Abstract
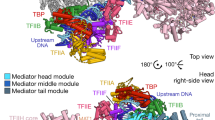
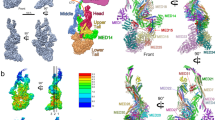
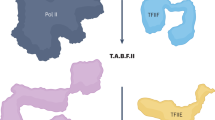
Mediator is a key regulator of eukaryotic transcription1, connecting activators and repressors bound to regulatory DNA elements with RNA polymerase II1,2,3,4 (Pol II). In the yeast Saccharomyces cerevisiae, Mediator comprises 25 subunits with a total mass of more than one megadalton (refs 5, 6) and is organized into three modules, called head, middle/arm and tail7,8,9. Our understanding of Mediator assembly and its role in regulating transcription has been impeded so far by limited structural information. Here we report the crystal structure of the essential Mediator head module (seven subunits, with a mass of 223 kilodaltons) at a resolution of 4.3 ångströms. Our structure reveals three distinct domains, with the integrity of the complex centred on a bundle of ten helices from five different head subunits. An intricate pattern of interactions within this helical bundle ensures the stable assembly of the head subunits and provides the binding sites for general transcription factors and Pol II. Our structural and functional data suggest that the head module juxtaposes transcription factor IIH and the carboxy-terminal domain of the largest subunit of Pol II, thereby facilitating phosphorylation of the carboxy-terminal domain of Pol II. Our results reveal architectural principles underlying the role of Mediator in the regulation of gene expression.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Kornberg, R. D. Mediator and the mechanism of transcriptional activation. Trends Biochem. Sci. 30, 235–239 (2005)
Conaway, R. C., Sato, S., Tomomori-Sato, C., Yao, T. & Conaway, J. W. The mammalian Mediator complex and its role in transcriptional regulation. Trends Biochem. Sci. 30, 250–255 (2005)
Boube, M., Joulia, L., Cribbs, D. L. & Bourbon, H. M. Evidence for a mediator of RNA polymerase II transcriptional regulation conserved from yeast to man. Cell 110, 143–151 (2002)
Malik, S. & Roeder, R. G. The metazoan Mediator co-activator complex as an integrative hub for transcriptional regulation. Nature Rev. Genet. 11, 761–772 (2010)
Bjorklund, S. & Gustafsson, C. M. The yeast Mediator complex and its regulation. Trends Biochem. Sci. 30, 240–244 (2005)
Guglielmi, B. et al. A high resolution protein interaction map of the yeast Mediator complex. Nucleic Acids Res. 32, 5379–5391 (2004)
Asturias, F. J., Jiang, Y. W., Myers, L. C., Gustafsson, C. M. & Kornberg, R. D. Conserved structures of mediator and RNA polymerase II holoenzyme. Science 283, 985–987 (1999)
Davis, J. A., Takagi, Y., Kornberg, R. D. & Asturias, F. A. Structure of the yeast RNA polymerase II holoenzyme: Mediator conformation and polymerase interaction. Mol. Cell 10, 409–415 (2002)
Cai, G., Imasaki, T., Takagi, Y. & Asturias, F. J. Mediator structural conservation and implications for the regulation mechanism. Structure 17, 559–567 (2009)
Takagi, Y. et al. Head module control of mediator interactions. Mol. Cell 23, 355–364 (2006)
Nonet, M. L. & Young, R. A. Intragenic and extragenic suppressors of mutations in the heptapeptide repeat domain of Saccharomyces cerevisiae RNA polymerase II. Genetics 123, 715–724 (1989)
Thompson, C. M., Koleske, A. J., Chao, D. M. & Young, R. A. A multisubunit complex associated with the RNA polymerase II CTD and TATA-binding protein in yeast. Cell 73, 1361–1375 (1993)
Thompson, C. M. & Young, R. A. General requirement for RNA polymerase II holoenzymes in vivo . Proc. Natl Acad. Sci. USA 92, 4587–4590 (1995)
Holstege, F. C. et al. Dissecting the regulatory circuitry of a eukaryotic genome. Cell 95, 717–728 (1998)
Takagi, Y. & Kornberg, R. D. Mediator as a general transcription factor. J. Biol. Chem. 281, 80–89 (2006)
Cai, G. et al. Mediator head module structure and functional interactions. Nature Struct. Mol. Biol. 17, 273–279 (2010)
Esnault, C. et al. Mediator-dependent recruitment of TFIIH modules in preinitiation complex. Mol. Cell 31, 337–346 (2008)
Larivière, L. et al. Structure and TBP binding of the Mediator head subcomplex Med8–Med18–Med20. Nature Struct. Mol. Biol. 13, 895–901 (2006)
Kim, Y. J., Bjorklund, S., Li, Y., Sayre, M. H. & Kornberg, R. D. A multiprotein mediator of transcriptional activation and its interaction with the C-terminal repeat domain of RNA polymerase II. Cell 77, 599–608 (1994)
Svejstrup, J. Q. et al. Evidence for a mediator cycle at the initiation of transcription. Proc. Natl Acad. Sci. USA 94, 6075–6078 (1997)
Max, T., Sogaard, M. & Svejstrup, J. Q. Hyperphosphorylation of the C-terminal repeat domain of RNA polymerase II facilitates dissociation of its complex with mediator. J. Biol. Chem. 282, 14113–14120 (2007)
Payne, J. M., Laybourn, P. J. & Dahmus, M. E. The transition of RNA polymerase II from initiation to elongation is associated with phosphorylation of the carboxyl-terminal domain of subunit IIa. J. Biol. Chem. 264, 19621–19629 (1989)
Kang, J. S. et al. The structural and functional organization of the yeast mediator complex. J. Biol. Chem. 276, 42003–42010 (2001)
Koleske, A. J., Buratowski, S., Nonet, M. & Young, R. A. A novel transcription factor reveals a functional link between the RNA polymerase II CTD and TFIID. Cell 69, 883–894 (1992)
Fitzgerald, D. J. et al. Protein complex expression by using multigene baculoviral vectors. Nature Methods 3, 1021–1032 (2006)
