Abstract
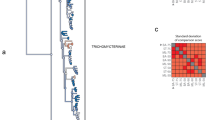
Evolutionary relationships among the eight major lineages of Mollusca have remained unresolved despite their diversity and importance. Previous investigations of molluscan phylogeny, based primarily on nuclear ribosomal gene sequences1,2,3 or morphological data4, have been unsuccessful at elucidating these relationships. Recently, phylogenomic studies using dozens to hundreds of genes have greatly improved our understanding of deep animal relationships5. However, limited genomic resources spanning molluscan diversity has prevented use of a phylogenomic approach. Here we use transcriptome and genome data from all major lineages (except Monoplacophora) and recover a well-supported topology for Mollusca. Our results strongly support the Aculifera hypothesis placing Polyplacophora (chitons) in a clade with a monophyletic Aplacophora (worm-like molluscs). Additionally, within Conchifera, a sister-taxon relationship between Gastropoda and Bivalvia is supported. This grouping has received little consideration and contains most (>95%) molluscan species. Thus we propose the node-based name Pleistomollusca. In light of these results, we examined the evolution of morphological characters and found support for advanced cephalization and shells as possibly having multiple origins within Mollusca.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
Capillary sequence data are available from the NCBI EST database (http://www.ncbi.nlm.nih.gov/projects/dbEST) under accession numbers JG454968.1–JG456874.1 and 454 sequence data are available from the NCBI SRA database (http://www.ncbi.nlm.nih.gov/sra) accession number SRA030407.1. Matrices and trees from this study are available from TreeBASE (http://www.treebase.org) accession number S11762.
References
Passamaneck, Y. J., Schander, C. & Halanych, K. M. Investigation of molluscan phylogeny using large-subunit and small-subunit nuclear rRNA sequences. Mol. Phylogenet. Evol. 32, 25–38 (2004)
Giribet, G. et al. Evidence for a clade composed of molluscs with serially repeated structures: Monoplacophorans are related to chitons. Proc. Natl. Acad. Sci. USA 103, 7723–7728 (2006)
Wilson, N. G., Rouse, G. W. & Giribet, G. Assessing the molluscan hypothesis Serialia (Monoplacophora+ Polyplacophora) using novel molecular data. Mol. Phylogenet. Evol. 54, 187–193 (2010)
Haszprunar, G. Is the Aplacophora monophyletic? A cladistic point of view. Am. Malacol. Bull. 15, 115–130 (2000)
Dunn, C. W. et al. Broad phylogenomic sampling improves resolution of the animal tree of life. Nature 452, 745–749 (2008)
Haszprunar, G., Schander, C. & Halanych, K. M. In Phylogeny and Evolution of the Mollusca (eds Ponder, W. & Lindberg, D. R. ) 19–32 (Univ. of California Press, 2008)
Todt, C., Okusu, A., Schander, C. & Schwabe, E. In Phylogeny and evolution of the Mollusca (eds Ponder, W. & Lindberg, D. R. ) 105–141 (Univ. of California Press, 2008)
Scheltema, A. H. Aplacophora as progenetic aculiferans and the coelomate origin of mollusks as the sister taxon of Sipuncula. Biol. Bull. 184, 57–78 (1993)
Salvini-Plawen, L. On the phylogenetic significance of the aplacophoran Mollusca. Iberus 21, 67–97 (2003)
Meyer, A., Todt, C., Mikkelson, N. & Lieb, B. Fast evolving 18S rRNA sequences from Solenogastres (Mollusca) resist standard PCR amplification and give new insights into mollusk substitution rate heterogeneity. BMC Evol. Biol. 10, 70 (2010)
Struck, T. H. et al. Phylogenomic analyses unravel annelid evolution. Nature 471, 95–98 (2011)
Sigwart, J. D. & Sutton, M. D. Deep molluscan phylogeny: synthesis of palaeontological and neontological data. Proc. R. Soc. B 274, 2413–2419 (2007)
