Abstract
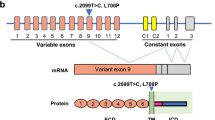
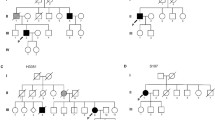
Amyotrophic lateral sclerosis (ALS) is a paralytic and usually fatal disorder caused by motor-neuron degeneration in the brain and spinal cord. Most cases of ALS are sporadic but about 5–10% are familial. Mutations in superoxide dismutase 1 (SOD1)1,2, TAR DNA-binding protein (TARDBP, also known as TDP43)3,4 and fused in sarcoma (FUS, also known as translocated in liposarcoma (TLS))5,6 account for approximately 30% of classic familial ALS. Mutations in several other genes have also been reported as rare causes of ALS or ALS-like syndromes7,8,9,10,11,12,13,14,15. The causes of the remaining cases of familial ALS and of the vast majority of sporadic ALS are unknown. Despite extensive studies of previously identified ALS-causing genes, the pathogenic mechanism underlying motor-neuron degeneration in ALS remains largely obscure. Dementia, usually of the frontotemporal lobar type, may occur in some ALS cases. It is unclear whether ALS and dementia share common aetiology and pathogenesis in ALS/dementia. Here we show that mutations in UBQLN2, which encodes the ubiquitin-like protein ubiquilin 2, cause dominantly inherited, chromosome-X-linked ALS and ALS/dementia. We describe novel ubiquilin 2 pathology in the spinal cords of ALS cases and in the brains of ALS/dementia cases with or without UBQLN2 mutations. Ubiquilin 2 is a member of the ubiquilin family, which regulates the degradation of ubiquitinated proteins. Functional analysis showed that mutations in UBQLN2 lead to an impairment of protein degradation. Therefore, our findings link abnormalities in ubiquilin 2 to defects in the protein degradation pathway, abnormal protein aggregation and neurodegeneration, indicating a common pathogenic mechanism that can be exploited for therapeutic intervention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Deng, H. X. et al. Amyotrophic lateral sclerosis and structural defects in Cu,Zn superoxide dismutase. Science 261, 1047–1051 (1993)
Rosen, D. R. et al. Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 362, 59–62 (1993)
Kabashi, E. et al. TARDBP mutations in individuals with sporadic and familial amyotrophic lateral sclerosis. Nature Genet. 40, 572–574 (2008)
Sreedharan, J. et al. TDP-43 mutations in familial and sporadic amyotrophic lateral sclerosis. Science 319, 1668–1672 (2008)
Kwiatkowski, T. J., Jr et al. Mutations in the FUS/TLS gene on chromosome 16 cause familial amyotrophic lateral sclerosis. Science 323, 1205–1208 (2009)
Vance, C. et al. Mutations in FUS, an RNA processing protein, cause familial amyotrophic lateral sclerosis type 6. Science 323, 1208–1211 (2009)
Chen, Y. Z. et al. DNA/RNA helicase gene mutations in a form of juvenile amyotrophic lateral sclerosis (ALS4). Am. J. Hum. Genet. 74, 1128–1135 (2004)
Greenway, M. J. et al. ANG mutations segregate with familial and ‘sporadic’ amyotrophic lateral sclerosis. Nature Genet. 38, 411–413 (2006)
Nishimura, A. L. et al. A mutation in the vesicle-trafficking protein VAPB causes late-onset spinal muscular atrophy and amyotrophic lateral sclerosis. Am. J. Hum. Genet. 75, 822–831 (2004)
Yang, Y. et al. The gene encoding alsin, a protein with three guanine-nucleotide exchange factor domains, is mutated in a form of recessive amyotrophic lateral sclerosis. Nature Genet. 29, 160–165 (2001)