Li, M. Z. & Elledge, S. J. Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nature Methods 4, 251–256 (2007)
Tempst, P., Geromanos, S., Elicone, C. & Erdjument-Bromage, H. Improvements in microsequencer performance for low picomole sequence analysis. Methods 6, 248–261 (1994)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Adams, P. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Sheldrick, G. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008)
McCoy, A. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007)
Cowtan, K. Recent developments in classical density modification. Acta Crystallogr. D 66, 470–478 (2010)
Cowtan, K. Modified phased translation functions and their application to molecular-fragment location. Acta Crystallogr. D 54, 750–756 (1998)
Emsley, P., Lohkamp, B., Scott, W. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Schröder, G. F., Levitt, M. & Brunger, A. T. Super-resolution biomolecular crystallography with low-resolution data. Nature 464, 1218–1222 (2010)
Bryson, K. et al. Protein structure prediction servers at University College London. Nucleic Acids Res. 33, W36–W38 (2005)
Pettersen, E. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Stoffler, G. & Stoffler-Meilicke, M. The Ultrastructure of Macromolecular Complexes Studied with Antibodies 409–455 (De Gruyter, 1983)
Tischendorf, G. W., Zeichhardt, H. & Stoffler, G. Determination of the location of proteins L14, L17, L18, L19, L22, L23 on the surface of the 50S ribosomal subunit of Escherichia coli by immune electron microscopy. Mol. Gen. Genet. 134, 187–208 (1974)
Scheres, S. H. et al. Maximum-likelihood multi-reference refinement for electron microscopy images. J. Mol. Biol. 348, 139–149 (2005)
Sorzano, C. O. et al. XMIPP: a new generation of an open-source image processing package for electron microscopy. J. Struct. Biol. 148, 194–204 (2004)
Brignole, E. J., Smith, S. & Asturias, F. J. Conformational flexibility of metazoan fatty acid synthase enables catalysis. Nature Struct. Mol. Biol. 16, 190–197 (2009)
Acknowledgements
We thank L. Messerle for providing the Ta6Br14 metal cluster, T. Hurley for reading the manuscript, M. Georgiadis for discussions, C. Kaplan for his advice on yeast genetics, T. Earnest for giving us beam time and L. Fabrizio for assisting with the N-terminal sequence analysis. We thank the CCP4 summer school, funded by the NCI (Y1-CO-1020), and NIGMS (Y1-GM-1104) for their assistance with the twinning data analysis. Y.T. thanks the instructors on ‘The X-Ray Methods Course’ at Cold Spring Harbor Laboratory. This work was supported by US National Science Foundation grant MCB 0843026 (Y.T.); the American Heart Association 0735395N (Y.T.); a Human Frontier Science Program long-term fellowship (T.I.); NIH grants R01 GM67167 (F.J.A.) and GM36659 (R.D.K.); NCI Cancer Center Support Grant P30 CA08748 (to the MSKCC Microchemistry and Proteomics Core Laboratory); and the European Commission Framework Program 7 projects INSTRUCT and P-CUBE (I.B.). X-ray data were collected at the GM/CA-CAT at the Advanced Photon Source, Argonne National Laboratory. GM/CA-CAT is funded by the NIH (Y1-CO-1020 and Y1-GM-1104) and the Advanced Photon Source is supported by the DOE (DE-AC02-06CH11357). Portions of this research were carried out at the Stanford Synchrotron Radiation Lightsource, supported by the DOE and the NIH, and at the Advanced Light Source, supported by the DOE (DE-AC02-05CH11231).
Author information
Authors and Affiliations
Contributions
T.I., I.B. and Y.T. implemented the MultiBac system. T.I. was mainly responsible for protein complex preparation, crystallization, data collection, data analysis and model building in collaboration with Y.T. T.I., H.E.-B. and P.T. carried out mass spectroscopy analysis. G. Calero, G.L.K. and Y.T. carried out the initial crystallization and data collection, supervised by R.D.K.. Y.T., T.I. and F.C. designed and carried out expression of the mutant head modules and their biochemical characterization. Y.T. and K.Y. designed and carried out the yeast genetic experiment. Y.T. carried out the in vitro transcription assay and the CTD kinase assay; G. Cai, K.-L.T. and F.J.A. carried out the electron microscopy study on the head module and its mutants. T.I., F.J.A. and Y.T. discussed and interpreted all results. Y.T. supervised the X-ray, biochemical and yeast genetic work, and wrote the manuscript in collaboration with T.I., I.B., F.J.A. and R.D.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Text 1-7, Supplementary Tables 1-4, Supplementary Fgures 1-19 with legends and additional references. (PDF 2568 kb)
Rights and permissions
About this article
Cite this article
Imasaki, T., Calero, G., Cai, G. et al. Architecture of the Mediator head module. Nature 475, 240–243 (2011). https://doi.org/10.1038/nature10162
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10162
This article is cited by
-
The Mediator complex as a master regulator of transcription by RNA polymerase II
Nature Reviews Molecular Cell Biology (2022)
-
Structure of the human Mediator–RNA polymerase II pre-initiation complex
Nature (2021)
-
Crystal structure of human Mediator subunit MED23
Nature Communications (2018)
-
Transcription regulation by the Mediator complex
Nature Reviews Molecular Cell Biology (2018)
-
Anaerobic crystallization of proteins
Biophysical Reviews (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.