Abstract
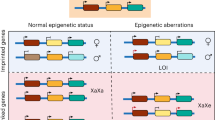
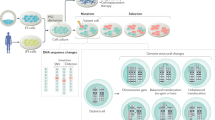
Induced pluripotent stem cells (iPSCs) offer immense potential for regenerative medicine and studies of disease and development. Somatic cell reprogramming involves epigenomic reconfiguration, conferring iPSCs with characteristics similar to embryonic stem (ES) cells. However, it remains unknown how complete the reestablishment of ES-cell-like DNA methylation patterns is throughout the genome. Here we report the first whole-genome profiles of DNA methylation at single-base resolution in five human iPSC lines, along with methylomes of ES cells, somatic cells, and differentiated iPSCs and ES cells. iPSCs show significant reprogramming variability, including somatic memory and aberrant reprogramming of DNA methylation. iPSCs share megabase-scale differentially methylated regions proximal to centromeres and telomeres that display incomplete reprogramming of non-CG methylation, and differences in CG methylation and histone modifications. Lastly, differentiation of iPSCs into trophoblast cells revealed that errors in reprogramming CG methylation are transmitted at a high frequency, providing an iPSC reprogramming signature that is maintained after differentiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Primary accessions
Sequence Read Archive
Data deposits
Analysed datasets can be browsed and downloaded from http://neomorph.salk.edu/ips_methylomes. Sequence data for MethylC-Seq, RNA-Seq and Chip-Seq experiments have been submitted to the NCBI SRA database under the accession numbers SRA023829.2 and SRP000941.
References
Soldner, F. et al. Parkinson’s disease patient-derived induced pluripotent stem cells free of viral reprogramming factors. Cell 136, 964–977 (2009)
Yamanaka, S. A fresh look at iPS cells. Cell 137, 13–17 (2009)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007)
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007)
Park, I. et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature 451, 141–146 (2008)
Yu, J. et al. Human induced pluripotent stem cells free of vector and transgene sequences. Science 324, 797–801 (2009)
Zhao, X. Y. et al. iPS cells produce viable mice through tetraploid complementation. Nature 461, 86–90 (2009)
Boland, M. J. et al. Adult mice generated from induced pluripotent stem cells. Nature 461, 91–94 (2009)
Guenther, M. G. et al. Chromatin structure and gene expression programs of human embryonic and induced pluripotent stem cells. Cell Stem Cell 7, 249–257 (2010)
Deng, J. et al. Targeted bisulfite sequencing reveals changes in DNA methylation associated with nuclear reprogramming. Nature Biotechnol. 27, 353–360 (2009)
Doi, A. et al. Differential methylation of tissue- and cancer-specific CpG island shores distinguishes human induced pluripotent stem cells, embryonic stem cells and fibroblasts. Nature Genet. 41, 1350–1353 (2009)
Kim, K. et al. Epigenetic memory in induced pluripotent stem cells. Nature 467, 285–290 (2010)
Polo, J. M. et al. Cell type of origin influences the molecular and functional properties of mouse induced pluripotent stem cells. Nature Biotechnol. 28, 848–855 (2010)
Stadtfeld, M. et al. Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature 465, 175–181 (2010)
Miura, K. et al. Variation in the safety of induced pluripotent stem cell lines. Nature Biotechnol. 27, 743–745 (2009)
Hu, B. Y. et al. Neural differentiation of human induced pluripotent stem cells follows developmental principles but with variable potency. Proc. Natl Acad. Sci. USA 107, 4335–4340 (2010)
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009)
Sugii, S. et al. Human and mouse adipose-derived cells support feeder-independent induction of pluripotent stem cells. Proc. Natl Acad. Sci. USA 108, 3558–3563 (2010)
Daley, G. et al. Broader implications of defining standards for the pluripotency of iPSCs. Cell Stem Cell 4, 200–201 (2009)
Xu, R. H. et al. BMP4 initiates human embryonic stem cell differentiation to trophoblast. Nature Biotechnol. 20, 1261–1264 (2002)
Cedar, H. & Bergman, Y. Linking DNA methylation and histone modification: patterns and paradigms. Nature Rev. Genet. 10, 295–304 (2009)
Chodavarapu, R. K. et al. Relationship between nucleosome positioning and DNA methylation. Nature 466, 388–392 (2010)
Laurent, L. et al. Dynamic changes in the human methylome during differentiation. Genome Res. 20, 320–331 (2010)
Hawkins, R. D. et al. Distinct epigenomic landscapes of pluripotent and lineage-committed human cells. Cell Stem Cell 6, 479–491 (2010)