Chow, C. Y. et al. Deleterious variants of FIG4, a phosphoinositide phosphatase, in patients with ALS. Am. J. Hum. Genet. 84, 85–88 (2009)
Maruyama, H. et al. Mutations of optineurin in amyotrophic lateral sclerosis. Nature 465, 223–226 (2010)
Ticozzi, N. et al. Paraoxonase gene mutations in amyotrophic lateral sclerosis. Ann. Neurol. 68, 102–107 (2010)
Mitchell, J. et al. Familial amyotrophic lateral sclerosis is associated with a mutation in D-amino acid oxidase. Proc. Natl Acad. Sci. USA 107, 7556–7561 (2010)
Johnson, J. O. et al. Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 68, 857–864 (2010)
Lansbury, P. T. & Lashuel, H. A. A century-old debate on protein aggregation and neurodegeneration enters the clinic. Nature 443, 774–779 (2006)
Deng, H. X. et al. FUS-immunoreactive inclusions are a common feature in sporadic and non-SOD1 familial amyotrophic lateral sclerosis. Ann. Neurol. 67, 739–748 (2010)
Neumann, M. et al. Ubiquitinated TDP-43 in frontotemporal lobar degeneration and amyotrophic lateral sclerosis. Science 314, 130–133 (2006)
Shibata, N. et al. Intense superoxide dismutase-1 immunoreactivity in intracytoplasmic hyaline inclusions of familial amyotrophic lateral sclerosis with posterior column involvement. J. Neuropathol. Exp. Neurol. 55, 481–490 (1996)
Mackenzie, I. R. et al. Pathological TDP-43 distinguishes sporadic amyotrophic lateral sclerosis from amyotrophic lateral sclerosis with SOD1 mutations. Ann. Neurol. 61, 427–434 (2007)
Neumann, M. et al. Frontotemporal lobar degeneration with FUS pathology. Brain 132, 2922–2931 (2009)
Urwin, H. et al. FUS pathology defines the majority of tau- and TDP-43-negative frontotemporal lobar degeneration. Acta Neuropathol. 120, 33–41 (2010)
Nonaka, T., Kametani, F., Arai, T., Akiyama, H. & Hasegawa, M. Truncation and pathogenic mutations facilitate the formation of intracellular aggregates of TDP-43. Hum. Mol. Genet. 18, 3353–3364 (2009)
Ko, H. S., Uehara, T., Tsuruma, K. & Nomura, Y. Ubiquilin interacts with ubiquitylated proteins and proteasome through its ubiquitin-associated and ubiquitin-like domains. FEBS Lett. 566, 110–114 (2004)
Dantuma, N. P., Lindsten, K., Glas, R., Jellne, M. & Masucci, M. G. Short-lived green fluorescent proteins for quantifying ubiquitin/proteasome-dependent proteolysis in living cells. Nature Biotechnol. 18, 538–543 (2000)
Kay, B. K., Williamson, M. P. & Sudol, M. The importance of being proline: the interaction of proline-rich motifs in signaling proteins with their cognate domains. FASEB J. 14, 231–241 (2000)
Aitio, O. et al. Recognition of tandem PxxP motifs as a unique Src homology 3-binding mode triggers pathogen-driven actin assembly. Proc. Natl Acad. Sci. USA 107, 21743–21748 (2010)
Haapasalo, A. et al. Emerging role of Alzheimer’s disease-associated ubiquilin-1 in protein aggregation. Biochem. Soc. Trans. 38, 150–155 (2010)
Kim, S. H. et al. Potentiation of amyotrophic lateral sclerosis (ALS)-associated TDP-43 aggregation by the proteasome-targeting factor, ubiquilin 1. J. Biol. Chem. 284, 8083–8092 (2009)
Aguzzi, A. & O'Connor, T. Protein aggregation diseases: pathogenicity and therapeutic perspectives. Nature Rev. Drug Discov. 9, 237–248 (2010)
Brooks, B. R., Miller, R. G., Swash, M. & Munsat, T. L. El Escorial revisited: revised criteria for the diagnosis of amyotrophic lateral sclerosis. Amyotroph. Lateral Scler. Other Motor Neuron Disord. 1, 293–299 (2000)
