Abstract
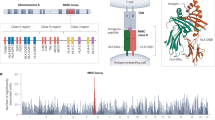
The HLA-C locus is distinct relative to the other classical HLA class I loci in that it has relatively limited polymorphism1, lower expression on the cell surface2,3, and more extensive ligand–receptor interactions with killer-cell immunoglobulin-like receptors4. A single nucleotide polymorphism (SNP) 35 kb upstream of HLA-C (rs9264942; termed −35) associates with control of HIV5,6,7, and with levels of HLA-C messenger RNA transcripts8 and cell-surface expression7, but the mechanism underlying its varied expression is unknown. We proposed that the −35 SNP is not the causal variant for differential HLA-C expression, but rather is marking another polymorphism that directly affects levels of HLA-C7. Here we show that variation within the 3′ untranslated region (UTR) of HLA-C regulates binding of the microRNA hsa-miR-148 to its target site, resulting in relatively low surface expression of alleles that bind this microRNA and high expression of HLA-C alleles that escape post-transcriptional regulation. The 3′ UTR variant associates strongly with control of HIV, potentially adding to the effects of genetic variation encoding the peptide-binding region of the HLA class I loci. Variation in HLA-C expression adds another layer of diversity to this highly polymorphic locus that must be considered when deciphering the function of these molecules in health and disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Zemmour, J. & Parham, P. Distinctive polymorphism at the HLA-C locus: implications for the expression of HLA-C. J. Exp. Med. 176, 937–950 (1992)
McCutcheon, J. A., Gumperz, J., Smith, K. D., Lutz, C. T. & Parham, P. Low HLA-C expression at cell surfaces correlates with increased turnover of heavy chain mRNA. J. Exp. Med. 181, 2085–2095 (1995)
Snary, D., Barnstable, C. J., Bodmer, W. F. & Crumpton, M. J. Molecular structure of human histocompatibility antigens: the HLA-C series. Eur. J. Immunol. 7, 580–585 (1977)
Bashirova, A. A., Martin, M. P., McVicar, D. W. & Carrington, M. The killer immunoglobulin-like receptor gene cluster: tuning the genome for defense. Annu. Rev. Genomics Hum. Genet. 7, 277–300 (2006)
Fellay, J. et al. A whole-genome association study of major determinants for host control of HIV-1. Science 317, 944–947 (2007)
International HIV Controllers Study The major genetic determinants of HIV-1 control affect HLA class I peptide presentation. Science 310, 1551–1557 (2010)
Thomas, R. et al. HLA-C cell surface expression and control of HIV/AIDS correlate with a variant upstream of HLA-C . Nature Genet. 41, 1290–1294 (2009)
Stranger, B. E. et al. Genome-wide associations of gene expression variation in humans. PLoS Genet. 1, e78 (2005)
Lewis, B. P., Burge, C. B. & Bartel, D. P. Conserved seed pairing, often flanked by adenosines, indicates that thousands of human genes are microRNA targets. Cell 120, 15–20 (2005)
Giraldez, A. J. et al. Zebrafish MiR-430 promotes deadenylation and clearance of maternal mRNAs. Science 312, 75–79 (2006)
Lim, L. P. et al. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 433, 769–773 (2005)
Wu, L., Fan, J. & Belasco, J. G. MicroRNAs direct rapid deadenylation of mRNA. Proc. Natl Acad. Sci. USA 103, 4034–4039 (2006)
Corrah, T. W. et al. A reappraisal of the relationship between the HIV-1-protective single nucleotide polymorphism 35 kb upstream of the HLA-C gene and surface HLA-C expression. J. Virol. 85, 3367–3374 (2011)
Schaefer, M. R. et al. A novel trafficking signal within the HLA-C cytoplasmic tail allows regulated expression upon differentiation of macrophages. J. Immunol. 180, 7804–7817 (2008)
Kralovicova, J. & Vorechovsky, I. Position-dependent repression and promotion of DQB1 intron 3 splicing by GGGG motifs. J. Immunol. 176, 2381–2388 (2006)
Krangel, M. S. Secretion of HLA-A and -B antigens via an alternative RNA splicing pathway. J. Exp. Med. 163, 1173–1190 (1986)
Castelli, E. C. et al. In silico analysis of microRNAs targeting the HLA-G 3′ untranslated region alleles and haplotypes. Hum. Immunol. 70, 1020–1025 (2009)
Tan, Z. et al. Allele-specific targeting of microRNAs to HLA-G and risk of asthma. Am. J. Hum. Genet. 81, 829–834 (2007)
Apps, R., Gardner, L. & Moffett, A. A critical look at HLA-G. Trends Immunol. 29, 313–321 (2008)
Phair, J. et al. Acquired immune deficiency syndrome occurring within 5 years of infection with human immunodeficiency virus type-1: the Multicenter AIDS Cohort Study. J. Acquir. Immune Defic. Syndr. 5, 490–496 (1992)
