. J. Biol. Chem. 269, 26568–26574 (1994)
Cliften, P. F., Park, J. Y., Davis, B. P., Jang, S. H. & Jaehning, J. A. Identification of three regions essential for interaction between a sigma-like factor and core RNA polymerase. Genes Dev. 11, 2897–2909 (1997)
Savkina, M., Temiakov, D., McAllister, W. T. & Anikin, M. Multiple functions of yeast mitochondrial transcription factor Mtf1p during initiation. J. Biol. Chem. 285, 3957–3964 (2010)
Leslie, A. G. W. Joint CCP4 and ESF-EACMB Newsletter Protein Crystallogr. No. 26 (Daresbury Laboratory, 1992)
Evans, P. Scaling and assessment of data quality. Acta Crystallogr. D Biol. Crystallogr. 62, 72–82 (2006)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997)
Acknowledgements
We thank A. Cheung and D. Kostrewa for help with crystallography, and W. T. McAllister and M. Anikin for the critical reading of the manuscript. We acknowledge the crystallization facility at the Max Planck Institute of Biochemistry, Martinsried. Part of this work was performed at the Swiss Light Source at the Paul Scherrer Institut, Villigen, Switzerland. Use of the Advanced Photon Source at Argonne National Laboratory was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under contract no. DE-AC02-06CH11357. Use of the Lilly Research Laboratories Collaborative Access Team (LRL-CAT) beamline at Sector 31 of the Advanced Photon Source was provided by Eli Lilly Company, which operates the facility. P.C. was supported by the Deutsche Forschungsgemeinschaft, SFB646, TR5, FOR1068, the Nanosystems Initiative Munich NIM, an Advanced Investigator Grant from the European Research Council ERC, and the Jung-Stiftung. D.T. was supported by the UMDNJ Foundation, grant no. PC25-11.
Author information
Authors and Affiliations
Contributions
M.S. and D.L. cloned mtRNAP variants. M.S., D.L., D.T. and Y.I.M. performed RNAP purification and biochemical assays. R.R. and D.T. prepared the crystals. R.R. performed structure determination and modelling. P.C. and D.T. designed and supervised the project and prepared the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-4 with legends, Supplementary Table 1 and Supplementary References. (PDF 4695 kb)
Rights and permissions
About this article
Cite this article
Ringel, R., Sologub, M., Morozov, Y. et al. Structure of human mitochondrial RNA polymerase. Nature 478, 269–273 (2011). https://doi.org/10.1038/nature10435
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10435
This article is cited by
-
Mechanisms and regulation of human mitochondrial transcription
Nature Reviews Molecular Cell Biology (2024)
-
N-Acryloylindole-alkyne (NAIA) enables imaging and profiling new ligandable cysteines and oxidized thiols by chemoproteomics
Nature Communications (2023)
-
Mitochondrien: wie die Gene im Kraftwerk der Zelle aktiviert werden
BIOspektrum (2022)
-
POLRMT mutations impair mitochondrial transcription causing neurological disease
Nature Communications (2021)
-
Small-molecule inhibitors of human mitochondrial DNA transcription
Nature (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.