Scheltema, A. H. & Ivanov, D. L. An aplacophoran postlarva with iterated dorsal groups of spicules and skeletal similarities to Paleozoic fossils. Invertebr. Biol. 121, 1–10 (2002)
Nielsen, C., Haszprunar, G., Ruthensteiner, B. & Wanninger, A. Early development of the aplacophoran mollusc Chaetoderma. Acta Zool. 88, 231–247 (2007)
Todt, C. & Wanninger, A. Of tests, trochs, shells, and spicules: Development of the basal mollusk Wirenia argentea (Solenogastres) and its bearing on the evolution of trochozoan larval key features. Front. Zool. 7, 6 (2010)
Scheltema, A. H. & Schander, C. Exoskeletons: tracing molluscan evolution. Venus 65, 19–26 (2006)
Meyer, A., Witek, A. & Lieb, B. Selecting ribosomal protein genes for invertebrate phylogenetic inferences: how many genes to resolve the Mollusca? Method. Ecol. Evol. 2, 34–42 (2011)
Wanninger, A. & Haszprunar, G. Muscle development in Antalis entalis (Mollusca, Scaphopoda) and its significance for scaphopod relationships. J. Morphol. 254, 53–64 (2002)
Lundin, K., Schander, C. & Todt, C. Ultrastructure of epidermal cilia and ciliary rootlets in Scaphopoda. J. Molluscan Stud. 75, 69–73 (2008)
Moroz, L. L. On the independent origins of complex brains and neurons. Brain Behav. Evol. 74, 177–190 (2009)
Simone, L. R. L. Filogenia das superfamílias de Caenogastropoda (Mollusca) com base em morfologia comparativa. PhD thesis, Univ. São Paulo. (2000)
Jörger, K. M. et al. On the origin of Acochlidia and other enigmatic euthyneuran gastropods, with implications for the systematics of Heterobranchia. BMC Evol. Biol. 10, 323 (2010)
Caron, J. B., Scheltema, A., Schander, C. & Rudkin, D. A soft-bodied mollusc with radula from the Middle Cambrian Burgess Shale. Nature 442, 159–163 (2006)
Scheltema, A. H., Kerth, K. & Kuzirian, A. M. Original molluscan radula: comparisons among Aplacophora, Polyplacophora, Gastropoda, and the Cambrian fossil Wiwaxia corrugata. J. Morphol. 257, 219–245 (2003)
Forment, J. et al. EST2uni: an open, parallel tool for automated EST analysis and database creation, with a data mining web interface and microarray expression data integration. BMC Bioinformatics 9, 5 (2008)
Ebersberger, I., Strauss, S. & Von Haeseler, A. HaMStR: Profile hidden markov model based search for orthologs in ESTs. BMC Evol. Biol. 9, 157 (2009)
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690 (2006)
Lartillot, N. & Philippe, H. A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Mol. Biol. Evol. 21, 1095–1109 (2004)
Smith, S. A. & Dunn, C. W. Phyutility: a phyloinformatics tool for trees, alignments and molecular data. Bioinformatics 24, 715–716 (2008)
Shimodaira, H. An approximately unbiased test of phylogenetic tree selection. Syst. Biol. 51, 492–508 (2002)
Chou, H. H. & Holmes, M. H. DNA sequence quality trimming and vector removal. Bioinformatics 17, 1093–1104 (2001)
Huang, X. & Madan, A. CAP3: a DNA sequence assembly program. Genome Res. 9, 868–877 (1999)
Lottaz, C., Iseli, C., Jongeneel, C. V. & Bucher, P. Modeling sequencing errors by combining Hidden Markov models. Bioinformatics 19, (2003)
O’Brien, K. P., Remm, M. & Sonnhammer, E. L. Inparanoid: a comprehensive database of eukaryotic orthologs. Nucleic Acids Res. 33, D476–D480 (2005)
Katoh, K., Kuma, K., Toh, H. & Miyata, T. MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res. 33, 511–518 (2005)
Misof, B. & Misof, K. A Monte Carlo approach successfully identifies randomness in multiple sequence alignments: a more objective means of data exclusion. Syst. Biol. (2009)
Roure, B., Rodriguez-Ezpeleta, N. & Philippe, H. SCaFoS: a tool for selection, concatenation and fusion of sequences for phylogenomics. BMC Evol. Biol. 7, (2007)
Okusu, A. & Giribet, G. New 18S rRNA sequences from neomenioid aplacophorans and the possible origin of persistent exogenous contamination. J. Molluscan Stud. 69, 385–387 (2003)
