Abstract
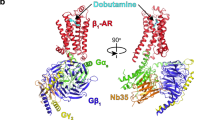
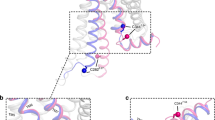
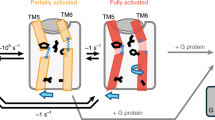
G-protein-coupled receptors (GPCRs) are eukaryotic integral membrane proteins that modulate biological function by initiating cellular signalling in response to chemically diverse agonists. Despite recent progress in the structural biology of GPCRs1, the molecular basis for agonist binding and allosteric modulation of these proteins is poorly understood. Structural knowledge of agonist-bound states is essential for deciphering the mechanism of receptor activation, and for structure-guided design and optimization of ligands. However, the crystallization of agonist-bound GPCRs has been hampered by modest affinities and rapid off-rates of available agonists. Using the inactive structure of the human β2 adrenergic receptor (β2AR) as a guide, we designed a β2AR agonist that can be covalently tethered to a specific site on the receptor through a disulphide bond. The covalent β2AR-agonist complex forms efficiently, and is capable of activating a heterotrimeric G protein. We crystallized a covalent agonist-bound β2AR–T4L fusion protein in lipid bilayers through the use of the lipidic mesophase method2, and determined its structure at 3.5 Å resolution. A comparison to the inactive structure and an antibody-stabilized active structure (companion paper3) shows how binding events at both the extracellular and intracellular surfaces are required to stabilize an active conformation of the receptor. The structures are in agreement with long-timescale (up to 30 μs) molecular dynamics simulations showing that an agonist-bound active conformation spontaneously relaxes to an inactive-like conformation in the absence of a G protein or stabilizing antibody.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Rosenbaum, D. M., Rasmussen, S. G. & Kobilka, B. K. The structure and function of G-protein-coupled receptors. Nature 459, 356–363 (2009)
Caffrey, M. Crystallizing membrane proteins for structure determination: use of lipidic mesophases. Annu. Rev. Biophys. 38, 29–51 (2009)
Rasmussen, S. G. F. et al. Structure of a nanobody-stabilized active state of the β2 adrenoceptor. Nature doi:10.1038/nature09648 (this issue)
Li, J., Edwards, P. C., Burghammer, M., Villa, C. & Schertler, G. F. Structure of bovine rhodopsin in a trigonal crystal form. J. Mol. Biol. 343, 1409–1438 (2004)
Palczewski, K. et al. Crystal structure of rhodopsin: a G protein-coupled receptor. Science 289, 739–745 (2000)
Park, J. H., Scheerer, P., Hofmann, K. P., Choe, H. W. & Ernst, O. P. Crystal structure of the ligand-free G-protein-coupled receptor opsin. Nature 454, 183–187 (2008)
Scheerer, P. et al. Crystal structure of opsin in its G-protein-interacting conformation. Nature 455, 497–502 (2008)
Deupi, X. & Kobilka, B. K. Energy landscapes as a tool to integrate GPCR structure, dynamics, and function. Physiology (Bethesda) 25, 293–303 (2010)
Dohlman, H. G., Caron, M. G., Strader, C. D., Amlaiky, N. & Lefkowitz, R. J. Identification and sequence of a binding site peptide of the β 2 adrenergic receptor. Biochemistry 27, 1813–1817 (1988)
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein coupled receptors. Meth. Neurosci. 25, 366–428 (1995)
Buck, E. & Wells, J. A. Disulfide trapping to localize small-molecule agonists and antagonists for a G protein-coupled receptor. Proc. Natl Acad. Sci. USA 102, 2719–2724 (2005)
Kolb, H. C., VanNieuwenhze, M. S. & Sharpless, K. B. Catalytic asymmetric dihydroxylation. Chem. Rev. 94, 2483–2547 (1994)
Martinelli, M. J. et al. Catalytic regioselective sulfonylation of α-chelatable alcohols: scope and mechanistic insight. J. Am. Chem. Soc. 124, 3578–3585 (2002)
Whorton, M. R. et al. A monomeric G protein-coupled receptor isolated in a high-density lipoprotein particle efficiently activates its G protein. Proc. Natl Acad. Sci. USA 104, 7682–7687 (2007)
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nature Protocols 4, 706–731 (2009)
Cherezov, V., Peddi, A., Muthusubramaniam, L., Zheng, Y. F. & Caffrey, M. A robotic system for crystallizing membrane and soluble proteins in lipidic mesophases. Acta Crystallogr. D 60, 1795–1807 (2004)
Rosenbaum, D. M. et al. GPCR engineering yields high-resolution structural insights into β2-adrenergic receptor function. Science 318, 1266–1273 (2007)
