Abstract
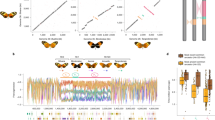
Supergenes are tight clusters of loci that facilitate the co-segregation of adaptive variation, providing integrated control of complex adaptive phenotypes1. Polymorphic supergenes, in which specific combinations of traits are maintained within a single population, were first described for ‘pin’ and ‘thrum’ floral types in Primula1 and Fagopyrum2, but classic examples are also found in insect mimicry3,4,5 and snail morphology6. Understanding the evolutionary mechanisms that generate these co-adapted gene sets, as well as the mode of limiting the production of unfit recombinant forms, remains a substantial challenge7,8,9,10. Here we show that individual wing-pattern morphs in the polymorphic mimetic butterfly Heliconius numata are associated with different genomic rearrangements at the supergene locus P. These rearrangements tighten the genetic linkage between at least two colour-pattern loci that are known to recombine in closely related species9,10,11, with complete suppression of recombination being observed in experimental crosses across a 400-kilobase interval containing at least 18 genes. In natural populations, notable patterns of linkage disequilibrium (LD) are observed across the entire P region. The resulting divergent haplotype clades and inversion breakpoints are found in complete association with wing-pattern morphs. Our results indicate that allelic combinations at known wing-patterning loci have become locked together in a polymorphic rearrangement at the P locus, forming a supergene that acts as a simple switch between complex adaptive phenotypes found in sympatry. These findings highlight how genomic rearrangements can have a central role in the coexistence of adaptive phenotypes involving several genes acting in concert, by locally limiting recombination and gene flow.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Data deposits
GenBank accessions for BAC clone sequences: FP885863, FP476061, FP565803, FP476023, CU856181, FP885878, FP476047, FP885857, CU856182, CU655868, FP885879, FP885861, FP885880, FP885855, CU914733, FP475989, CU655869, CU914734, CU633161, CU638865, CU856175, FP884220 and FP236755. Accessions for 1364 marker sequences: JN173798–JN175161.
References
Mather, K. The genetical architecture of heterostyly in Primula sinensis. Evolution 4, 340–352 (1950)
Garber, R. J. & Quisenberry, K. S. The inheritance of length of style in buckwheat. J. Agric. Res. 34, 181–183 (1927)
Clarke, C. A., Sheppard, P. M. & Thornton, I. W. B. The genetics of the mimetic butterfly Papilio memnon. Philos. Trans. R. Soc. Lond. B 254, 37–89 (1968)
Brown, K. S. & Benson, W. W. Adaptive polymorphism associated with multiple Müllerian mimicry in Heliconius numata. Biotropica 6, 205–228 (1974)
Nijhout, H. F. Polymorphic mimicry in Papilio dardanus: mosaic dominance, big effects, and origins. Evol. Dev. 5, 579–592 (2003)
Murray, J. & Clarke, B. Supergenes in polymorphic land snails—examples from genus Partula. Genetics 74, S188–S189 (1973)
Kirkpatrick, M. & Barton, N. Chromosome inversions, local adaptation and speciation. Genetics 173, 419–434 (2006)
Manfield, I. W. et al. Molecular characterization of DNA sequences from the Primula vulgaris S-locus. J. Exp. Bot. 56, 1177–1188 (2005)
Joron, M. et al. A conserved supergene locus controls colour pattern diversity in Heliconius butterflies. PLoS Biol. 4, e303 (2006)
Baxter, S. W. et al. Genomic hotspots for adaptation: the population genetics of Müllerian mimicry in the Heliconius melpomene clade. PLoS Genet. 6, e1000794 (2010)
Ferguson, L. et al. Characterization of a hotspot for mimicry: Assembly of a butterfly wing transcriptome to genomic sequence at the HmYb/Sb locus. Mol. Ecol. 19, 240–254 (2010)
Hoffmann, A. A. & Rieseberg, L. H. Revisiting the impact of inversions in evolution: from population genetic markers to drivers of adaptive shifts and speciation? Annu. Rev. Ecol. Evol. Syst. 39, 21–42 (2008)