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2 – approximately maximum-likelihood trees for large alignments. PLoS ONE 5, (2010)
Acknowledgements
We thank W. Jones and K. T. Fielman for help with cDNA library preparation, R. M. Jennings, N. Mikkelsen, and the crews of the RV Håkon Mosby, RV Hans Brattstrom and RV Laurence M. Gould for assistance collecting aplacophorans, and J. C. Havird, P. J. Krug, S. C. Kempf, D. R. Lindberg, M. V. Matz, L. R. Page and T. H. Struck for discussions. D. Speiser kindly shared the photo of Argopecten. F. W. Goetz, A. Gracey and M. L. Blaxter kindly provided sequence quality data for Dreissena rostriformis, Mytilus californianus and Lumbricus rubellus, respectively. We thank A. Di Cosmo, P. Burbach, V. Rehder, W. Wright and R. Gillette for providing samples of Octopus, Loligo, Helisoma, Dolabrifera and Pleurobranchaea as well as sharing some sequencing cost for these species. We also thank D. Young and the Alabama Supercomputer Authority for access to computational resources. The genomes of Capitella teleta, Helobdella robusta, Lottia gigantea and Nematostella vectensis were produced by the US Department of Energy Joint Genome Institute in collaboration with the user community. This work was supported by National Science Foundation (NSF) grants (0744649 and 0821622) to K.M.H., National Institute of Health (NIH) grants (1RO1NS06076, 1R01GM097502, R21 RR025699, R21DA030118) and the McKnight Brain Research Foundation to L.L.M., the Deep Metazoan Phylogeny (DMP) program of the German Science Foundation (Li 998/9-1) to B.L., and The University of Bergen (Norway) free researcher initiated project grant to C.T. (project no. 226270). This work represents contributions 82 and 4 to the Auburn University (AU) Marine Biology Program and Molette Biology Laboratory for Environmental and Climate Change Studies, respectively.
Author information
Authors and Affiliations
Contributions
K.M.H., C.T., B.L., C.S. and K.M.K. conceived and designed this study. K.M.H., L.L.M., B.L. and C.T. supervised cDNA preparation and sequencing. L.L.M., A.B.K., K.M.K., J.T.C. and A.M. prepared and sequenced cDNA. K.M.K., J.T.C., S.R.S. and M.R.C. developed the bioinformatics pipeline. K.M.K. performed phylogenetic and ancestral state reconstruction analyses. K.M.K. and J.T.C. prepared the figures. C.S., C.T. and K.M.K. modified the morphological character matrix. A.B.K., K.M.K. and A.M. submitted sequences to GenBank. All authors contributed in preparing the Letter.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Results, Supplementary References, Supplementary Tables 1-7 and Supplementary Figures 1-16 with legends. (PDF 14738 kb)
Rights and permissions
About this article
Cite this article
Kocot, K., Cannon, J., Todt, C. et al. Phylogenomics reveals deep molluscan relationships. Nature 477, 452–456 (2011). https://doi.org/10.1038/nature10382
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10382
This article is cited by
-
A chromosome-level genome assembly of a deep-sea symbiotic Aplacophora mollusc Chaetoderma sp.
Scientific Data (2024)
-
Comparative proteomics of the shell matrix proteins of Nautilus pompilius and the conchiferans provide insights into mollusk shell evolution at the molecular level
Marine Biology (2023)
-
Diversification of the aquaporin family in geographical isolated oyster species promote the adaptability to dynamic environments
BMC Genomics (2022)
-
Elemental analyses reveal distinct mineralization patterns in radular teeth of various molluscan taxa
Scientific Reports (2022)
-
Mito-nuclear coevolution and phylogenetic artifacts: the case of bivalve mollusks
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.