Abstract
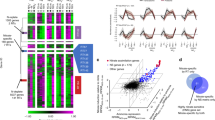
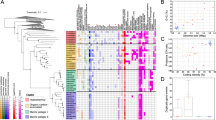
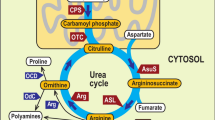
Diatoms dominate the biomass of phytoplankton in nutrient-rich conditions and form the basis of some of the world’s most productive marine food webs1,2,3,4. The diatom nuclear genome contains genes with bacterial and plastid origins as well as genes of the secondary endosymbiotic host (the exosymbiont5)1,6,7,8,9,10, yet little is known about the relative contribution of each gene group to diatom metabolism. Here we show that the exosymbiont-derived ornithine-urea cycle, which is similar to that of metazoans but is absent in green algae and plants, facilitates rapid recovery from prolonged nitrogen limitation. RNA-interference-mediated knockdown of a mitochondrial carbamoyl phosphate synthase impairs the response of nitrogen-limited diatoms to nitrogen addition. Metabolomic analyses indicate that intermediates in the ornithine-urea cycle are particularly depleted and that both the tricarboxylic acid cycle and the glutamine synthetase/glutamate synthase cycles are linked directly with the ornithine-urea cycle. Several other depleted metabolites are generated from ornithine-urea cycle intermediates by the products of genes laterally acquired from bacteria. This metabolic coupling of bacterial- and exosymbiont-derived proteins seems to be fundamental to diatom physiology because the compounds affected include the major diatom osmolyte proline12 and the precursors for long-chain polyamines required for silica precipitation during cell wall formation11. So far, the ornithine-urea cycle is only known for its essential role in the removal of fixed nitrogen in metazoans. In diatoms, this cycle serves as a distribution and repackaging hub for inorganic carbon and nitrogen and contributes significantly to the metabolic response of diatoms to episodic nitrogen availability. The diatom ornithine-urea cycle therefore represents a key pathway for anaplerotic carbon fixation into nitrogenous compounds that are essential for diatom growth and for the contribution of diatoms to marine productivity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Falkowski, P. G. et al. The evolution of modern eukaryotic phytoplankton. Science 305, 354–360 (2004)
Falkowski, P. G. & Oliver, M. J. Mix and match: how climate selects phytoplankton. Nature Rev. Microbiol. 5, 813–819 (2007)
Nelson, D. M., Treguer, P., Brzezinski, M. A., Leynaert, A. & Queguiner, B. Production and dissolution of biogenic silica in the ocean - revised global estimates, comparison with regional data and relationship to biogenic sedimentation. Glob. Biogeochem. Cycles 9, 359–372 (1995)
Smetacek, V. Diatoms and the ocean carbon cycle. Protist 150, 25–32 (1999)
Hamm, C. & Smetacek, V. in Evolution of Primary Producers in the Sea (eds Falkowski, P. G. & Knoll, A. H. ) (Academic Press, 2007)
Allen, A. E., Vardi, A. & Bowler, C. An ecological and evolutionary context for integrated nitrogen metabolism and related signaling pathways in marine diatoms. Curr. Opin. Plant Biol. 9, 264–273 (2006)
Bowler, C. et al. The Phaeodactylum genome reveals the evolutionary history of diatom genomes. Nature 456, 239–244 (2008)
Moustafa, A. et al. Genomic footprints of a cryptic plastid endosymbiosis in diatoms. Science 324, 1724–1726 (2009)
Bowler, C., Vardi, A. & Allen, A. E. Oceanographic and biogeochemical insights from diatom genomes. Ann. Rev. Mar. Sci. 2, 333–365 (2010)
Allen, A. E. et al. Whole-cell response of the pennate diatom Phaeodactylum tricornutum to iron starvation. Proc. Natl Acad. Sci. USA 105, 10438–10443 (2008)
Kröger, N. & Poulsen, N. Diatoms—from cell wall biogenesis to nanotechnology. Annu. Rev. Genet. 42, 83–107 (2008)
Krell, A., Funck, D., Plettner, I., John, U. & Dieckmann, G. Regulation of proline metabolism under salt stress in the psychrophilic diatom Fragilariopsis cylindrus (Bacillariophyceae). J. Phycol. 43, 753–762 (2007)
Anderson, P. M. Glutamine- and N-acetylglutamate-dependent carbamoyl phosphate synthetase in elasmobranchs. Science 208, 291–293 (1980)
