Abstract
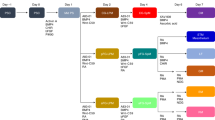
Studies in embryonic development have guided successful efforts to direct the differentiation of human embryonic and induced pluripotent stem cells (PSCs) into specific organ cell types in vitro1,2. For example, human PSCs have been differentiated into monolayer cultures of liver hepatocytes and pancreatic endocrine cells3,4,5,6 that have therapeutic efficacy in animal models of liver disease7,8 and diabetes9, respectively. However, the generation of complex three-dimensional organ tissues in vitro remains a major challenge for translational studies. Here we establish a robust and efficient process to direct the differentiation of human PSCs into intestinal tissue in vitro using a temporal series of growth factor manipulations to mimic embryonic intestinal development10. This involved activin-induced definitive endoderm formation11, FGF/Wnt-induced posterior endoderm pattering, hindgut specification and morphogenesis12,13,14, and a pro-intestinal culture system15,16 to promote intestinal growth, morphogenesis and cytodifferentiation. The resulting three-dimensional intestinal ‘organoids’ consisted of a polarized, columnar epithelium that was patterned into villus-like structures and crypt-like proliferative zones that expressed intestinal stem cell markers17. The epithelium contained functional enterocytes, as well as goblet, Paneth and enteroendocrine cells. Using this culture system as a model to study human intestinal development, we identified that the combined activity of WNT3A and FGF4 is required for hindgut specification whereas FGF4 alone is sufficient to promote hindgut morphogenesis. Our data indicate that human intestinal stem cells form de novo during development. We also determined that NEUROG3, a pro-endocrine transcription factor that is mutated in enteric anendocrinosis18, is both necessary and sufficient for human enteroendocrine cell development in vitro. PSC-derived human intestinal tissue should allow for unprecedented studies of human intestinal development and disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Mayhew, C. N. & Wells, J. M. Converting human pluripotent stem cells into β-cells: recent advances and future challenges. Curr. Opin. Organ Transplant. 15, 54–60 (2010)
Spence, J. R. & Wells, J. M. Translational embryology: using embryonic principles to generate pancreatic endocrine cells from embryonic stem cells. Dev. Dyn. 236, 3218–3227 (2007)
Cai, J. et al. Directed differentiation of human embryonic stem cells into functional hepatic cells. Hepatology 45, 1229–1239 (2007)
D’Amour, K. A. et al. Production of pancreatic hormone-expressing endocrine cells from human embryonic stem cells. Nature Biotechnol. 24, 1392–1401 (2006)
Song, Z. et al. Efficient generation of hepatocyte-like cells from human induced pluripotent stem cells. Cell Res. 19, 1233–1242 (2009)
Zhang, D. et al. Highly efficient differentiation of human ES cells and iPS cells into mature pancreatic insulin-producing cells. Cell Res. 19, 429–438 (2009)
Basma, H. et al. Differentiation and transplantation of human embryonic stem cell-derived hepatocytes. Gastroenterology 136, 990–999 (2008)
Touboul, T. et al. Generation of functional hepatocytes from human embryonic stem cells under chemically defined conditions that recapitulate liver development. Hepatology 51, 1754–1765 (2010)
Kroon, E. et al. Pancreatic endoderm derived from human embryonic stem cells generates glucose-responsive insulin-secreting cells in vivo . Nature Biotechnol. 26, 443–452 (2008)
Zorn, A. M. & Wells, J. M. Vertebrate endoderm development and organ formation. Annu. Rev. Cell Dev. Biol. 25, 221–251 (2009)
D’Amour, K. A. et al. Efficient differentiation of human embryonic stem cells to definitive endoderm. Nature Biotechnol. 23, 1534–1541 (2005)
Dessimoz, J., Opoka, R., Kordich, J. J., Grapin-Botton, A. & Wells, J. M. FGF signaling is necessary for establishing gut tube domains along the anterior-posterior axis in vivo . Mech. Dev. 123, 42–55 (2006)
McLin, V. A., Rankin, S. A. & Zorn, A. M. Repression of Wnt/β-catenin signaling in the anterior endoderm is essential for liver and pancreas development. Development 134, 2207–2217 (2007)
Wells, J. M. & Melton, D. A. Early mouse endoderm is patterned by soluble factors from adjacent germ layers. Development 127, 1563–1572 (2000)
Gracz, A. D., Ramalingam, S. & Magness, S. T. Sox9 expression marks a subset of CD24-expressing small intestine epithelial stem cells that form organoids in vitro . Am. J. Physiol. Gastrointest. Liver Physiol. 298, G590–G600 (2010)
Sato, T. et al. Single Lgr5 stem cells build crypt-villus structures in vitro without a mesenchymal niche. Nature 459, 262–265 (2009)
de Santa Barbara, P., van den Brink, G. R. & Roberts, D. J. Development and differentiation of the intestinal epithelium. Cell. Mol. Life Sci. 60, 1322–1332 (2003)
Wang, J. et al. Mutant neurogenin-3 in congenital malabsorptive diarrhea. N. Engl. J. Med. 355, 270–280 (2006)
Cao, L. et al. Intestinal lineage commitment of embryonic stem cells. Differentiationdoi:10.1016/j.diff.2010.09.182 (in the press)
Torihashi, S. et al. Gut-like structures from mouse embryonic stem cells as an in vitro model for gut organogenesis preserving developmental potential after transplantation. Stem Cells 24, 2618–2626 (2006)
