Abstract
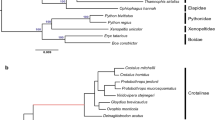
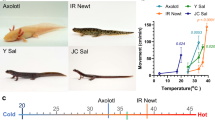
Vampire bats (Desmodus rotundus) are obligate blood feeders that have evolved specialized systems to suit their sanguinary lifestyle1,2,3. Chief among such adaptations is the ability to detect infrared radiation as a means of locating hotspots on warm-blooded prey. Among vertebrates, only vampire bats, boas, pythons and pit vipers are capable of detecting infrared radiation1,4. In each case, infrared signals are detected by trigeminal nerve fibres that innervate specialized pit organs on the animal’s face5,6,7,8,9,10. Thus, vampire bats and snakes have taken thermosensation to the extreme by developing specialized systems for detecting infrared radiation. As such, these creatures provide a window into the molecular and genetic mechanisms underlying evolutionary tuning of thermoreceptors in a species-specific or cell-type-specific manner. Previously, we have shown that snakes co-opt a non-heat-sensitive channel, vertebrate TRPA1 (transient receptor potential cation channel A1), to produce an infrared detector6. Here we show that vampire bats tune a channel that is already heat-sensitive, TRPV1, by lowering its thermal activation threshold to about 30 °C. This is achieved through alternative splicing of TRPV1 transcripts to produce a channel with a truncated carboxy-terminal cytoplasmic domain. These splicing events occur exclusively in trigeminal ganglia, and not in dorsal root ganglia, thereby maintaining a role for TRPV1 as a detector of noxious heat in somatic afferents. This reflects a unique organization of the bat Trpv1 gene that we show to be characteristic of Laurasiatheria mammals (cows, dogs and moles), supporting a close phylogenetic relationship with bats. These findings reveal a novel molecular mechanism for physiological tuning of thermosensory nerve fibres.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Gene Expression Omnibus
Data deposits
Deep sequencing data are archived under GEO accession number GSE28243. GenBank accession numbers are JN006855 (D. rotundus TRPV1-S), JN006856 (D. rotundus TRPV1-L), JN006857 (D. rotundus TRPA1), JN006858 (C. brevicauda TRPA1), JN006859 (C. brevicauda TRPV1-L), JN006860 (C. brevicauda TRPV1-S), JN006861 (Scapanus orarius TRPV1-L), JN006862 (S. orarius TRPV1-S), JN006863 (Pteropus rodricensis intron), JN006864 (D. rotundus intron), JN006865 (C. brevicauda intron), JN006866 (P. vampyrus intron), JN006867 (Rousettus aegyptiacus intron) and JN006868 (S. orarius intron).
References
Kurten, L. & Schmidt, U. Thermoperception in the common vampire bat (Desmodus rotundus). J. Comp. Physiol. 146, 223–228 (1982)
Schutt, B. Dark Banquet: Blood and the Curious Lives of Blood-Feeding Creatures (Three Rivers Press, 2008)
Tellgren-Roth, A. et al. Keeping the blood flowing — plasminogen activator genes and feeding behavior in vampire bats. Naturwissenschaften 96, 39–47 (2009)
Campbell, A. L., Naik, R. R., Sowards, L. & Stone, M. O. Biological infrared imaging and sensing. Micron 33, 211–225 (2002)
Bullock, T. H. & Cowles, R. B. Physiology of an infrared receptor: the facial pit of pit vipers. Science 115, 541–543 (1952)
Gracheva, E. O. et al. Molecular basis of infrared detection by snakes. Nature 464, 1006–1011 (2010)
Kurten, L., Schmidt, U. & Schafer, K. Warm and cold receptors in the nose of the vampire bat Desmodus rotundus . Naturwissenschaften 71, 327–328 (1984)
Molenaar, G. J. The sensory trigeminal system of a snake in the possession of infrared receptors. II. The central projections of the trigeminal nerve. J. Comp. Neurol. 179, 137–151 (1978)
Bakken, G. S. & Krochmal, A. R. The imaging properties and sensitivity of the facial pits of pitvipers as determined by optical and heat-transfer analysis. J. Exp. Biol. 210, 2801–2810 (2007)
Safer, A. B. & Grace, M. S. Infrared imaging in vipers: differential responses of crotaline and viperine snakes to paired thermal targets. Behav. Brain Res. 154, 55–61 (2004)
Kishida, R., Goris, R. C., Terashima, S. & Dubbeldam, J. L. A suspected infrared-recipient nucleus in the brainstem of the vampire bat, Desmodus rotundus . Brain Res. 322, 351–355 (1984)
Schafer, K., Braun, H. A. & Kurten, L. Analysis of cold and warm receptor activity in vampire bats and mice. Pflugers Arch. 412, 188–194 (1988)
Caterina, M. J. et al. The capsaicin receptor: a heat-activated ion channel in the pain pathway. Nature 389, 816–824 (1997)
Jordt, S. E., McKemy, D. D. & Julius, D. Lessons from peppers and peppermint: the molecular logic of thermosensation. Curr. Opin. Neurobiol. 13, 487–492 (2003)
