Abstract
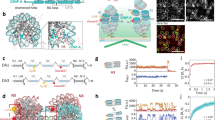
During cell division, chromosomes are segregated to nascent daughter cells by attaching to the microtubules of the mitotic spindle through the kinetochore. Kinetochores are assembled on a specialized chromatin domain called the centromere, which is characterized by the replacement of nucleosomal histone H3 with the histone H3 variant centromere protein A (CENP-A). CENP-A is essential for centromere and kinetochore formation in all eukaryotes but it is unknown how CENP-A chromatin directs centromere and kinetochore assembly1. Here we generate synthetic CENP-A chromatin that recapitulates essential steps of centromere and kinetochore assembly in vitro. We show that reconstituted CENP-A chromatin when added to cell-free extracts is sufficient for the assembly of centromere and kinetochore proteins, microtubule binding and stabilization, and mitotic checkpoint function. Using chromatin assembled from histone H3/CENP-A chimaeras, we demonstrate that the conserved carboxy terminus of CENP-A is necessary and sufficient for centromere and kinetochore protein recruitment and function but that the CENP-A targeting domain—required for new CENP-A histone assembly2—is not. These data show that two of the primary requirements for accurate chromosome segregation, the assembly of the kinetochore and the propagation of CENP-A chromatin, are specified by different elements in the CENP-A histone. Our unique cell-free system enables complete control and manipulation of the chromatin substrate and thus presents a powerful tool to study centromere and kinetochore assembly.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Cheeseman, I. M. & Desai, A. Molecular architecture of the kinetochore-microtubule interface. Nature Rev. Mol. Cell Biol. 9, 33–46 (2008)
Black, B. E. et al. Structural determinants for generating centromeric chromatin. Nature 430, 578–582 (2004)
Black, B. E. & Bassett, E. A. The histone variant CENP-A and centromere specification. Curr. Opin. Cell Biol. 20, 91–100 (2008)
Sekulic, N., Bassett, E. A., Rogers, D. J. & Black, B. E. The structure of (CENP-A-H4)2 reveals physical features that mark centromeres. Nature 467, 347–351 (2010)
Carroll, C. W., Milks, K. J. & Straight, A. F. Dual recognition of CENP-A nucleosomes is required for centromere assembly. J. Cell Biol. 189, 1143–1155 (2010)
Carroll, C. W., Silva, M. C. C., Godek, K. M., Jansen, L. E. T. & Straight, A. F. Centromere assembly requires the direct recognition of CENP-A nucleosomes by CENP-N. Nature Cell Biol. 11, 896–902 (2009)
Dunleavy, E. M. et al. HJURP is a cell-cycle-dependent maintenance and deposition factor of CENP-A at centromeres. Cell 137, 485–497 (2009)
Foltz, D. R. et al. Centromere-specific assembly of CENP-A nucleosomes is mediated by HJURP. Cell 137, 472–484 (2009)
Hu, H. et al. Structure of a CENP-A-histone H4 heterodimer in complex with chaperone HJURP. Genes Dev. 25, 901–906 (2011)
Tachiwana, H. et al. Crystal structure of the human centromeric nucleosome containing CENP-A. Nature 10.1038/nature10258 (10 July, 2011)
Blower, M. D., Sullivan, B. A. & Karpen, G. H. Conserved organization of centromeric chromatin in flies and humans. Dev. Cell 2, 319–330 (2002)
Ribeiro, S. et al. A super-resolution map of the vertebrate kinetochore. Proc. Natl Acad. Sci. USA 107, 10484–10489 (2010)
Zinkowski, R. P., Meyne, J. & Brinkley, B. R. The centromere-kinetochore complex: a repeat subunit model. J. Cell Biol. 113, 1091–1110 (1991)
Huynh, V., Robinson, P. & Rhodes, D. A method for the in vitro reconstitution of a defined “30 nm” chromatin fibre containing stoichiometric amounts of the linker histone. J. Mol. Biol. 345, 957–968 (2005)
Luger, K., Rechsteiner, T. J., Flaus, A. J., Waye, M. M. & Richmond, T. J. Characterization of nucleosome core particles containing histone proteins made in bacteria. J. Mol. Biol. 272, 301–311 (1997)
Desai, A., Murray, A., Mitchison, T. J. & Walczak, C. E. The use of Xenopus egg extracts to study mitotic spindle assembly and function in vitro . Methods Cell Biol. 61, 385–412 (1999)
Hori, T. et al. CCAN makes multiple contacts with centromeric DNA to provide distinct pathways to the outer kinetochore. Cell 135, 1039–1052 (2008)
Kawashima, S. A., Yamagishi, Y., Honda, T., Ishiguro, K.-i. & Watanabe, Y. Phosphorylation of H2A by Bub1 prevents chromosomal instability through localizing shugoshin. Science 327, 172–177 (2010)
Kelly, A. E. et al. Survivin reads phosphorylated histone H3 threonine 3 to activate the mitotic kinase Aurora B. Science 330, 235–239 (2010)
Wang, F. et al. Histone H3 Thr-3 phosphorylation by Haspin positions Aurora B at centromeres in mitosis. Science 330, 231–235 (2010)