Hong, J., Salo, W. L., Lusty, C. J. & Anderson, P. M. Carbamoyl-phosphate synthetase-III, an evolutionary intermediate in the transition between glutamine-dependent and ammonia-dependent carbamoyl-phosphate synthetases. J. Mol. Biol. 243, 131–140 (1994)
Mommsen, T. P. & Walsh, P. J. Evolution of urea synthesis in vertebrates: the piscine connection. Science 243, 72–75 (1989)
Lawson, F. S., Charlebois, R. L. & Dillon, J. A. R. Phylogenetic analysis of carbamoylphosphate synthetase genes: Complex evolutionary history includes an internal duplication within a gene which can root the tree of life. Mol. Biol. Evol. 13, 970–977 (1996)
Guppy, M. The hibernating bear: why is it so hot, and why does it cycle urea through the gut. Trends Biochem. Sci. 11, 274–276 (1986)
Holden, H. M., Thoden, J. B. & Raushel, F. M. Carbamoyl phosphate synthetase: an amazing biochemical odyssey from substrate to product. Cell. Mol. Life Sci. 56, 507–522 (1999)
Armbrust, E. V. et al. The genome of the diatom Thalassiosira pseudonana: Ecology, evolution, and metabolism. Science 306, 79–86 (2004)
Ast, M. et al. Diatom plastids depend on nucleotide import from the cytosol. Proc. Natl Acad. Sci. USA 106, 3621–3626 (2009)
Maheswari, U. et al. Digital expression profiling of novel diatom transcripts provides insight into their biological functions. Genome Biol. 11, R85 (2010)
Esteban-Pretel, G. et al. Vitamin A deficiency increases protein catabolism and induces urea cycle enzymes in rats. J. Nutr. 140, 792–798 (2010)
Lee, B. et al. In vivo urea cycle flux distinguishes and correlates with phenotypic severity in disorders of the urea cycle. Proc. Natl Acad. Sci. USA 97, 8021–8026 (2000)
De Riso, V. et al. Gene silencing in the marine diatom Phaeodactylum tricornutum . Nucleic Acids Res. 37, e96 (2009)
Young, E. B. & Beardall, J. Photosynthetic function in Dunaliella tertiolecta (Chlorophyta) during a nitrogen starvation recovery cycle. J. Phycol. 39, 897–905 (2003)
Morris, S. M. Regulation of enzymes of the urea cycle and arginine metabolism. Annu. Rev. Nutr. 22, 87–105 (2002)
Nunes-Nesi, A., Fernie, A. R. & Stitt, M. Metabolic and signaling aspects underpinning the regulation of plant carbon nitrogen interactions. Mol. Plant 3, 973–96 (2010)
Parker, M. S., Mock, T. & Armbrust, E. V. Genomic insights into marine microalgae. Annu. Rev. Genet. 42, 619–645 (2008)
Lisec, J., Schauer, N., Kopka, J., Willmitzer, L. & Fernie, A. R. Gas chromatography mass spectrometry-based metabolite profiling in plants. Nature Protocols 1, 387–396 (2006)
Stamatakis, A. RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and models. Bioinformatics 22, 2688–2690 (2006)
Siaut, M. et al. Molecular toolbox for studying diatom biology in Phaeodactylum tricornutum . Gene 406, 23–35 (2007)
Falciatore, A., Casotti, R., Leblanc, C., Abrescia, C. & Bowler, C. Transformation of nonselectable reporter genes in marine diatoms. Mar. Biotechnol. 1, 239–251 (1999)
Schauer, N. et al. GC-MS libraries for the rapid identification of metabolites in complex biological samples. FEBS Lett. 579, 1332–1337 (2005)
R Development Core Team R: A language and environment for statistical computing. ISBN 3-900051-07-0 (R Foundation for Statistical Computing, 2008)
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002)
Gouy, M., Guindon, S. & Gascuel, O. SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building. Mol. Biol. Evol. 27, 221–224 (2010)
Schmidt, H. A., Strimmer, K., Vingron, M. & von Haeseler, A. TREE-PUZZLE: a maximum likelihood phylogenetic analysis using quartets and parallel computing. Bioinformatics 18, 502–504 (2002)
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003)
Lartillot, N. & Philippe, H. A Bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Mol. Biol. Evol. 21, 1095–1109 (2004)
Quang, L. S., Gascuel, O. & Lartillot, N. Empirical profile mixture models for phylogenetic reconstruction. Bioinformatics 24, 2317–2323 (2008)
