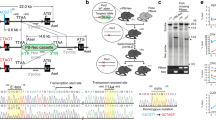
Abstract
Daily oscillations of gene expression underlie circadian behaviours in multicellular organisms1. While attention has been focused on transcriptional and post-translational mechanisms1,2,3, other post-transcriptional modes have been less clearly delineated. Here we report mutants of a novel Drosophila gene twenty-four (tyf) that show weak behavioural rhythms. Weak rhythms are accompanied by marked reductions in the levels of the clock protein Period (PER) as well as more modest effects on Timeless (TIM). Nonetheless, PER induction in pacemaker neurons can rescue tyf mutant rhythms. TYF associates with a 5′-cap-binding complex, poly(A)-binding protein (PABP), as well as per and tim transcripts. Furthermore, TYF activates reporter expression when tethered to reporter messenger RNA even in vitro. Taken together, these data indicate that TYF potently activates PER translation in pacemaker neurons to sustain robust rhythms, revealing a new and important role for translational control in the Drosophila circadian clock.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Doherty, C. J. & Kay, S. A. Circadian control of global gene expression patterns. Annu. Rev. Genet. 44, 419–444 (2010)
Zheng, X. & Sehgal, A. Probing the relative importance of molecular oscillations in the circadian clock. Genetics 178, 1147–1155 (2008)
Harms, E., Kivimae, S., Young, M. W. & Saez, L. Posttranscriptional and posttranslational regulation of clock genes. J. Biol. Rhythms 19, 361–373 (2004)
Dubruille, R. & Emery, P. A plastic clock: how circadian rhythms respond to environmental cues in Drosophila . Mol. Neurobiol. 38, 129–145 (2008)
Stoleru, D., Peng, Y., Agosto, J. & Rosbash, M. Coupled oscillators control morning and evening locomotor behaviour of Drosophila . Nature 431, 862–868 (2004)
Grima, B., Chelot, E., Xia, R. & Rouyer, F. Morning and evening peaks of activity rely on different clock neurons of the Drosophila brain. Nature 431, 869–873 (2004)
Renn, S. C., Park, J. H., Rosbash, M., Hall, J. C. & Taghert, P. H. A pdf neuropeptide gene mutation and ablation of PDF neurons each cause severe abnormalities of behavioral circadian rhythms in Drosophila . Cell 99, 791–802 (1999)
Peng, Y., Stoleru, D., Levine, J. D., Hall, J. C. & Rosbash, M. Drosophila free-running rhythms require intercellular communication. PLoS Biol. 1, e13 (2003)
Lin, Y., Stormo, G. D. & Taghert, P. H. The neuropeptide pigment-dispersing factor coordinates pacemaker interactions in the Drosophila circadian system. J. Neurosci. 24, 7951–7957 (2004)
Zeng, H., Hardin, P. E. & Rosbash, M. Constitutive overexpression of the Drosophila period protein inhibits period mRNA cycling. EMBO J. 13, 3590–3598 (1994)
Park, J. H. et al. Differential regulation of circadian pacemaker output by separate clock genes in Drosophila . Proc. Natl Acad. Sci. USA 97, 3608–3613 (2000)
Yang, Z. & Sehgal, A. Role of molecular oscillations in generating behavioral rhythms in Drosophila . Neuron 29, 453–467 (2001)
Frisch, B., Hardin, P. E., Hamblen-Coyle, M. J., Rosbash, M. & Hall, J. C. A promoterless period gene mediates behavioral rhythmicity and cyclical per expression in a restricted subset of the Drosophila nervous system. Neuron 12, 555–570 (1994)
Kim, E. Y., Ko, H. W., Yu, W., Hardin, P. E. & Edery, I. A. DOUBLETIME kinase binding domain on the Drosophila PERIOD protein is essential for its hyperphosphorylation, transcriptional repression, and circadian clock function. Mol. Cell. Biol. 27, 5014–5028 (2007)
Blanchardon, E. et al. Defining the role of Drosophila lateral neurons in the control of circadian rhythms in motor activity and eclosion by targeted genetic ablation and PERIOD protein overexpression. Eur. J. Neurosci. 13, 871–888 (2001)
Sonenberg, N. & Hinnebusch, A. G. Regulation of translation initiation in eukaryotes: mechanisms and biological targets. Cell 136, 731–745 (2009)
Kula-Eversole, E. et al. Surprising gene expression patterns within and between PDF-containing circadian neurons in Drosophila . Proc. Natl Acad. Sci. USA 107, 13497–13502 (2010)
Keryer-Bibens, C., Barreau, C. & Osborne, H. B. Tethering of proteins to RNAs by bacteriophage proteins. Biol. Cell 100, 125–138 (2008)