Pinho, C. & Hey, J. Divergence with gene flow: Models and data. Annu. Rev. Ecol. Evol. Syst. 41, 215–230 (2010)
Clark, R. et al. Colour pattern specification in the Mocker swallowtail Papilio dardanus: the transcription factor invected is a candidate for the mimicry locus H. Proc. R. Soc. B 275, 1181–1188 (2008)
Joron, M., Wynne, I. R., Lamas, G. & Mallet, J. Variable selection and the coexistence of multiple mimetic forms of the butterfly Heliconius numata. Evol. Ecol. 13, 721–754 (1999)
Turner, J. R. G. in The biology of butterflies Vol. 11 (eds Vane-Wright, R. I. & Ackery, P. R. ) 141–161 (Academic, 1984)
Charlesworth, D. & Charlesworth, B. Theoretical genetics of Batesian mimicry. II. Evolution of supergenes. J. Theor. Biol. 55, 305–324 (1975)
Alvarez, G. & Zapata, C. Conditions for protected inversion polymorphism under supergene selection. Genetics 146, 717–722 (1997)
Hatadani, L. M., Baptista, J. C. R., Souza, W. N. & Klaczko, L. B. Colour polymorphism in Drosophila mediopunctata: genetic (chromosomal) analysis and nonrandom association with chromosome inversions. Heredity 93, 525–534 (2004)
Lowry, D. B. & Willis, J. H. A widespread chromosomal inversion polymorphism contributes to a major life-history transition, local adaptation, and reproductive isolation. PLoS Biol. 8, e1000500 (2010)
Jones, R. T., Salazar, P., ffrench-Constant, R. H., Jiggins, C. D. & Joron, M. Evolution of a mimicry supergene from a multilocus architecture. Proc. R. Soc. B 10.1098/rspb.2011.0882 (2011)
Papa, R. et al. Highly conserved gene order and numerous novel repetitive elements in genomic regions linked to wing pattern variation in Heliconius butterflies. BMC Genomics 9, 345 (2008)
Pringle, E. G. et al. Synteny and chromosome evolution in the lepidoptera: Evidence from mapping in Heliconius melpomene. Genetics 177, 417–426 (2007)
Counterman, B. A. et al. Genomic hotspots for adaptation: the population genetics of Müllerian mimicry in Heliconius erato. PLoS Genet. 6, e1000796 (2010)
Dobzhansky, T. Genetics of natural populations. XIV. A response of certain gene arrangements in the third chromosome of Drosophila pseudoobscura to natural selection. Genetics 32, 142–160 (1947)
Saenko, S. V., Brakefield, P. M. & Beldade, P. Single locus affects embryonic segment polarity and multiple aspects of an adult evolutionary novelty. BMC Biol. 8, 111 (2010)
van't Hof, A. E., Edmonds, N., Dalíková, M., Marec, F. & Saccheri, I. J. Industrial melanism in British peppered moths has a singular and recent mutational origin. Science 332, 958–960 (2011)
Kronforst, M. R. et al. Linkage of butterfly mate preference and wing color preference cue at the genomic location of wingless. Proc. Natl Acad. Sci. USA 103, 6575–6580 (2006)
Chamberlain, N. L., Hill, R. I., Kapan, D. D., Gilbert, L. E. & Kronforst, M. R. Polymorphic butterfly reveals the missing link in ecological speciation. Science 326, 847–850 (2009)
Merrill, R. M., Van Schooten, B., Scott, J. A. & Jiggins, C. D. Pervasive genetic associations between traits causing reproductive isolation in Heliconius butterflies. Proc. R. Soc. B 278, 511–518 (2010)
Frazer, K. A., Pachter, L., Poliakov, A., Rubin, E. M. & Dubchak, I. VISTA: computational tools for comparative genomics. Nucleic Acids Res. 32, W273–W279 (2004)
Mayor, C. et al. VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16, 1046 (2000)
Cantarel, B. L. et al. MAKER: An easy-to-use annotation pipeline designed for emerging model organism genomes. Genome Res. 18, 188–196 (2008)
Chevreux, B. et al. Using the miraEST assembler for reliable and automated mRNA transcript assembly and SNP detection in sequenced ESTs. Genome Res. 14, 1147–1159 (2004)
Salazar, C., Jiggins, C. D., Taylor, J. E., Kronforst, M. R. & Linares, M. Gene flow and the genealogical history of Heliconius heurippa. BMC Evol. Biol. 8, 132 (2008)