Erhardt, A. et al. TMEM132D, a new candidate for anxiety phenotypes: evidence from human and mouse studies. Mol. Psychiatry advance online publication. 10.1038/mp.2010.4 (6 April 2010)
Yilmaz, G., Alexander, J. S., Erkuran Yilmaz, C. & Granger, D. N. Induction of neuro-protective/regenerative genes in stem cells infiltrating post-ischemic brain tissue. Exp. Transl. Stroke Med. 2, 11 (2010)
Li, M. Z. et al. Molecular mapping of developing dorsal horn-enriched genes by microarray and dorsal/ventral subtractive screening. Dev. Biol. 292, 555–564 (2006)
Chin, M. H. et al. Induced pluripotent stem cells and embryonic stem cells are distinguished by gene expression signatures. Cell Stem Cell 5, 111–123 (2009)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009)
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nature Biotechnol. 28, 511–515 (2010)
Ludwig, T. et al. Feeder-independent culture of human embryonic stem cells. Nature Methods 3, 637–646 (2006)
Ludwig, T. et al. Derivation of human embryonic stem cells in defined conditions. Nature Biotechnol. 24, 185–187 (2006)
O’Malley, R. C., Alonso, J. M., Kim, C. J., Leisse, T. J. & Ecker, J. R. An adapter ligation-mediated PCR method for high-throughput mapping of T-DNA inserts in the Arabidopsis genome. Nature Protocols 2, 2910–2917 (2007)
Chen, P., Cokus, S. J. & Pellegrini, M. B. S. Seeker: precise mapping for bisulfite sequencing. BMC Bioinformatics 11, 203 (2010)
Lister, R. et al. Highly integrated single-base resolution maps of the epigenome in Arabidopsis. Cell 133, 523–536 (2008)
Acknowledgements
We thank L. Zhang and G. Schroth for assistance with MethylC-Seq library sequencing. R.L. is supported by a California Institute for Regenerative Medicine Training Grant. M.P. is supported by a Catharina Foundation postdoctoral fellowship. R.D.H. is supported by an American Cancer Society Postdoctoral Fellowship. Y.K. is supported by the Japan Society for the Promotion of Science. This work was supported by grants from the following: Mary K. Chapman Foundation, the National Science Foundation (NSF) (NSF 0726408), the National Institutes of Health (NIH) (U01 ES017166, U01 1U01ES017166-01, DK062434), the California Institute for Regenerative Medicine (RB2-01530), the Morgridge Institute for Research and the Howard Hughes Medical Institute. We thank the NIH Roadmap Reference Epigenome Consortium (http://www.roadmapepigenomics.org/). This study was carried out as part of the NIH Roadmap Epigenomics Program.
Author information
Authors and Affiliations
Contributions
Experiments were designed by R.L., J.R.E., R.M.E., B.R., J.A.T., Y.S.K., R.Y., M.D. and R.D.H. Cells were grown by J.A.-B. and Y.S.K. MethylC-Seq and RNA-Seq experiments were conducted by R.L. and J.R.N. ChIP-Seq experiments were conducted by R.D.H. ChIP-Seq data analysis was performed by G.H., S.K. and R.D.H. Retroviral insertion site localization experiments were performed by R.O’M. and R.C. Sequencing data processing was performed by R.L. and G.H. Bioinformatic and statistical analyses were conducted by M.P., R.L. and G.H. R.S. performed data interpretation analyses. The manuscript was prepared by R.L., M.P. and J.R.E.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Figures
The file contains Supplementary Figures 1-17 with legends (PDF 9437 kb)
Supplementary Tables
The file contains Supplementary Tables 1-7 (XLS 178 kb)
Rights and permissions
About this article
Cite this article
Lister, R., Pelizzola, M., Kida, Y. et al. Hotspots of aberrant epigenomic reprogramming in human induced pluripotent stem cells. Nature 471, 68–73 (2011). https://doi.org/10.1038/nature09798
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09798
This article is cited by
-
Epigenetic regulation and factors that influence the effect of iPSCs-derived neural stem/progenitor cells (NS/PCs) in the treatment of spinal cord injury
Clinical Epigenetics (2024)
-
The Regenerative Microenvironment of the Tissue Engineering for Urethral Strictures
Stem Cell Reviews and Reports (2024)
-
Global effects of identity and aging on the human sperm methylome
Clinical Epigenetics (2023)
-
Single-cell DNA methylation sequencing by combinatorial indexing and enzymatic DNA methylation conversion
Cell & Bioscience (2023)
-
Vascular organoids: unveiling advantages, applications, challenges, and disease modelling strategies
Stem Cell Research & Therapy (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.