Tota, M. R., Candelore, M. R., Dixon, R. A. & Strader, C. D. Biophysical and genetic analysis of the ligand-binding site of the β-adrenoceptor. Trends Pharmacol. Sci. 12, 4–6 (1991)
Dror, R. O. et al. Identification of two distinct inactive conformations of the β2-adrenergic receptor reconciles structural and biochemical observations. Proc. Natl Acad. Sci. USA 106, 4689–4694 (2009)
Vilardaga, J. P., Bunemann, M., Krasel, C., Castro, M. & Lohse, M. J. Measurement of the millisecond activation switch of G protein-coupled receptors in living cells. Nature Biotechnol. 21, 807–812 (2003)
Vanni, S., Neri, M., Tavernelli, I. & Rothlisberger, U. A conserved protonation-induced switch can trigger “ionic-lock” formation in adrenergic receptors. J. Mol. Biol. 397, 1339–1349 (2010)
Ghanouni, P. et al. The effect of pH on β2 adrenoceptor function. Evidence for protonation-dependent activation. J. Biol. Chem. 275, 3121–3127 (2000)
Ghanouni, P. et al. Functionally different agonists induce distinct conformations in the G protein coupling domain of the β2 adrenergic receptor. J. Biol. Chem. 276, 24433–24436 (2001)
Ahuja, S. et al. Helix movement is coupled to displacement of the second extracellular loop in rhodopsin activation. Nature Struct. Mol. Biol. 16, 168–175 (2009)
Bokoch, M. P. et al. Ligand-specific regulation of the extracellular surface of a G-protein-coupled receptor. Nature 463, 108–112 (2010)
MacKerell, A. D., Jr et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998)
Shaw, D. E. et al. in Proceedings of the Conference on High Performance Computing, Networking, Storage and Analysis (ACM Press, 2009)
Yao, X. et al. Coupling ligand structure to specific conformational switches in the β2-adrenoceptor. Nature Chem. Biol. 2, 417–422 (2006)
Kozasa, T. & Gilman, A. G. Purification of recombinant G proteins from Sf9 cells by hexahistidine tagging of associated subunits. Characterization of α12 and inhibition of adenylyl cyclase by αz . J. Biol. Chem. 270, 1734–1741 (1995)
Kobilka, B. K. Amino and carboxyl terminal modifications to facilitate the production and purification of a G protein-coupled receptor. Anal. Biochem. 231, 269–271 (1995)
Chae, P. S. et al. Maltose–neopentyl glycol (MNG) amphiphiles for solubilization, stabilization and crystallization of membrane proteins. Nature Methods 7, 1003–1008 (2010)
Cheng, A., Hummel, B., Qiu, H. & Caffrey, M. A simple mechanical mixer for small viscous lipid-containing samples. Chem. Phys. Lipids 95, 11–21 (1998)
Leslie, A. G. The integration of macromolecular diffraction data. Acta Crystallogr. D 62, 48–57 (2006)
Collaborative Computational Project, number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Cryst. 40, 658–674 (2007)
Afonine, P. V., Grosse-Kunstleve, R. W. & Adams, P. D. A robust bulk-solvent correction and anisotropic scaling procedure. Acta Crystallogr. D 61, 850–855 (2005)
Blanc, E. et al. Refinement of severely incomplete structures with maximum likelihood in BUSTER-TNT . Acta Crystallogr. D 60, 2210–2221 (2004)
Weik, M. et al. Specific chemical and structural damage to proteins produced by synchrotron radiation. Proc. Natl Acad. Sci. USA 97, 623–628 (2000)
Rubenstein, R. C. et al. The hydrophobic tryptic core of the β-adrenergic receptor retains Gs regulatory activity in response to agonists and thiols. J. Biol. Chem. 262, 16655–16662 (1987)
Bowers, K. J. et al. in Proceedings of the 2006 ACM/IEEE Conference on Supercomputing (ACM Press, 2006)
Fahmy, K. et al. Protonation states of membrane-embedded carboxylic acid groups in rhodopsin and metarhodopsin II: a Fourier-transform infrared spectroscopy study of site-directed mutants. Proc. Natl Acad. Sci. USA 90, 10206–10210 (1993)
Vogel, R. et al. Functional role of the “ionic lock”–an interhelical hydrogen-bond network in family A heptahelical receptors. J. Mol. Biol. 380, 648–655 (2008)
Liapakis, G., Chan, W. C., Papadokostaki, M. & Javitch, J. A. Synergistic contributions of the functional groups of epinephrine to its affinity and efficacy at the β2 adrenergic receptor. Mol. Pharmacol. 65, 1181–1190 (2004)
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: an N⋅log(N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089 (1993)
Kräutler, V., van Gunsteren, W. F. & Hünenberger, P. H. A fast SHAKE algorithm to solve distance constraint equations for small molecules in molecular dynamics simulations. J. Comput. Chem. 22, 501–508 (2001)
Tuckerman, M., Berne, B. J. & Martyna, G. J. Reversible multiple time scale molecular dynamics. J. Chem. Phys. 97, 1990 (1992)