Anisimova, M. & Gascuel, O. Approximate likelihood-ratio test for branches: A fast, accurate, and powerful alternative. Syst. Biol. 55, 539–552 (2006)
Huelsenbeck, J. P. & Ronquist, F. MRBAYES: Bayesian inference of phylogeny. Bioinformatics 17, 754–755 (2001)
Ronquist, F. & Huelsenbeck, J. P. MRBAYES 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003)
Lartillot, N., Lepgae, T. & Blanquart, S. PhyloBayes 3: a Bayesian software package for phylogenetic reconstruction and molecular dating. Bioinformatics 25, 2286–2288 (2009)
Sanderson, M. J. A nonparametric approach to estimating divergence times in the absence of rate consistency. Mol. Biol. Evol. 19, 1218–1231 (1997)
Sanderson, M. J. Estimating absolute rates of molecular evolution and divergence times: a penalized likelihood approach. Mol. Biol. Evol. 19, 101–109 (2002)
Thorne, J. L., Kishino, H. & Painter, I. S. Estimating the rate of evolution of the rate of molecular evolution. Mol. Biol. Evol. 15, 1647–1657 (1998)
Berney, C. & Pawlowski, J. A molecular timescale for eukaryote evolution recalibrated with the continuous microfossil record. Proc. R. Soc. A/B 273, 1867–1872 (2006)
Benton, M. J. & Donoghue, P. C. J. Paleontological evidence to date the tree of life. Mol. Biol. Evol. 24, 26–53 (2007)
Acknowledgements
We thank A. Meichenin and C. Lichtlé for assistance with electron microscropy, J. C. Thomas for CPS purification and activity experiments, A. Falciatore for advice on RNAi constructs and A. Main for screening and evaluation of RNAi lines. This study was supported by the National Science Foundation (NSF-OCE-0722374, NSF-OCE-0727997, NSF-MCB-1024913) and JCVI internal funding to A.E.A., the European Commission Diatomics project and Agence Nationale de la Recherche (France) (C.B.) and the Czech Science Foundation (206/08/1423) (M.O.).
Author information
Authors and Affiliations
Contributions
A.E.A. and C.B. designed the study. A.E.A. performed CPS localization, confocal microscopy, protein purification, activity, overexpression and other laboratory experiments. A.E.A. and C.L.D. designed nitrogen-recovery experiments and the physiological characterization of the RNAi as well as wild-type experiments, which were performed by H.Z and C.L.D.. H.Z. generated and screened RNAi lines. H.Z. and H.H. performed long-term and short-term nitrogen-recovery and related qPCR experiments. D.A.J. ran qPCR reactions and assisted with analyses of qPCR data. M.O., A.H. and A.E.A. generated and analysed phylogenetic and molecular clock data. A.N-N. and A.R.F. performed metabolite profiling of samples collected from RNAi and wild-type cultures. J.P.M., C.L.D. and A.E.A. analysed qPCR, metabolite and western blot data in detail. A.E.A. wrote the paper with assistance from C.L.D., A.R.F., C.B. and M.O. All the authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-15 with legends and Supplementary Tables 1-5. (PDF 1770 kb)
Rights and permissions
About this article
Cite this article
Allen, A., Dupont, C., Oborník, M. et al. Evolution and metabolic significance of the urea cycle in photosynthetic diatoms. Nature 473, 203–207 (2011). https://doi.org/10.1038/nature10074
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature10074
This article is cited by
-
Bacterial transcriptional response to labile exometabolites from photosynthetic picoeukaryote Micromonas commoda
ISME Communications (2023)
-
Nutrient removal and biogas upgrade using co-cultivation of Chlorella vulgaris and three different bacteria under various GR24 concentrations by induction with 5-deoxystrigol
World Journal of Microbiology and Biotechnology (2023)
-
Review of phenotypic response of diatoms to salinization with biotechnological relevance
Hydrobiologia (2023)
-
Improving the genome and proteome annotations of the marine model diatom Thalassiosira pseudonana using a proteogenomics strategy
Marine Life Science & Technology (2023)
-
Early onset of urea synthesis and ammonia detoxification pathways in three terrestrially developing frogs
Journal of Comparative Physiology B (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.