Abstract
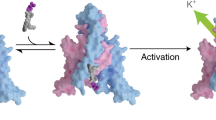
The regulation of ion channel activity by specific lipid molecules is widely recognized as an integral component of electrical signalling in cells1,2. In particular, phosphatidylinositol 4,5-bisphosphate (PIP2), a minor yet dynamic phospholipid component of cell membranes, is known to regulate many different ion channels2,3,4,5,6,7,8. PIP2 is the primary agonist for classical inward rectifier (Kir2) channels, through which this lipid can regulate a cell’s resting membrane potential2,7,8,9. However, the molecular mechanism by which PIP2 exerts its action is unknown. Here we present the X-ray crystal structure of a Kir2.2 channel in complex with a short-chain (dioctanoyl) derivative of PIP2. We found that PIP2 binds at an interface between the transmembrane domain (TMD) and the cytoplasmic domain (CTD). The PIP2-binding site consists of a conserved non-specific phospholipid-binding region in the TMD and a specific phosphatidylinositol-binding region in the CTD. On PIP2 binding, a flexible expansion linker contracts to a compact helical structure, the CTD translates 6 Å and becomes tethered to the TMD and the inner helix gate begins to open. In contrast, the small anionic lipid dioctanoyl glycerol pyrophosphatidic acid (PPA) also binds to the non-specific TMD region, but not to the specific phosphatidylinositol region, and thus fails to engage the CTD or open the channel. Our results show how PIP2 can control the resting membrane potential through a specific ion-channel-receptor–ligand interaction that brings about a large conformational change, analogous to neurotransmitter activation of ion channels at synapses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
Atomic coordinates and structure factors for the reported crystal structures have been deposited into the Protein Data Bank under accession codes 3SPI (wild-type PIP2), 3SPC (wild-type PPA), 3SPH (PIP2(I223L)), 3SPJ (apo(I223L)) and 3SPG (PIP2(R186A)).
References
Dart, C. Lipid microdomains and the regulation of ion channel function. J. Physiol. (Lond.) 588, 3169–3178 (2010)
Hilgemann, D. W., Feng, S. & Nasuhoglu, C. The complex and intriguing lives of PIP2 with ion channels and transporters. Sci. STKE 2001, re19 (2001)
Suh, B. C. & Hille, B. Regulation of ion channels by phosphatidylinositol 4,5-bisphosphate. Curr. Opin. Neurobiol. 15, 370–378 (2005)
Fujiwara, Y. & Kubo, Y. Regulation of the desensitization and ion selectivity of ATP-gated P2X2 channels by phosphoinositides. J. Physiol. (Lond.) 576, 135–149 (2006)
Logothetis, D. E., Jin, T., Lupyan, D. & Rosenhouse-Dantsker, A. Phosphoinositide-mediated gating of inwardly rectifying K+ channels. Pflügers Arch. Eur. J. Physiol. 455, 83–95 (2007)
Vaithianathan, T. et al. Direct regulation of BK channels by phosphatidylinositol 4,5-bisphosphate as a novel signaling pathway. J. Gen. Physiol. 132, 13–28 (2008)
Gamper, N. & Shapiro, M. S. Regulation of ion transport proteins by membrane phosphoinositides. Nature Rev. Neurosci. 8, 921–934 (2007)
Huang, C. L., Feng, S. & Hilgemann, D. W. Direct activation of inward rectifier potassium channels by PIP2 and its stabilization by Gβγ. Nature 391, 803–806 (1998)
Lopes, C. M. B. et al. Alterations in conserved Kir channel-PIP2 interactions underlie channelopathies. Neuron 34, 933–944 (2002)
Berridge, M. J. Inositol trisphosphate and calcium signalling. Nature 361, 315–325 (1993)
Monserrate, J. P. & York, J. D. Inositol phosphate synthesis and the nuclear processes they affect. Curr. Opin. Cell Biol. 22, 365–373 (2010)
Martin, T. F. PI(4,5)P2 regulation of surface membrane traffic. Curr. Opin. Cell Biol. 13, 493–499 (2001)
Cho, W. & Stahelin, R. V. Membrane-protein interactions in cell signaling and membrane trafficking. Annu. Rev. Biophys. Biomol. Struct. 34, 119–151 (2005)
Heo, W. D. et al. PI(3,4,5)P3 and PI(4,5)P2 lipids target proteins with polybasic clusters to the plasma membrane. Science 314, 1458–1461 (2006)