Budde, P. P., Kumagai, A., Dunphy, W. G. & Heald, R. Regulation of Op18 during spindle assembly in Xenopus egg extracts. J. Cell Biol. 153, 149–158 (2001)
Blow, J. J. & Laskey, R. A. Initiation of DNA replication in nuclei and purified DNA by a cell-free extract of Xenopus eggs. Cell 47, 577–587 (1986)
Minshull, J., Sun, H., Tonks, N. K. & Murray, A. W. A MAP kinase-dependent spindle assembly checkpoint in Xenopus egg extracts. Cell 79, 475–486 (1994)
Sawin, K. E. & Mitchison, T. J. Mitotic spindle assembly by two different pathways in vitro . J. Cell Biol. 112, 925–940 (1991)
Heald, R. et al. Self-organization of microtubules into bipolar spindles around artificial chromosomes in Xenopus egg extracts. Nature 382, 420–425 (1996)
Nicklas, R. B., Ward, S. C. & Gorbsky, G. J. Kinetochore chemistry is sensitive to tension and may link mitotic forces to a cell cycle checkpoint. J. Cell Biol. 130, 929–939 (1995)
Rieder, C. L., Cole, R. W., Khodjakov, A. & Sluder, G. The checkpoint delaying anaphase in response to chromosome monoorientation is mediated by an inhibitory signal produced by unattached kinetochores. J. Cell Biol. 130, 941–948 (1995)
Milks, K. J., Moree, B. & Straight, A. F. Dissection of CENP-C-directed centromere and kinetochore assembly. Mol. Biol. Cell 20, 4246–4255 (2009)
Foltz, D. R. et al. The human CENP-A centromeric nucleosome-associated complex. Nature Cell Biol. 8, 458–469 (2006)
McClelland, S. E. et al. The CENP-A NAC/CAD kinetochore complex controls chromosome congression and spindle bipolarity. EMBO J. 26, 5033–5047 (2007)
Luger, K., Rechsteiner, T. & Richmond, T. Preparation of nucleosome core particle from recombinant histones. Methods Enzymol. 304, 3–19 (1999)
Lowary, P. T. & Widom, J. New DNA sequence rules for high affinity binding to histone octamer and sequence-directed nucleosome positioning. J. Mol. Biol. 276, 19–42 (1998)
Murray, A. W. Cell cycle extracts. Methods Cell Biol. 36, 581–605 (1991)
Kim, S., Song, E., Lee, K. & Ferrell, J., Jr Multisite M-phase phosphorylation of Xenopus Wee1A. Mol. Cell. Biol. 25, 10580–10590 (2005)
Hannak, E. & Heald, R. Investigating mitotic spindle assembly and function in vitro using Xenopus laevis egg extracts. Nature Protocols 1, 2305–2314 (2006)
Field, C. M., Oegema, K., Zheng, Y., Mitchison, T. J. & Walczak, C. E. Purification of cytoskeletal proteins using peptide antibodies. Methods Enzymol. 298, 525–541 (1998)
Osborne, J. W. Best Practices in Quantitative Methods (Sage Publications, 2008)
Acknowledgements
The authors would like to thank A.F.S. laboratory members for support and comments, J. E. Ferrell, A. Murray, R.-H. Chen, G. Kops and P. T. Stukenberg for providing antibodies. D. Rhodes, P. Robinson, K. Luger, J. Hansen, G. Narlikar and J. Yang for providing reagents and advice. A.G. was supported by a postdoctoral fellowship from the German Research Foundation (DFG). C.W.C. was supported by a postdoctoral fellowship from the Helen Hay Whitney Foundation and the American Heart Association (AHA). B.M. was supported by T32GM007276, C.J.F. was supported by a Stanford Graduate Fellowship and this work was supported by National Institutes of Health (NIH) R01GM074728 to A.F.S.
Author information
Authors and Affiliations
Contributions
A.G. and A.F.S. designed the experiments and wrote the manuscript. A.G. performed all the experiments. C.W.C. purified the CENP-A/H3 chimaeras and assembled arrays containing chimaeric proteins, analysed Xenopus cenp-n binding to human CENP-A mononucleosomes and provided advice. B.M. generated Xenopus centromere protein antibodies and C.J.F. designed and wrote the image analysis software for quantitative analysis.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
The file contains Supplementary Figures 1-10 with legends. (PDF 3910 kb)
Rights and permissions
About this article
Cite this article
Guse, A., Carroll, C., Moree, B. et al. In vitro centromere and kinetochore assembly on defined chromatin templates. Nature 477, 354–358 (2011). https://doi.org/10.1038/nature10379
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10379
This article is cited by
-
The CENP-A centromere targeting domain facilitates H4K20 monomethylation in the nucleosome by structural polymorphism
Nature Communications (2019)
-
DNA replication acts as an error correction mechanism to maintain centromere identity by restricting CENP-A to centromeres
Nature Cell Biology (2019)
-
Structure of the inner kinetochore CCAN complex assembled onto a centromeric nucleosome
Nature (2019)
-
Phosphorylation of CENP-A on serine 7 does not control centromere function
Nature Communications (2019)
-
High-resolution mapping of centromeric protein association using APEX-chromatin fibers
Epigenetics & Chromatin (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.