Abstract
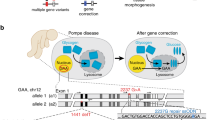
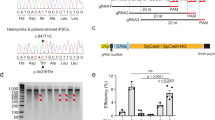
Human induced pluripotent stem cells (iPSCs) represent a unique opportunity for regenerative medicine because they offer the prospect of generating unlimited quantities of cells for autologous transplantation, with potential application in treatments for a broad range of disorders1,2,3,4. However, the use of human iPSCs in the context of genetically inherited human disease will require the correction of disease-causing mutations in a manner that is fully compatible with clinical applications3,5. The methods currently available, such as homologous recombination, lack the necessary efficiency and also leave residual sequences in the targeted genome6. Therefore, the development of new approaches to edit the mammalian genome is a prerequisite to delivering the clinical promise of human iPSCs. Here we show that a combination of zinc finger nucleases (ZFNs)7 and piggyBac8,9 technology in human iPSCs can achieve biallelic correction of a point mutation (Glu342Lys) in the α1-antitrypsin (A1AT, also known as SERPINA1) gene that is responsible for α1-antitrypsin deficiency. Genetic correction of human iPSCs restored the structure and function of A1AT in subsequently derived liver cells in vitro and in vivo. This approach is significantly more efficient than any other gene-targeting technology that is currently available and crucially prevents contamination of the host genome with residual non-human sequences. Our results provide the first proof of principle, to our knowledge, for the potential of combining human iPSCs with genetic correction to generate clinically relevant cells for autologous cell-based therapies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Gene Expression Omnibus
Data deposits
Exome sequence data have been deposited at the European Genome-Phenome Archive (http://www.ebi.ac.uk/ega/) hosted by the European Bioinformatics Institute under accession EGAS00001000055. CGH and SNP array data have been deposited with EBI ArrayExpress (http://www.ebi.ac.uk/arrayexpress/) under accession number E-MEXP-3316 and with Gene Expression Omnibus (http://www.ncbi.nlm.nih.gov/geo/) under accession number GSE31035, respectively.
References
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007)
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007)
Stadtfeld, M. & Hochedlinger, K. Induced pluripotency: history, mechanisms, and applications. Genes Dev. 24, 2239–2263 (2010)
Hanna, J. et al. Treatment of sickle cell anemia mouse model with iPS cells generated from autologous skin. Science 318, 1920–1923 (2007)
Fairchild, P. J. The challenge of immunogenicity in the quest for induced pluripotency. Nature Rev. Immunol. 10, 868–875 (2010)
Tenzen, T., Zembowicz, F. & Cowan, C. A. Genome modification in human embryonic stem cells. J. Cell. Physiol. 222, 278–281 (2010)
Urnov, F. D., Rebar, E. J., Holmes, M. C., Zhang, H. S. & Gregory, P. D. Genome editing with engineered zinc finger nucleases. Nature Rev. Genet. 11, 636–646 (2010)
Yusa, K., Zhou, L., Li, M. A., Bradley, A. & Craig, N. L. A hyperactive piggyBac transposase for mammalian applications. Proc. Natl Acad. Sci. USA 108, 1531–1536 (2011)
Wang, W. et al. Chromosomal transposition of PiggyBac in mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 105, 9290–9295 (2008)
Hockemeyer, D. et al. Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nature Biotechnol. 27, 851–857 (2009)
van der Weyden, L., Adams, D. J. & Bradley, A. Tools for targeted manipulation of the mouse genome. Physiol. Genomics 11, 133–164 (2002)
Meier, I. D. et al. Short DNA sequences inserted for gene targeting can accidentally interfere with off-target gene expression. FASEB J. 24, 1714–1724 (2010)
Lacoste, A., Berenshteyn, F. & Brivanlou, A. H. An efficient and reversible transposable system for gene delivery and lineage-specific differentiation in human embryonic stem cells. Cell Stem Cell 5, 332–342 (2009)
Fraser, M. J., Ciszczon, T., Elick, T. & Bauser, C. Precise excision of TTAA-specific lepidopteran transposons piggyBac (IFP2) and tagalong (TFP3) from the baculovirus genome in cell lines from two species of Lepidoptera. Insect Mol. Biol. 5, 141–151 (1996)
Yusa, K., Rad, R., Takeda, J. & Bradley, A. Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon. Nature Methods 6, 363–369 (2009)
Rashid, S. T. et al. Modeling inherited metabolic disorders of the liver using human induced pluripotent stem cells. J. Clin. Invest. 120, 3127–3136 (2010)
Perlmutter, D. H. Autophagic disposal of the aggregation-prone protein that causes liver inflammation and carcinogenesis in α-1-antitrypsin deficiency. Cell Death Differ. 16, 39–45 (2009)
Gooptu, B. & Lomas, D. A. Conformational pathology of the serpins: themes, variations, and therapeutic strategies. Annu. Rev. Biochem. 78, 147–176 (2009)
Zou, J. et al. Gene targeting of a disease-related gene in human induced pluripotent stem and embryonic stem cells. Cell Stem Cell 5, 97–110 (2009)
Mitalipova, M. M. et al. Preserving the genetic integrity of human embryonic stem cells. Nature Biotechnol. 23, 19–20 (2005)
Baker, D. E. et al. Adaptation to culture of human embryonic stem cells and oncogenesis in vivo . Nature Biotechnol. 25, 207–215 (2007)
Lefort, N. et al. Human embryonic stem cells reveal recurrent genomic instability at 20q11.21. Nature Biotechnol. 26, 1364–1366 (2008)
