Abstract
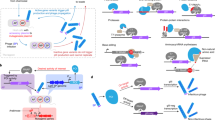
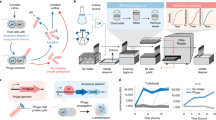
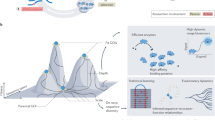
Laboratory evolution has generated many biomolecules with desired properties, but a single round of mutation, gene expression, screening or selection, and replication typically requires days or longer with frequent human intervention1. Because evolutionary success is dependent on the total number of rounds performed2, a means of performing laboratory evolution continuously and rapidly could dramatically enhance its effectiveness3. Although researchers have accelerated individual steps in the evolutionary cycle4,5,6,7,8,9, the only previous example of continuous directed evolution was the landmark study of Wright and Joyce10, who continuously evolved RNA ligase ribozymes with an in vitro replication cycle that unfortunately cannot be easily adapted to other biomolecules. Here we describe a system that enables the continuous directed evolution of gene-encoded molecules that can be linked to protein production in Escherichia coli. During phage-assisted continuous evolution (PACE), evolving genes are transferred from host cell to host cell through a modified bacteriophage life cycle in a manner that is dependent on the activity of interest. Dozens of rounds of evolution can occur in a single day of PACE without human intervention. Using PACE, we evolved T7 RNA polymerase (RNAP) variants that recognize a distinct promoter, initiate transcripts with ATP instead of GTP, and initiate transcripts with CTP. In one example, PACE executed 200 rounds of protein evolution over the course of 8 days. Starting from undetectable activity levels in two of these cases, enzymes with each of the three target activities emerged in less than 1 week of PACE. In all three cases, PACE-evolved polymerase activities exceeded or were comparable to that of the wild-type T7 RNAP on its wild-type promoter, representing improvements of up to several hundred-fold. By greatly accelerating laboratory evolution, PACE may provide solutions to otherwise intractable directed evolution problems and address novel questions about molecular evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Yuan, L., Kurek, I., English, J. & Keenan, R. Laboratory-directed protein evolution. Microbiol. Mol. Biol. Rev 69, 373–392 (2005)
Voigt, C. A., Kauffman, S. & Wang, Z. G. Rational evolutionary design: the theory of in vitro protein evolution. Adv. Protein Chem. 55, 79–160 (2000)
Mills, D. R., Peterson, R. L. & Spiegelman, S. An extracellular Darwinian experiment with a self-duplicating nucleic acid molecule. Proc. Natl Acad. Sci. USA 58, 217–224 (1967)
Wang, L., Jackson, W. C., Steinbach, P. A. & Tsien, R. Y. Evolution of new nonantibody proteins via iterative somatic hypermutation. Proc. Natl Acad. Sci. USA 101, 16745–16749 (2004)
Camps, M., Naukkarinen, J., Johnson, B. P. & Loeb, L. A. Targeted gene evolution in Escherichia coli using a highly error-prone DNA polymerase I. Proc. Natl Acad. Sci. USA 100, 9727–9732 (2003)
Makeyev, E. V. & Bamford, D. H. Evolutionary potential of an RNA virus. J. Virol. 78, 2114–2120 (2004)
Davis, J. N. & van den Pol, A. N. Viral mutagenesis as a means for generating novel proteins. J. Virol. 84, 1625–1630 (2009)
Das, A. T. et al. Viral evolution as a tool to improve the tetracycline-regulated gene expression system. J. Biol. Chem. 279, 18776–18782 (2004)
Wang, H. H. et al. Programming cells by multiplex genome engineering and accelerated evolution. Nature 460, 894–898 (2009)
Wright, M. C. & Joyce, G. F. Continuous in vitro evolution of catalytic function. Science 276, 614–617 (1997)
Husimi, Y. Selection and evolution of bacteriophages in cellstat. Adv. Biophys. 25, 1–43 (1989)
Smith, G. P. Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science 228, 1315–1317 (1985)
Riechmann, L. & Holliger, P. The C-terminal domain of TolA is the coreceptor for filamentous phage infection of E. coli . Cell 90, 351–360 (1997)
Nelson, F. K., Friedman, S. M. & Smith, G. P. Filamentous phage DNA cloning vectors: a noninfective mutant with a nonpolar deletion in gene III. Virology 108, 338–350 (1981)
Rakonjac, J. & Model, P. Roles of pIII in filamentous phage assembly. J. Mol. Biol. 282, 25–41 (1998)
Calendar, R. The Bacteriophages (Oxford Univ. Press, 2006)
Vidal, M. & Legrain, P. Yeast forward and reverse ‘n’-hybrid systems. Nucleic Acids Res. 27, 919–929 (1999)