Veleri, S., Brandes, C., Helfrich-Forster, C., Hall, J. C. & Stanewsky, R. A self-sustaining, light-entrainable circadian oscillator in the Drosophila brain. Curr. Biol. 13, 1758–1767 (2003)
Staiger, D. & Koster, T. Spotlight on post-transcriptional control in the circadian system. Cell. Mol. Life Sci. doi:10.1007/s00018–010–0513–5. (in the press)
Dockendorff, T. C. et al. Drosophila lacking dfmr1 activity show defects in circadian output and fail to maintain courtship interest. Neuron 34, 973–984 (2002)
Sofola, O. et al. The Drosophila FMRP and LARK RNA-binding proteins function together to regulate eye development and circadian behavior. J. Neurosci. 28, 10200–10205 (2008)
Kadener, S. et al. A role for microRNAs in the Drosophila circadian clock. Genes Dev. 23, 2179–2191 (2009)
Nagoshi, E. et al. Dissecting differential gene expression within the circadian neuronal circuit of Drosophila . Nature Neurosci. 13, 60–68 (2010)
So, W. V. & Rosbash, M. Post-transcriptional regulation contributes to Drosophila clock gene mRNA cycling. EMBO J. 16, 7146–7155 (1997)
Stanewsky, R., Jamison, C. F., Plautz, J. D., Kay, S. A. & Hall, J. C. Multiple circadian-regulated elements contribute to cycling period gene expression in Drosophila . EMBO J. 16, 5006–5018 (1997)
Majercak, J., Sidote, D., Hardin, P. E. & Edery, I. How a circadian clock adapts to seasonal decreases in temperature and day length. Neuron 24, 219–230 (1999)
Suri, V., Lanjuin, A. & Rosbash, M. TIMELESS-dependent positive and negative autoregulation in the Drosophila circadian clock. EMBO J. 18, 675–686 (1999)
Suri, V., Hall, J. C. & Rosbash, M. Two novel doubletime mutants alter circadian properties and eliminate the delay between RNA and protein in Drosophila . J. Neurosci. 20, 7547–7555 (2000)
Green, C. B. et al. Loss of Nocturnin, a circadian deadenylase, confers resistance to hepatic steatosis and diet-induced obesity. Proc. Natl Acad. Sci. USA 104, 9888–9893 (2007)
Sharma, Y., Cheung, U., Larsen, E. W. & Eberl, D. F. PPTGAL, a convenient GAL4 P-element vector for testing expression of enhancer fragments in Drosophila . Genesis 34, 115–118 (2002)
Lykke-Andersen, J., Shu, M. D. & Steitz, J. A. Human Upf proteins target an mRNA for nonsense-mediated decay when bound downstream of a termination codon. Cell 103, 1121–1131 (2000)
Kaneko, M. & Hall, J. C. Neuroanatomy of cells expressing clock genes in Drosophila: transgenic manipulation of the period and timeless genes to mark the perikarya of circadian pacemaker neurons and their projections. J. Comp. Neurol. 422, 66–94 (2000)
Cheng, Y., Gvakharia, B. & Hardin, P. E. Two alternatively spliced transcripts from the Drosophila period gene rescue rhythms having different molecular and behavioral characteristics. Mol. Cell. Biol. 18, 6505–6514 (1998)
Zhao, J. et al. Drosophila clock can generate ectopic circadian clocks. Cell 113, 755–766 (2003)
Osterwalder, T., Yoon, K. S., White, B. H. & Keshishian, H. A conditional tissue-specific transgene expression system using inducible GAL4 . Proc. Natl Acad. Sci. USA 98, 12596–12601 (2001)
Groth, A. C., Fish, M., Nusse, R. & Calos, M. P. Construction of transgenic Drosophila by using the site-specific integrase from phage phiC31. Genetics 166, 1775–1782 (2004)
Markstein, M., Pitsouli, C., Villalta, C., Celniker, S. E. & Perrimon, N. Exploiting position effects and the gypsy retrovirus insulator to engineer precisely expressed transgenes. Nature Genet. 40, 476–483 (2008)
Lim, C. et al. Clockwork orange encodes a transcriptional repressor important for circadian-clock amplitude in Drosophila . Curr. Biol. 17, 1082–1089 (2007)
Lim, C. et al. Functional role of CREB-binding protein in the circadian clock system of Drosophila melanogaster . Mol. Cell. Biol. 27, 4876–4890 (2007)
Matsumoto, A. et al. A functional genomics strategy reveals clockwork orange as a transcriptional regulator in the Drosophila circadian clock. Genes Dev. 21, 1687–1700 (2007)
Houl, J. H., Yu, W., Dudek, S. M. & Hardin, P. E. Drosophila CLOCK is constitutively expressed in circadian oscillator and non-oscillator cells. J. Biol. Rhythms 21, 93–103 (2006)