Brown, K. S. An illustrated key to the silvaniform Heliconius (Lepidoptera: Nymphalidae) with descriptions of new subspecies. Trans. Am. Entomol. Soc. 102, 373–484 (1976)
Zaykin, D. V., Pudovkin, A. & Weir, B. S. Correlation-based inference for linkage disequilibrium with multiple alleles. Genetics 180, 533–545 (2008)
Stephens, M. & Donnelly, P. A comparison of Bayesian methods for haplotype reconstruction from population genotype data. Am. J. Hum. Genet. 73, 1162–1169 (2003)
Polzin, T. & Daneshmand, S. V. On Steiner trees and minimum spanning trees in hypergraphs. Oper. Res. Lett. 31, 12–20 (2003)
Forster, P., Torroni, A., Renfrew, C. & Rohl, A. Phylogenetic star contraction applied to Asian and Papuan mtDNA evolution. Mol. Biol. Evol. 18, 1864–1881 (2001)
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009)
Hudson, R. R., Slatkin, M. & Maddison, W. P. Estimation of levels of gene flow from DNA-sequence data. Genetics 132, 583–589 (1992)
Acknowledgements
We thank M. Blaxter and D. Charlesworth for advice throughout the study; The GenePool and S. Humphray for DNA sequencing; S. Kumar and A. Papanicolaou for bioinformatics support; M. Beltrán, A. Bulski, M. Veuille and the Botanique-Entomologie-Mycologie molecular facility (BoEM) for laboratory support; S. Johnston for genome-size estimates in H. numata; D. Obbard for providing R scripts; M. Abanto, S. Gallusser, C. Ramírez, L. de Silva, J. Barbut, B. Gilles and G. Lamas for help with butterfly rearing, fieldwork and collecting permits; and the Peruvian National Institute of Natural Resources (INRENA) for granting collecting and export permits (076-2007-INRENA-IFFS-DCB). Fieldwork in French Guiana was supported by a CNRS ‘Nouragues Research Grant’. This work was supported by an EMBO long-term fellowship (ALTF-431-2004), EMBO-matching funds from NWO (Netherlands), a Royal Society University Research Fellowship (516002.K5917/ROG), a CNRS grant (ATIP Biodiversité 2008, France) and a European Research Council Starting Grant (ERC-Stg ‘MimEvol’) to M.J., a BBSRC grant (BBE0118451) to C.D.J. and R.H.ff.-C., a Leverhulme Trust grant (F/00144AY) to R.H.ff.-C., and a Royal Society University Research Fellowship and a Leverhulme Research Leadership grant to C.D.J.
Author information
Authors and Affiliations
Contributions
M.J., C.D.J. and R.H.ff.-C. designed the study and contributed to all stages of the project. M.J., L. Frezal and R.T.J. performed the principal experiments and data analysis, with assistance from N.L.C., S.W.B., S.F.L., M.B., C.S., L. Ferguson, C.R.H., A.W. and P.A.W. BAC clone sequencing was carried out by C.D., R.G., C.L., R.C., H.B., S.S., J.R., M.C.J. and M.A.Q. M.J., A.W., C.D.J. and R.H.ff.-C. co-wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-6 with legends, Supplementary Tables 1-9 and additional references. (PDF 2716 kb)
Rights and permissions
About this article
Cite this article
Joron, M., Frezal, L., Jones, R. et al. Chromosomal rearrangements maintain a polymorphic supergene controlling butterfly mimicry. Nature 477, 203–206 (2011). https://doi.org/10.1038/nature10341
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10341
This article is cited by
-
Genetic architecture of acute hyperthermia resistance in juvenile rainbow trout (Oncorhynchus mykiss) and genetic correlations with production traits
Genetics Selection Evolution (2023)
-
A novel tetra-primer ARMS-PCR approach for the molecular karyotyping of chromosomal inversion 2Ru in the main malaria vectors Anopheles gambiae and Anopheles coluzzii
Parasites & Vectors (2023)
-
Reconstruction of hundreds of reference ancestral genomes across the eukaryotic kingdom
Nature Ecology & Evolution (2023)
-
Inversions maintain differences between migratory phenotypes of a songbird
Nature Communications (2023)
-
Dominance mechanisms in supergene alleles controlling butterfly wing pattern variation: insights from gene expression in Heliconius numata
Heredity (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.