Ramsey, I. S., Delling, M. & Clapham, D. E. An introduction to TRP channels. Annu. Rev. Physiol. 68, 619–647 (2006)
Black, D. L. Mechanisms of alternative pre-messenger RNA splicing. Annu. Rev. Biochem. 72, 291–336 (2003)
Will, C. L. & Luhrmann, R. Splicing of a rare class of introns by the U12-dependent spliceosome. Biol. Chem. 386, 713–724 (2005)
Murphy, W. J. et al. Molecular phylogenetics and the origins of placental mammals. Nature 409, 614–618 (2001)
Murphy, W. J. et al. Resolution of the early placental mammal radiation using Bayesian phylogenetics. Science 294, 2348–2351 (2001)
Nishihara, H., Hasegawa, M. & Okada, N. Pegasoferae, an unexpected mammalian clade revealed by tracking ancient retroposon insertions. Proc. Natl Acad. Sci. USA 103, 9929–9934 (2006)
Asher, R. J., Bennett, N. & Lehmann, T. The new framework for understanding placental mammal evolution. Bioessays 31, 853–864 (2009)
Prasad, A. B., Allard, M. W. & Green, E. D. Confirming the phylogeny of mammals by use of large comparative sequence data sets. Mol. Biol. Evol. 25, 1795–1808 (2008)
Pettigrew, J. D. et al. Phylogenetic relations between microbats, megabats and primates (Mammalia: Chiroptera and Primates). Phil. Trans. R. Soc. Lond. B 325, 489–559 (1989)
Spence, R., Gerlach, G., Lawrence, C. & Smith, C. The behaviour and ecology of the zebrafish, Danio rerio . Biol. Rev. Camb. Phil. Soc. 83, 13–34 (2008)
Grabowski, P. J. & Black, D. L. Alternative RNA splicing in the nervous system. Prog. Neurobiol. 65, 289–308 (2001)
Prescott, E. D. & Julius, D. A modular PIP2 binding site as a determinant of capsaicin receptor sensitivity. Science 300, 1284–1288 (2003)
Chuang, H. H., Neuhausser, W. M. & Julius, D. The super-cooling agent icilin reveals a mechanism of coincidence detection by a temperature-sensitive TRP channel. Neuron 43, 859–869 (2004)
DeCoursey, T. E. & Cherny, V. V. Temperature dependence of voltage-gated H+ currents in human neutrophils, rat alveolar epithelial cells, and mammalian phagocytes. J. Gen. Physiol. 112, 503–522 (1998)
Notredame, C., Higgins, D. G. & Heringa, J. J. Mol. Biol. 302, 205–217 (2000)
Ronquist, F. & Huelsenbeck, J. P. MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003)
Acknowledgements
We thank Y. Kelly and J. Poblete for technical assistance, C. Sehnert for help with bovine tissue collection, M. Suzawa and H. Ingraham for providing zebrafish mRNA, A. Walsh for providing megabat blood samples, the Centro Técnico de Producción Socialista Florentino for providing access to Hato Piñero (Cojedes, Venezuela) for animal collection and J. Nassar for providing access to laboratory material required for specimen collection. This work was supported by a Ruth L. Kirschstein National Research Service Award (GM080853; N.T.I.), a Pathway to Independence Fellowship from the UCSF CVRI (E.O.G.), the Howard Hughes Medical Institute (J.S.W.), and grants from NIH, including P01 AG010770 (J.S.W.) and NS047723 and NS055299 (D.J.).
Author information
Authors and Affiliations
Contributions
E.O.G., J.F.C.-M. and N.T.I. designed and performed experiments and analysed data. N.T.I. and J.S.W. developed analytical tools and analysed data. J.A.G.-C., C.I.A. and C.M. collected bat species and obtained tissues for analysis. E.O.G., J.F.C.-M. and D.J. wrote the manuscript with discussion and contributions from all authors. J.S.W. and D.J. provided advice and guidance throughout.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
This file contains Supplementary Figures 1-15 with legends. (PDF 3132 kb)
Rights and permissions
About this article
Cite this article
Gracheva, E., Cordero-Morales, J., González-Carcacía, J. et al. Ganglion-specific splicing of TRPV1 underlies infrared sensation in vampire bats. Nature 476, 88–91 (2011). https://doi.org/10.1038/nature10245
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10245
This article is cited by
-
Alternative splicing and environmental adaptation in wild house mice
Heredity (2024)
-
The physiology of alternative splicing
Nature Reviews Molecular Cell Biology (2023)
-
Molecular mechanism of hyperactivation conferred by a truncation of TRPA1
Nature Communications (2023)
-
Two single-point mutations in Ankyrin Repeat one drastically change the threshold temperature of TRPV1
Nature Communications (2023)
-
Recent Progress of Bio-inspired Camouflage Materials: From Visible to Infrared Range
Chemical Research in Chinese Universities (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.