Picot, M., Cusumano, P., Klarsfeld, A., Ueda, R. & Rouyer, F. Light activates output from evening neurons and inhibits output from morning neurons in the Drosophila circadian clock. PLoS Biol. 5, e315 (2007)
Eulalio, A., Behm-Ansmant, I., Schweizer, D. & Izaurralde, E. P-body formation is a consequence, not the cause, of RNA-mediated gene silencing. Mol. Cell. Biol. 27, 3970–3981 (2007)
Roy, G., Miron, M., Khaleghpour, K., Lasko, P. & Sonenberg, N. The Drosophila poly(A) binding protein-interacting protein, dPaip2, is a novel effector of cell growth. Mol. Cell. Biol. 24, 1143–1154 (2004)
Nakamura, A., Sato, K. & Hanyu-Nakamura, K. Drosophila cup is an eIF4E binding protein that associates with Bruno and regulates oskar mRNA translation in oogenesis. Dev. Cell 6, 69–78 (2004)
Monzo, K. et al. Fragile X mental retardation protein controls trailer hitch expression and cleavage furrow formation in Drosophila embryos. Proc. Natl Acad. Sci. USA 103, 18160–18165 (2006)
Satterfield, T. F. & Pallanck, L. J. Ataxin-2 and its Drosophila homolog, ATX2, physically assemble with polyribosomes. Hum. Mol. Genet. 15, 2523–2532 (2006)
Wessel, D. & Flugge, U. I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. 138, 141–143 (1984)
Thoma, C., Ostareck-Lederer, A. & Hentze, M. W. A poly(A) tail-responsive in vitro system for cap- or IRES-driven translation from HeLa cells. Methods Mol. Biol. 257, 171–180 (2004)
Castagnetti, S., Hentze, M. W., Ephrussi, A. & Gebauer, F. Control of oskar mRNA translation by Bruno in a novel cell-free system from Drosophila ovaries. Development 127, 1063–1068 (2000)
Acknowledgements
We thank I. Edery, J. Hall, H. Keshishian, M. Rosbash, F. Rouyer, A. Sehgal, the Bloomington Drosophila stock center, Harvard Exelixis Drosophila stock collection, KAIST GenExel Drosophila library and the National Institute of Genetics for Drosophila strains; P. Hardin, E. Izaurralde, A. Nakamura, M. Rosbash and N. Sonenberg for antibodies; J. Lykke-Andersen for plasmids; K. E. Duncan for suggestions on in vitro translation assays. This work was supported by grants from the Brain Research Center of the 21st Century Frontier Research Program through the National Research Foundation of Korea funded by the Ministry of Education, Science and Technology, the Republic of Korea (J.C.) and from the National Institutes of Health (R01NS059042, R01NS052903, R01MH067870; R.A.)
Author information
Authors and Affiliations
Contributions
R.A. and J.C. conceived the study; R.A., C.L. and J.C. designed the experiments; C.L. (under the supervision of R.A.) and J.L (under the supervision of J.C.) jointly completed Figs 1 and 2, Supplementary Figs 1, 4, 8, 14 and Supplementary Tables 2 and 3; J.L., S.M.P. and S.K.J. performed and analysed the experiments in Supplementary Fig. 13; J.L., C.C. and J.K. performed the genome-wide behavioural screen; C.C. performed GST pull-down studies in Supplementary Fig. 12a; V.L.K. performed PDF quantification analysis in Supplementary Fig. 5b; C.L. performed and analysed experiments in all remaining Figures, Supplementary Figures and Tables; C.L. and R.A. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures1-15 with legends and Supplementary Tables 1-5. (PDF 5471 kb)
Rights and permissions
About this article
Cite this article
Lim, C., Lee, J., Choi, C. et al. The novel gene twenty-four defines a critical translational step in the Drosophila clock. Nature 470, 399–403 (2011). https://doi.org/10.1038/nature09728
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09728
This article is cited by
-
Molecular mechanisms and physiological importance of circadian rhythms
Nature Reviews Molecular Cell Biology (2020)
-
Nuclear Envelope Protein MAN1 Regulates the Drosophila Circadian Clock via Period
Neuroscience Bulletin (2019)
-
A high-fat diet impacts memory and gene expression of the head in mated female Drosophila melanogaster
Journal of Comparative Physiology B (2019)
-
Molecular modulators of the circadian clock: lessons from flies and mice
Cellular and Molecular Life Sciences (2017)
-
CRTC Potentiates Light-independent timeless Transcription to Sustain Circadian Rhythms in Drosophila
Scientific Reports (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.