Baker, K. et al. Chemical complementation: a reaction-independent genetic assay for enzyme catalysis. Proc. Natl Acad. Sci. USA 99, 16537–16542 (2002)
Fijalkowska, I. J. & Schaaper, R. M. Mutants in the Exo I motif of Escherichia coli dnaQ: defective proofreading and inviability due to error catastrophe. Proc. Natl Acad. Sci. USA 93, 2856–2861 (1996)
Opperman, T., Murli, S., Smith, B. T. & Walker, G. C. A model for a umuDC-dependent prokaryotic DNA damage checkpoint. Proc. Natl Acad. Sci. USA 96, 9218–9223 (1999)
Raskin, C. A., Diaz, G., Joho, K. & McAllister, W. T. Substitution of a single bacteriophage T3 residue in bacteriophage T7 RNA polymerase at position 748 results in a switch in promoter specificity. J. Mol. Biol. 228, 506–515 (1992)
Ikeda, R. A., Chang, L. L. & Warshamana, G. S. Selection and characterization of a mutant T7 RNA polymerase that recognizes an expanded range of T7 promoter-like sequences. Biochemistry 32, 9115–9124 (1993)
Raskin, C. A., Diaz, G. A. & McAllister, W. T. T7 RNA polymerase mutants with altered promoter specificities. Proc. Natl Acad. Sci. USA 90, 3147–3151 (1993)
Vidal-Aroca, F. et al. One-step high-throughput assay for quantitative detection of beta-galactosidase activity in intact Gram-negative bacteria, yeast, and mammalian cells. Biotechniques 40, 433–438 (2006)
Imburgio, D., Rong, M., Ma, K. & McAllister, W. T. Studies of promoter recognition and start site selection by T7 RNA polymerase using a comprehensive collection of promoter variants. Biochemistry 39, 10419–10430 (2000)
Brieba, L. G., Padilla, R. & Sousa, R. Role of T7 RNA polymerase His784 in start site selection and initial transcription. Biochemistry 41, 5144–5149 (2002)
Kuzmine, I., Gottlieb, P. A. & Martin, C. T. Binding of the priming nucleotide in the initiation of transcription by T7 RNA polymerase. J. Biol. Chem. 278, 2819–2823 (2003)
Cheetham, G. M., Jeruzalmi, D. & Steitz, T. A. Structural basis for initiation of transcription from an RNA polymerase-promoter complex. Nature 399, 80–83 (1999)
Martin, C. T. & Coleman, J. E. Kinetic analysis of T7 RNA polymerase-promoter interactions with small synthetic promoters. Biochemistry 26, 2690–2696 (1987)
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nature Methods 6, 343–345 (2009)
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA 97, 6640–6645 (2000)
Ichetovkin, I. E., Abramochkin, G. & Shrader, T. E. Substrate recognition by the leucyl/phenylalanyl-tRNA-protein transferase. Conservation within the enzyme family and localization to the trypsin-resistant domain. J Biol Chem 272, 33009–33014 (1997)
Acknowledgements
This work was supported by National Institutes of Health/National Institute of General Medical Sciences R01 GM065400 and by HHMI. K.M.E. acknowledges graduate research fellowships from the Hertz Foundation and the National Science Foundation. J.C.C. was supported by the Harvard Chemical Biology Graduate Program. We thank B. Dorr for assistance with phage generation modelling, E. Curtis for suggestions and V. D’Souza for plasmid pT7-911Q.
Author information
Authors and Affiliations
Contributions
K.M.E., J.C.C. and D.R.L. designed the experiments. K.M.E. designed and built the PACE apparatus. K.M.E. and J.C.C. performed the experiments. All authors analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Results, Supplementary Figures 1-12 with legends, Supplementary Tables 1-3 and additional references. (PDF 997 kb)
Rights and permissions
About this article
Cite this article
Esvelt, K., Carlson, J. & Liu, D. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011). https://doi.org/10.1038/nature09929
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09929
This article is cited by
-
Ultrahigh-throughput screening-assisted in vivo directed evolution for enzyme engineering
Biotechnology for Biofuels and Bioproducts (2024)
-
Enhancing prime editor activity by directed protein evolution in yeast
Nature Communications (2024)
-
T7 phage-assisted evolution of riboswitches using error-prone replication and dual selection
Scientific Reports (2024)
-
Advances in recombinant protease production: current state and perspectives
World Journal of Microbiology and Biotechnology (2024)
-
Engineered bacterial orthogonal DNA replication system for continuous evolution
Nature Chemical Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.