Abstract
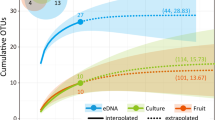
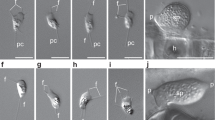
Fungi are the principal degraders of biomass in terrestrial ecosystems and establish important interactions with plants and animals1,2,3. However, our current understanding of fungal evolutionary diversity is incomplete4 and is based upon species amenable to growth in culture1. These culturable fungi are typically yeast or filamentous forms, bound by a rigid cell wall rich in chitin. Evolution of this body plan was thought critical for the success of the Fungi, enabling them to adapt to heterogeneous habitats and live by osmotrophy: extracellular digestion followed by nutrient uptake5. Here we investigate the ecology and cell biology of a previously undescribed and highly diverse form of eukaryotic life that branches with the Fungi, using environmental DNA analyses combined with fluorescent detection via DNA probes. This clade is present in numerous ecosystems including soil, freshwater and aquatic sediments. Phylogenetic analyses using multiple ribosomal RNA genes place this clade with Rozella, the putative primary branch of the fungal kingdom1. Tyramide signal amplification coupled with group-specific fluorescence in situ hybridization reveals that the target cells are small eukaryotes of 3–5 μm in length, capable of forming a microtubule-based flagellum. Co-staining with cell wall markers demonstrates that representatives from the clade do not produce a chitin-rich cell wall during any of the life cycle stages observed and therefore do not conform to the standard fungal body plan5. We name this highly diverse clade the cryptomycota in anticipation of formal classification.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
James, T. Y. et al. Reconstructing the early evolution of Fungi using a six-gene phylogeny. Nature 443, 818–822 (2006)
Pirozynski, K. A. & Malloch, D. W. The origin of land plants: a matter of mycotrophism. Biosystems 6, 153–164 (1975)
Wang, B. & Qiu, Y. L. Phylogenetic distribution and evolution of mycorrhizas in land plants. Mycorrhiza 16, 299–363 (2006)
Hawksworth, D. L. The magnitude of fungal diversity: the 1.5 million species estimate revisited. Mycol. Res. 105, 1422–1432 (2001)
Bartnicki-Garcia, S. Evolutionary Biology of the Fungi (eds Rayner, A. D. M., Brasier, C. M. & Moore, D. ) 389–403 (Cambridge University Press, 1987)
van Hannen, E. J., Mooij, W., van Agterveld, M. P., Gons, H. J. & Laanbroek, H. J. Detritus-dependent development of the microbial community in an experimental system: qualitative analysis by denaturing gradient gel electrophoresis. Appl. Environ. Microbiol. 65, 2478–2484 (1999)
Lepère, C., Domaizon, I. & Debroas, D. Unexpected importance of potential parasites in the composition of freshwater small-eukaryote community. Appl. Environ. Microbiol. 74, 2940–2949 (2008)
Lefèvre, E. et al. Unveiling fungal zooflagellates as members of freshwater picoeukaryotes: evidence from a molecular diversity study in a deep meromictic lake. Environ. Microbiol. 9, 61–71 (2007)
Amaral Zettler, L. A., Nerad, T., O'Kelly, C. J. & Sogin, M. L. The nucleariid amoebae: more protists at the animal-fungal boundary. J. Eukaryot. Microbiol. 48, 293–297 (2001)
Lara, E., Moreira, D. & Lopez-Garcia, P. Environmental clade LKM11 and Rozella form the deepest branching clade of Fungi. Protist 161, 116–121 (2010)
Massana, R. & Pedrós-Alió, C. Unveiling new microbial eukaryotes in the surface ocean. Curr. Opin. Microbiol. 11, 213–218 (2008)
Bass, D., Richards, T. A., Matthai, L., Marsh, V. & Cavalier-Smith, T. DNA evidence for global dispersal and probable endemicity of protozoa. BMC Evol. Biol. 7, 162 (2007)
Fuchs, B. M., Glöckner, F. O., Wulf, J. & Amann, R. Unlabeled helper oligonucleotides increase the in situ accessibility to 16S rRNA of fluorescently labeled oligonucleotide probes. Appl. Environ. Microbiol. 66, 3603–3607 (2000)
Kim, E. et al. Newly identified and diverse plastid-bearing branch on the eukaryotic tree of life. Proc. Natl Acad. Sci. USA 108, 1496–1500 (2011)
Mangot, J. F., Lepère, C., Bouvier, C., Debroas, D. & Domaizon, I. Community structure and dynamics of small eukaryotes targeted by new oligonucleotide probes: new insight into the lacustrine microbial food web. Appl. Environ. Microbiol. 75, 6373–6381 (2009)
Woods, A. et al. Definition of individual components within the cytoskeleton of Trypanosoma brucei by a library of monoclonal antibodies. J. Cell Sci. 93, 491–500 (1989)
Webster, J. & Weber, W. S. Introduction to Fungi 3rd edn (Cambridge University Press, 2007)