Hilgemann, D. W. & Ball, R. Regulation of cardiac Na+,Ca2+ exchange and KATP potassium channels by PIP2 . Science 273, 956–959 (1996)
Stanfield, P. R., Nakajima, S. & Nakajima, Y. Constitutively active and G-protein coupled inward rectifier K+ channels: Kir2.0 and Kir3.0. Rev. Physiol. Biochem. Pharmacol. 145, 47–179 (2002)
Rohács, T. et al. Specificity of activation by phosphoinositides determines lipid regulation of Kir channels. Proc. Natl Acad. Sci. USA 100, 745–750 (2003)
Enkvetchakul, D., Jeliazkova, I. & Nichols, C. G. Direct modulation of Kir channel gating by membrane phosphatidylinositol 4,5-bisphosphate. J. Biol. Chem. 280, 35785–35788 (2005)
Tao, X., Avalos, J. L., Chen, J. & MacKinnon, R. Crystal structure of the eukaryotic strong inward-rectifier K+ channel Kir2.2 at 3.1 Å resolution. Science 326, 1668–1674 (2009)
Zhang, H., He, C., Yan, X., Mirshahi, T. & Logothetis, D. E. Activation of inwardly rectifying K+ channels by distinct PtdIns(4,5)P2 interactions. Nature Cell Biol. 1, 183–188 (1999)
Cheng, W. W., D’Avanzo, N., Doyle, D. A. & Nichols, C. G. Dual-mode phospholipid regulation of human inward rectifying potassium channels. Biophys. J. 100, 620–628 (2011)
Clarke, O. B. et al. Domain reorientation and rotation of an intracellular assembly regulate conduction in Kir potassium channels. Cell 141, 1018–1029 (2010)
Pegan, S. et al. Cytoplasmic domain structures of Kir2.1 and Kir3.1 show sites for modulating gating and rectification. Nature Neurosci. 8, 279–287 (2005)
Vagin, A. & Teplyakov, A. An approach to multi-copy search in molecular replacement. Acta Crystallogr. D 56, 1622–1624 (2000)
Collaborative Computational Projet 4 The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Xie, L.-H., John, S., a, Ribalet, B. & Weiss, J. N. Activation of inwardly rectifying potassium (Kir) channels by phosphatidylinosital-4,5-bisphosphate (PIP2): interaction with other regulatory ligands. Prog. Biophys. Mol. Biol. 94, 320–335 (2007)
Cohen, S. X. et al. Towards complete validated models in the next generation of ARP/wARP. Acta Crystallogr. D 60, 2222–2229 (2004)
Delano, W. L. PyMOL. 〈http://www.pymol.org〉 (Delano Scientific, 2002)
Acknowledgements
We thank staff members at NSLS X29 and X25, Brookhaven National Laboratory for beamline assistance, members of the Gadsby laboratory (Rockefeller University) for help in Xenopus oocyte preparation, R. Molday (University of British Columbia) for providing the anti-1D4 tag cell line and members of the MacKinnon laboratory for helpful suggestions. R.M. is an investigator in the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
S.B.H. purified and crystallized the protein; collected, processed and refined crystallographic data, and performed electrophysiology experiments. X.T. aided in experimental design and provided assistance in all aspects of the project. R.M. designed the study and analysed data. All authors wrote and discussed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-7 with legends and Supplementary Table 1. (PDF 894 kb)
Rights and permissions
About this article
Cite this article
Hansen, S., Tao, X. & MacKinnon, R. Structural basis of PIP2 activation of the classical inward rectifier K+ channel Kir2.2. Nature 477, 495–498 (2011). https://doi.org/10.1038/nature10370
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10370
This article is cited by
-
Structure of an open KATP channel reveals tandem PIP2 binding sites mediating the Kir6.2 and SUR1 regulatory interface
Nature Communications (2024)
-
Conformational plasticity of NaK2K and TREK2 potassium channel selectivity filters
Nature Communications (2023)
-
Disruption of ER ion homeostasis maintained by an ER anion channel CLCC1 contributes to ALS-like pathologies
Cell Research (2023)
-
Subunit gating resulting from individual protonation events in Kir2 channels
Nature Communications (2023)
-
Regulation of membrane protein structure and function by their lipid nano-environment
Nature Reviews Molecular Cell Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.