van der Flier, L. G. et al. Transcription factor achaete scute-like 2 controls intestinal stem cell fate. Cell 136, 903–912 (2009)
Groneberg, D. A., Doring, F., Eynott, P. R., Fischer, A. & Daniel, H. Intestinal peptide transport: ex vivo uptake studies and localization of peptide carrier PEPT1. Am. J. Physiol. Gastrointest. Liver Physiol. 281, G697–G704 (2001)
Haveri, H. et al. Transcription factors GATA-4 and GATA-6 in normal and neoplastic human gastrointestinal mucosa. BMC Gastroenterol. 8, 9 (2008)
McLin, V. A., Henning, S. J. & Jamrich, M. The role of the visceral mesoderm in the development of the gastrointestinal tract. Gastroenterology 136, 2074–2091 (2009)
Ormestad, M. et al. Foxf1 and Foxf2 control murine gut development by limiting mesenchymal Wnt signaling and promoting extracellular matrix production. Development 133, 833–843 (2006)
Kosinski, C. et al. Indian hedgehog regulates intestinal stem cell fate through epithelial-mesenchymal interactions during development. Gastroenterology 139, 893–903 (2010)
Jenny, M. et al. Neurogenin3 is differentially required for endocrine cell fate specification in the intestinal and gastric epithelium. EMBO J. 21, 6338–6347 (2002)
Lee, C. S., Perreault, N., Brestelli, J. E. & Kaestner, K. H. Neurogenin 3 is essential for the proper specification of gastric enteroendocrine cells and the maintenance of gastric epithelial cell identity. Genes Dev. 16, 1488–1497 (2002)
Lopez-Diaz, L. et al. Intestinal Neurogenin 3 directs differentiation of a bipotential secretory progenitor to endocrine cell rather than goblet cell fate. Dev. Biol. 309, 298–305 (2007)
Ootani, A. et al. Sustained in vitro intestinal epithelial culture within a Wnt-dependent stem cell niche. Nature Med. 15, 701–706 (2009)
Zhou, Q., Brown, J., Kanarek, A., Rajagopal, J. & Melton, D. A. In vivo reprogramming of adult pancreatic exocrine cells to β-cells. Nature 455, 627–632 (2008)
Ludwig, T. E. et al. Feeder-independent culture of human embryonic stem cells. Nature Methods 3, 637–646 (2006)
Ludwig, T. E. et al. Derivation of human embryonic stem cells in defined conditions. Nature Biotechnol. 24, 185–187 (2006)
Lambert, P. F. et al. Using an immortalized cell line to study the HPV life cycle in organotypic “raft” cultures. Methods Mol. Med. 119, 141–155 (2005)
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007)
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006)
Richards, M., Tan, S. P., Tan, J. H., Chan, W. K. & Bongso, A. The transcriptome profile of human embryonic stem cells as defined by SAGE. Stem Cells 22, 51–64 (2004)
Thomson, J. A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998)
Acknowledgements
We thank members of the laboratory, D. Wiginton and C. Wylie for input. We also thank M. Kofron, T. Stefader and R. Lang for assistance with imaging. Vectors and antibodies were from D. Melton (Addgene no. 19410, 19413), S. Yamanaka (17217–17220), C. Baum (OCT4, KLF4, SOX4, MYC lenti), and I. Manabe (KLF5 antibody). This work was supported by the Juvenile Diabetes Research Foundation JDRF-2-2003-530 (J.M.W.) and NIH, R01GM072915 (J.M.W.); R01DK080823A1 and S1 (A.M.Z. and J.M.W.); R03 DK084167 and R01 CA142826 (N.F.S.), F32 DK83202-01 and T32 HD07463 (J.R.S.). We also acknowledge core support for viral vectors, microarrays (supported by P30 DK078392), karyotyping and the Pluripotent Stem Cell Facility (supported by U54 RR025216).
Author information
Authors and Affiliations
Contributions
J.M.W. and J.R.S. conceived the study and experimental design, performed and analysed experiments and co-wrote the manuscript. S.A.R., M.F.K. and J.E.V. performed experiments. C.N.M., M.F.K., K.T., V.V.K., J.E.V., E.E.H. and S.I.W. provided reagents, conceptual and/or technical support in generating and characterizing iPSC lines and intestinal organoids. N.F.S. and A.M.Z. provided additional conceptual and experimental support and co-funded the project. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-15 with legends, Supplementary Tables 1-3 and additional references. (PDF 15443 kb)
Rights and permissions
About this article
Cite this article
Spence, J., Mayhew, C., Rankin, S. et al. Directed differentiation of human pluripotent stem cells into intestinal tissue in vitro. Nature 470, 105–109 (2011). https://doi.org/10.1038/nature09691
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09691
This article is cited by
-
Functional intestinal monolayers from organoids derived from human iPS cells for drug discovery research
Stem Cell Research & Therapy (2024)
-
Chemically-defined and scalable culture system for intestinal stem cells derived from human intestinal organoids
Nature Communications (2024)
-
Human primordial germ cell-like cells specified from resetting precursors develop in human hindgut organoids
Nature Protocols (2024)
-
Insights on Three Dimensional Organoid Studies for Stem Cell Therapy in Regenerative Medicine
Stem Cell Reviews and Reports (2024)
-
Challenges and opportunities in inflammatory bowel disease: from current therapeutic strategies to organoid-based models
Inflammation Research (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.