Held, A. A. The zoospore of Rozella allomycis: ultrastructure. Can. J. Bot. 53, 2212–2232 (1975)
Held, A. A. Rozella and Rozellopsis: naked endoparasitic fungi which dress up as their hosts. Bot. Rev. 47, 451–515 (1981)
Bulawa, C. E. Genetics and molecular biology of chitin synthesis in fungi. Annu. Rev. Microbiol. 47, 505–534 (1993)
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5, 113 (2004)
Galtier, N., Gouy, M. & Gautier, C. SEAVIEW and PHYLO_WIN: two graphic tools for sequence alignment and molecular phylogeny. Comput. Appl. Biosci. 12, 543–548 (1996)
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003)
Lockhart, P. J., Steel, M. A., Hendy, M. D. & Penny, D. Recovering evolutionary trees under a more realistic model of sequence evolution. Mol. Biol. Evol. 11, 605–612 (1994)
Ronquist, F. & Huelsenbeck, J. P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574 (2003)
Bass, D. et al. Yeast forms dominate fungal diversity in the deep oceans. Proc. R. Soc. Lond. B 274, 3069–3077 (2007)
Vandenkoornhuyse, P., Baldauf, S. L., Leyval, C., Straczek, J. & Young, J. P. W. Extensive fungal diversity in plant roots. Science 295, 2051 (2002)
Ludwig, W. et al. ARB: a software environment for sequence data. Nucleic Acids Res. 32, 1363–1371 (2004)
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35, 7188–7196 (2007)
Benson, D. A., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J. & Wheeler, D. L. GenBank. Nucleic Acids Res. 34, D16–20 (2006)
James, T. Y. et al. A molecular phylogeny of the flagellated fungi (Chytridiomycota) and description of a new phylum (Blastocladiomycota). Mycologia 98, 860–871 (2006)
Keane, T. M. et al. Assessment of methods for amino acid matrix selection and their use on empirical data shows that ad hoc assumptions for choice of matrix are not justified. BMC Evol. Biol. 6, 29 (2004)
Lanave, C., Preparata, G., Saccone, C. & Serio, G. A new method for calculating evolutionary substitution rates. J. Mol. Evol. 20, 86–93 (1984)
Foster, P. G. & Hickey, D. A. Compositional bias may affect both DNA-based and protein-based phylogenetic reconstructions. J. Mol. Evol. 48, 284–290 (1999)
Swofford, D. L. PAUP*. Phylogenetic Analysis Using Parsimony (*and other methods). Version 4 (Sinauer Associates, 2002)
Baschien, C., Manz, W., Neu, T. R. & Szewzyk, U. Fluorescence in situ hybrization of freshwater fungi. Int. Rev. Hydrobiol. 86, 371–381 (2001)
Acknowledgements
We thank: N. J. Talbot for advice, K. Gull for the TAT1 antibody, L. Guillou for access to curated SSU database and the Broad Institute of the Massachusetts Institute of Technology and Harvard for making their Rhizopus and Batrachochytrium genome sequence data publicly available. T.A.R. thanks the Leverhulme Trust for fellowship support. This work was primarily supported by a Natural Environment Research Council grant UK (NE/F011709/1). Additional support came from the Systematic Research Fund (awarded by the Systematics Association and the Linnean Society) to T.A.R., project FLAME (CGL2010-16304, MICINN) to R. M. and the BioMarKs project (European Funding Agencies from the ERA-net program BiodivERsA) to T.A.R. and R.M.
Author information
Authors and Affiliations
Contributions
This study was conceived by T.A.R. and M.D.M.J. with assistance from D.B. and R.M.. M.D.M.J. performed the molecular biology experiments with assistance from I.F. (FISH), C.G. (immunolocalization) and M.J.E. (microscopy). T.A.R. performed the bioinformatics and phylogenetic analysis. T.A.R. and M.D.M.J. wrote the paper with assistance from D.B. and R.M.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-7 with legends and Supplementary Tables 1-6. (PDF 23617 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Jones, M., Forn, I., Gadelha, C. et al. Discovery of novel intermediate forms redefines the fungal tree of life. Nature 474, 200–203 (2011). https://doi.org/10.1038/nature09984
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09984
This article is cited by
-
Effects of oxygen availability on mycobenthic communities of marine coastal sediments
Scientific Reports (2023)
-
Taxonomy, phylogeny and evolution of freshwater Hypocreomycetidae (Sordariomycetes)
Fungal Diversity (2023)
-
Intercomparison of Two Fluorescent Dyes to Visualize Parasitic Fungi (Chytridiomycota) on Phytoplankton
Microbial Ecology (2023)
-
Evolution of fungal phenotypic disparity
Nature Ecology & Evolution (2022)
-
Carbon assimilating fungi from surface ocean to subseafloor revealed by coupled phylogenetic and stable isotope analysis
The ISME Journal (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.