Spits, C. et al. Recurrent chromosomal abnormalities in human embryonic stem cells. Nature Biotechnol. 26, 1361–1363 (2008)
Touboul, T. et al. Generation of functional hepatocytes from human embryonic stem cells under chemically defined conditions that recapitulate liver development. Hepatology 51, 1754–1765 (2010)
Fusaki, N., Ban, H., Nishiyama, A., Saeki, K. & Hasegawa, M. Efficient induction of transgene-free human pluripotent stem cells using a vector based on Sendai virus, an RNA virus that does not integrate into the host genome. Proc. Jpn. Acad., Ser. B, Phys. Biol. Sci. 85, 348–362 (2009)
Watanabe, K. et al. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nature Biotechnol. 25, 681–686 (2007)
Cadinanos, J. & Bradley, A. Generation of an inducible and optimized piggyBac transposon system. Nucleic Acids Res. 35, e87 (2007)
Skarnes, W. C. et al. A conditional knockout resource for the genome-wide study of mouse gene function. Nature 474, 337–342 (2011)
Liu, P., Jenkins, N. A. & Copeland, N. G. A highly efficient recombineering-based method for generating conditional knockout mutations. Genome Res. 13, 476–484 (2003)
Urnov, F. D. et al. Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435, 646–651 (2005)
Beerli, R. R. & Barbas, C. F. III Engineering polydactyl zinc-finger transcription factors. Nature Biotechnol. 20, 135–141 (2002)
Pavletich, N. P. & Pabo, C. O. Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 Å. Science 252, 809–817 (1991)
Guschin, D. Y. et al. A rapid and general assay for monitoring endogenous gene modification. Methods Mol. Biol. 649, 247–256 (2010)
Varela, I. et al. Exome sequencing identifies frequent mutation of the SWI/SNF complex gene PBRM1 in renal carcinoma. Nature 469, 539–542 (2011)
The 1000 Genomes Project Consortium . A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010)
Seki, T. et al. Generation of induced pluripotent stem cells from human terminally differentiated circulating T cells. Cell Stem Cell 7, 11–14 (2010)
Acknowledgements
We thank A. Klug and M. Minczuk for their advice, M. A. Li for comments on the manuscript, P. Ellis, N. Hammond and C. McGee for CGH analysis, the Sanger Institute sequencing facility for exome sequencing, N. Conte and S. Rice for assistance with bioinformatic analysis, M. Alexander for her help with cell culture reagents. We also thank L. Zhang, S. Hinkley and the production group for ZFN assembly and validation, K. Tong and X. Meng for technical assistance, J. C. Miller and E. Leung for ZFN off-target site analysis and S. Abrahamson and P. D. Gregory for careful reading of the manuscript. This work was supported by the Wellcome Trust (WT077187; A.B.), the MRC Senior non-clinical fellowship and the Cambridge Hospitals National Institute for Health Research Biomedical Research Center (L.V.), the Medical Research Council and Papworth NHS Trust (D.A.L.), the Bill and Melinda Gates Foundation, Inserm and Institut Pasteur (H.S.-M.) and Japan Science and Technology Agency (N.F.). K.Y. is supported by a postdoctoral fellowship of Japan Society for the Promotion of Science. S.T.R. and F.J.R. are Wellcome Trust Clinical Training Fellows. I.V. is supported by a fellowship from the International Human Frontiers Science Program Organization.
Author information
Authors and Affiliations
Contributions
K.Y. and S.T.R. are joint first authors. D.A.L., A.B. and L.V. contributed equally to this work. K.Y., S.T.R., D.A.L., A.B. and L.V. conceived the research and wrote the manuscript with comments from all authors. K.Y. performed gene correction in mouse and human iPSCs and conducted all experiments using piggyBac in Cambridge, UK. S.T.R., E.M., A.O., N.R.F.H., F.J.R., G.A. and S.J.M. performed in vitro phenotypic analysis of corrected human iPSCs. S.T.R., H.S.-M., S.D. and J.P.D.S. performed in vivo work. I.V. performed data analysis of exome sequencing. P.Q.-L., D.E.P. and M.C.H. generated and validated ZFNs. N.F. and M.H. generated Sendai virus vectors.
Corresponding authors
Ethics declarations
Competing interests
[Competing financial interests: P.-Q.L., D.E.P. and M.C.H. are employees of Sangamo BioSciences. N.F. and M.H. are employees of DNAVEC. All other authors declare no competing financial interests.]
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-8 with legends, Supplementary Tables 1-5, Supplementary Text and additional references. (PDF 1907 kb)
Rights and permissions
About this article
Cite this article
Yusa, K., Rashid, S., Strick-Marchand, H. et al. Targeted gene correction of α1-antitrypsin deficiency in induced pluripotent stem cells. Nature 478, 391–394 (2011). https://doi.org/10.1038/nature10424
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10424
This article is cited by
-
Stem cell therapy for hepatocellular carcinoma and end-stage liver disease
Journal of the Egyptian National Cancer Institute (2023)
-
CRISPR-Cas9 genetic screen leads to the discovery of L-Moses, a KAT2B inhibitor that attenuates Tunicamycin-mediated neuronal cell death
Scientific Reports (2023)
-
Mutant huntingtin confers cell-autonomous phenotypes on Huntington’s disease iPSC-derived microglia
Scientific Reports (2023)
-
Cas9-induced large deletions and small indels are controlled in a convergent fashion
Nature Communications (2022)
-
Reconstitution of the oocyte transcriptional network with transcription factors
Nature (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.