Neary, D. et al. Frontotemporal lobar degeneration: a consensus on clinical diagnostic criteria. Neurology 51, 1546–1554 (1998)
Cairns, N. J. et al. Neuropathologic diagnostic and nosologic criteria for frontotemporal lobar degeneration: consensus of the consortium for frontotemporal lobar degeneration. Acta Neuropathol. 114, 5–22 (2007)
Mackenzie, I. R. et al. Heterogeneity of ubiquitin pathology in frontotemporal lobar degeneration: classification and relation to clinical phenotype. Acta Neuropathol. 112, 539–549 (2006)
Deng, H. X. et al. Conversion to the amyotrophic lateral sclerosis phenotype is associated with intermolecular linked insoluble aggregates of SOD1 in mitochondria. Proc. Natl Acad. Sci. USA 103, 7142–7147 (2006)
Fecto, F. et al. Mutant TRPV4-mediated toxicity is linked to increased constitutive function in axonal neuropathies. J. Biol. Chem. 286, 17281–17291 (2011)
Acknowledgements
This study was supported by the National Institute of Neurological Disorders and Stroke (NS050641), the Les Turner ALS Foundation, the Vena E. Schaff ALS Research Fund, the Harold Post Research Professorship, the Herbert and Florence C. Wenske Foundation, the David C. Asselin MD Memorial Fund, the Help America Foundation and the Les Turner ALS Foundation/Herbert C. Wenske Foundation Professorship. F.F. has support from NIH (T32 AG20506). K.A. is a postdoctoral fellow of the Blazeman Foundation for ALS. G.H.G. received travel funds from MND Scotland. We thank N. Dantuma for the UPS reporter plasmid (through Addgene) and the staff of the Northwestern University Robert H. Lurie Comprehensive Cancer Center flow cytometry core facility for technical assistance. Imaging work was performed at the Northwestern University Cell Imaging Facility, supported by NCI CCSG P30 CA060553 awarded to the Robert H Lurie Comprehensive Cancer Center.
Author information
Authors and Affiliations
Contributions
T.S. conceived and supervised this project. W.C., S.-T.H., Y.Y., H.J., M.H., H.-X.D. and T.S. did the sequencing analysis. S.T.H., E.R., J.L.H., M.P.-V. and T.S. performed linkage analysis. K.M.B., G.H.G., F.F., G.H.J., H.Z., E.H.B., K.A., E.M., H.-X.D. and T.S. performed immunohistochemical, confocal and pathological analysis. F.F., Y.S. and H.-X.D. performed functional analysis. N.S., S.D. and T.S. collected family information and coordinated this study. K.M.B., G.H.J., B.R.B., R.L.S. and T.S. did clinical studies. H.-X.D. and T.S. analysed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-18 with legends, Supplementary Tables 1-2 and Supplementary Case Notes. (PDF 2329 kb)
Rights and permissions
About this article
Cite this article
Deng, HX., Chen, W., Hong, ST. et al. Mutations in UBQLN2 cause dominant X-linked juvenile and adult-onset ALS and ALS/dementia. Nature 477, 211–215 (2011). https://doi.org/10.1038/nature10353
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10353
This article is cited by
-
Glutathionylation on RNA-binding proteins: a regulator of liquid‒liquid phase separation in the pathogenesis of amyotrophic lateral sclerosis
Experimental & Molecular Medicine (2023)
-
Amyotrophic lateral sclerosis: translating genetic discoveries into therapies
Nature Reviews Genetics (2023)
-
Ubiquilin-2 regulates pathological alpha-synuclein
Scientific Reports (2023)
-
Mutation spectrum of chinese amyotrophic lateral sclerosis patients with frontotemporal dementia
Orphanet Journal of Rare Diseases (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.