Emu, B. et al. Phenotypic, functional, and kinetic parameters associated with apparent T-cell control of human immunodeficiency virus replication in individuals with and without antiretroviral treatment. J. Virol. 79, 14169–14178 (2005)
Li, H., Wright, P. W. & Anderson, S. K. Identification and analysis of novel transcripts and promoters in the human killer cell immunoglobulin-like receptor (KIR) genes. Methods Mol. Biol. 612, 377–391 (2010)
Braud, V. M., Allan, D. S., Wilson, D. & McMichael, A. J. TAP- and tapasin-dependent HLA-E surface expression correlates with the binding of an MHC class I leader peptide. Curr. Biol. 8, 1–10 (1998)
Tilanus, M. G. J. in Immunobiology of the Human MHC. Proceedings of the 13th International Histocompatibility Workshop and Conference (ed. Hansen, J. ) Vol. 1, 304–416 (IHWG Press, 2002)
Shimizu, Y. & DeMars, R. Production of human cells expressing individual transferred HLA-A,-B,-C genes using an HLA-A,-B,-C null human cell line. J. Immunol. 142, 3320–3328 (1989)
Graham, F. L., Smiley, J., Russell, W. C. & Nairn, R. Characteristics of a human cell line transformed by DNA from human adenovirus type 5. J. Gen. Virol. 36, 59–72 (1977)
Setini, A. et al. Distinctive features of the α1-domain alpha helix of HLA-C heavy chains free of β2-microglobulin. Hum. Immunol. 46, 69–81 (1996)
Acknowledgements
This project has been funded in part with federal funds from the National Cancer Institute, National Institutes of Health (NIH), under contracts HHSN261200800001E, N02-CP-55504, R01-DA04334 and R01-DA12568. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the US Government. This research was supported in part by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research and the Cancer Inflammation Program Project Award for the year 2009, a grant from the Bill & Melinda Gates Foundation as part of the Collaboration for AIDS Vaccine Discovery, and the Mark and Lisa Schwartz Foundation. We would also like to thank the patients and investigators involved in the Multicenter AIDS Cohort Study (the MACS is funded by the National Institute of Allergy and Infectious Diseases, with supplemental funding from the National Cancer Institute and the National Heart, Lung and Blood Institute (grants UO1-AI-35042, 5-MO1-RR-00722 (GCRC), UO1-AI-35043, UO1-AI-37984, UO1-AI-35039, UO1-AI-35040, UO1-AI-37613 and UO1-AI-35041), the Swiss HIV Cohort Study (see Supplementary Note 4 for the list of members), supported by the Swiss National Science Foundation grant number 33CSC0-108787, and the SCOPE study, which was funded by the UL1 RR024131 (Clinical and Translational Sciences Award) and P30 AI27763 (Center for AIDS Research) grants. We thank S. Anderson, G. O’Connor and R. Thomas for advice, A. Kronfli and K. Ramakrishnan for assistance in plasmid and genomic DNA preparations, A. McFarland for western blots, V. Braud for the DT9 antibody, R. Johnson and G. Nelson for statistical advice and T. Covell for administrative assistance.
Author information
Authors and Affiliations
Contributions
S.K. and R.S. performed and evaluated the miRNA experiments. S.K., R.S., H.A.Y. and M.C. designed the study. M.C. directed the study. S.K., R.S. and M.C. wrote the manuscript. X.G., Y.Y., S.B. and M.M. genotyped HLA. Statistical analysis was performed by Y.Q. The clinical samples and data were contributed to by P.H., S.G.D., D.D., A.T., D.G., S.W., F.P. and B.W. Intellectual input was provided by all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-10 with legends, Supplementary Tables 1-4, Supplementary Notes and additional references. (PDF 2205 kb)
Rights and permissions
About this article
Cite this article
Kulkarni, S., Savan, R., Qi, Y. et al. Differential microRNA regulation of HLA-C expression and its association with HIV control. Nature 472, 495–498 (2011). https://doi.org/10.1038/nature09914
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09914
This article is cited by
-
HBV genotype-dependent association of HLA variants with the serodecline of HBsAg in chronic hepatitis B patients
Scientific Reports (2023)
-
Cumulative effects of weakly repressive regulatory regions in the 3’ UTR maintain PD-1 expression homeostasis in mammals
Communications Biology (2023)
-
Comparison between qPCR and RNA-seq reveals challenges of quantifying HLA expression
Immunogenetics (2023)
-
Haplotype structures and polymorphisms of dog leukocyte antigen (DLA) class I loci shaped by intralocus and interlocus recombination events
Immunogenetics (2022)
-
Single-cell RNA-sequencing reveals distinct immune cell subsets and signaling pathways in IgA nephropathy
Cell & Bioscience (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.