Shan, Y., Klepeis, J. L., Eastwood, M. P., Dror, R. O. & Shaw, D. E. Gaussian split Ewald: a fast Ewald mesh method for molecular simulation. J. Chem. Phys. 122, 054101 (2005)
Mackerell, A. D., Jr, Feig, M. & Brooks, C. L., III Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004)
Beglov, D. & Roux, B. Finite representation of an infinite bulk system: solvent boundary potential for computer simulations. J. Chem. Phys. 100, 9050–9063 (1994)
Klauda, J. B. et al. Update of the CHARMM all-atom additive force field for lipids: validation on six lipid types. J. Phys. Chem. B 114, 7830–7843 (2010)
Vanommeslaeghe, K. et al. CHARMM general force field: a force field for drug-like molecules compatible with the CHARMM all-atom additive biological force fields. J. Comput. Chem. 31, 671–690 (2010)
Tu, T. et al. in Proceedings of the 2008 ACM/IEEE Conference on Supercomputing (ACM Press, 2008)
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996)
Acknowledgements
We acknowledge support from National Institutes of Health Grants NS028471 and GM083118 (B.K.K.), GM56169 (W.I.W.), P01 GM75913 (S.H.G), GM75915 and P50GM073210 (M.C), and P60DK-20572 (R.K.S.), the Mathers Foundation (B.K.K and W.I.W), the Lundbeck Foundation (Junior Group Leader Fellowship, S.G.F.R.),Science Foundation Ireland (07/IN.1/B1836) and FP7 COST Action CM0902 (M.C.), the Bavaria California Technology Center (P.G.), and the University of Michigan Biomedical Sciences Scholars Program (R.K.S). We thank Stefan Löber, Harald Hübner, Albert Pan and Paul Maragakis for discussion and suggestions. We thank Foon Sun Thian for help with insect cell expression.
Author information
Authors and Affiliations
Contributions
D.M.R. designed project and agonists, did binding assays to characterize agonists, developed purification, optimized crystallization conditions, optimized construct, purified protein, grew and harvested crystals, collected data, solved structure, refined structure, wrote manuscript. C.Z. performed G protein activation assays for covalent agonist, prepared recombinant baculovirus, performed large-scale expression of recombinant β2AR in insect cells, purified protein, grew crystals. J.L. helped to optimize crystallization conditions, grew and harvested crystals, collected data. D.A. was involved in LCP optimization, harvested crystals, collected data. R.H. synthesized agonists. S.G.F.R. identified the use of MNG3 detergent for β2AR stabilization and assisted with manuscript preparation. H.J.C. assisted with data processing and refinement. R.K.S. and B.D. provided ApoA1 and Gs protein for functional characterization of the covalent ligand. P.S.C. and S.G. provided MNG3 detergent for stabilization of purified β2AR. D.H.A. and R.O.D. performed and analyzed MD simulations; R.O.D. and D.E.S. oversaw MD simulations and analysis. W.I.W. assisted with data processing and refinement and with manuscript preparation. M.C. helped to guide the LCP crystallization efforts at Stanford and in Ireland, and oversaw automated lipidic cubic phase crystallography screens. P.G. designed the strategy for the synthesis of the covalent agonist. B.K. was responsible for the overall project strategy and management, oversaw manuscript preparation, and assisted with synchrotron data collection.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-7 with legends, Supplementary Table 1, Supplementary Methods and Data and additional references. (PDF 2854 kb)
Rights and permissions
About this article
Cite this article
Rosenbaum, D., Zhang, C., Lyons, J. et al. Structure and function of an irreversible agonist-β2 adrenoceptor complex. Nature 469, 236–240 (2011). https://doi.org/10.1038/nature09665
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09665
This article is cited by
-
Dynamic Nature of Proteins is Critically Important for Their Function: GPCRs and Signal Transducers
Applied Magnetic Resonance (2024)
-
Constrained catecholamines gain β2AR selectivity through allosteric effects on pocket dynamics
Nature Communications (2023)
-
The dynamics of agonist-β2-adrenergic receptor activation induced by binding of GDP-bound Gs protein
Nature Chemistry (2023)
-
A Vaccinia-based system for directed evolution of GPCRs in mammalian cells
Nature Communications (2023)
-
2D SIFt: a matrix of ligand-receptor interactions
Journal of Cheminformatics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.