Abstract
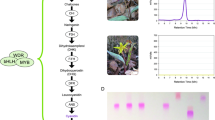
Species formation generates biological diversity and occurs when traits evolve that prevent gene flow between populations. Discerning the number and distribution of genes underlying these traits and, in a few cases, identifying the genes involved, has greatly enhanced our understanding over the past 15 years of species formation (reviewed by Noor and Feder1 and Wolf et al.2). However, this work has almost exclusively focused on traits that restrict gene flow between populations that have evolved as a by-product of genetic divergence between geographically isolated populations. By contrast, little is known about the characteristics of genes associated with reinforcement, the process by which natural selection directly favours restricted gene flow during the formation of species. Here we identify changes in two genes that appear to cause a flower colour change in Phlox drummondii, which previous work has shown contributes to reinforcement. Both changes involve cis-regulatory mutations to genes in the anthocyanin biosynthetic pathway (ABP). Because one change is recessive whereas the other is dominant, hybrid offspring produce an intermediate flower colour that is visited less by pollinators, and is presumably maladaptive. Thus genetic change selected to increase prezygotic isolation also appears to result in increased postzygotic isolation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Data deposits
The DNA sequences reported here are deposited in GenBank under accession numbers HQ127319–HQ127344 and HQ323688–HQ323691.
References
Noor, M. A. F. & Feder, J. L. Speciation genetics: evolving approaches. Nature Rev. Genet. 7, 851–861 (2006)
Wolf, J. B. W., Lindell, J. & Backstrom, N. Speciation genetics: current status and evolving approaches. Phil. Trans. R. Soc. B 365, 1717–1733 (2010)
Dobzhansky, T. Genetics and the Origin of Species (Columbia University Press, 1937)
Howard, D. J. in Hybrid Zones and the Evolutionary Process (ed. Harrison, R. G.) 46–69 (Oxford University Press, 1993)
Servedio, M. R. & Noor, M. A. F. The role of reinforcement in speciation: theory and data. Annu. Rev. Ecol. Evol. Syst. 34, 339–364 (2003)
Butlin, R. Speciation by reinforcement. Trends Ecol. Evol. 2, 8–13 (1987)
Felsenstein, J. Skepticism towards Santa Rosalia, or why are there so few kinds of animals. Evolution 35, 124–138 (1981)
Ortiz-Barrientos, D., Grealy, A. & Nosil, P. The genetics and ecology of reinforcement implications for the evolution of prezygotic isolation in sympatry and beyond. Year Evol. Biol. 2009, 156–182 (2009)
Pfennig, K. S. & Pfennig, D. W. Character displacement: ecological and reproductive responses to a common evolutionary problem. Q. Rev. Biol. 84, 253–276 (2009)
Ortiz-Barrientos, D., Counterman, B. A. & Noor, M. A. F. The genetics of speciation by reinforcement. PLoS Biol. 2, 2256–2263 (2004)
Saether, S. A. et al. Sex chromosome-linked species recognition and evolution of reproductive isolation in flycatchers. Science 318, 95–97 (2007)
Levin, D. A. Reproductive character displacement in Phlox . Evolution 39, 1275–1281 (1985)
Levin, D. A. Hybridization between annual species of Phlox – population structure. Am. J. Bot. 54, 1122-&. (1967)
Levin, D. A. Interspecific hybridization, heterozygosity and gene exchange in Phlox . Evolution 29, 37–51 (1975)
Ruane, L. G. & Donohue, K. Pollen competition and environmental effects on hybridization dynamics between Phlox drummondii and Phlox cuspidata . Evol. Ecol. 22, 229–241 (2008)
Gonnet, J. F. CIELab measurement, a precise communication in flower color: an example with carnation (Dianthus Caryophyllus) cultivars. J. Hortic. Sci. 68, 499–510 (1993)
Holton, T. A. & Cornish, E. C. Genetics and biochemistry of anthocyanin biosynthesis. Plant Cell 7, 1071–1083 (1995)
Wittkopp, P. J., Haerum, B. K. & Clark, A. G. Evolutionary changes in cis and trans gene regulation. Nature 430, 85–88 (2004)
Koes, R., Verweij, W. & Quattrocchio, F. Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci. 10, 236–242 (2005)
Kirkpatrick, M. Reinforcement and divergence under assortative mating. Proc. R. Soc. Lond. B 267, 1649–1655 (2000)
Haldane, J. B. S. A mathematical theory of natural and artificial selection, part 1. Trans. Cambr. Phil. Soc. 23, 19–41 (1924)
Orr, H. A. & Betancourt, A. J. Haldane’s sieve and adaptation from the standing genetic variation. Genetics 157, 875–884 (2001)
Lowry, D. B., Modliszewski, J. L., Wright, K. M., Wu, C. A. & Willis, J. H. The strength and genetic basis of reproductive isolating barriers in flowering plants. Phil. Trans. R. Soc. B 363, 3009–3021 (2008)
Nosil, P., Vines, T. H. & Funk, D. J. Perspective: reproductive isolation caused by natural selection against immigrants from divergent habitats. Evolution 59, 705–719 (2005)
Dambroski, H. R. et al. The genetic basis for fruit odor discrimination in Rhagoletis flies and its significance for sympatric host shifts. Evolution 59, 1953–1964 (2005)
Lowry, D. B. & Willis, J. H. A widespread chromosomal inversion polymorphism contributes to a major life-history transition, local adaptation, and reproductive isolation. PLoS Biol. 8, e1000500 (2010)
Levin, D. A. The exploitation of pollinators by species and hybrids of Phlox . Evolution 24, 367–377 (1970)
Hoballah, M. E. et al. Single gene-mediated shift in pollinator attraction in Petunia . Plant Cell 19, 779–790 (2007)
Schwinn, K. et al. A small family of MYB-regulatory genes controls floral pigmentation intensity and patterning in the genus Antirrhinum . Plant Cell 18, 831–851 (2006)
Des Marais, D. L. & Rausher, M. D. Parallel evolution at multiple levels in the origin of hummingbird pollinated flowers in Ipomoea . Evolution 64, 2044–2054 (2010)
Kelly, A. J. & Willis, J. H. Polymorphic microsatellite loci in Mimulus guttatus and related species. Mol. Ecol. 7, 769–774 (1998)
Mouritzen, P. et al. The ProbeLibrary™-expression profiling 99% of all human genes using only 90 dual-labeled real-time PCR probes. Biotechniques 37, 492–495 (2004)
Rieu, I. & Powers, S. J. Real-time quantitative RT-PCR: design, calculations, and statistics. Plant Cell 21, 1031–1033 (2009)
Ahmadian, A. et al. Single-nucleotide polymorphism analysis by pyrosequencing. Anal. Biochem. 280, 103–110 (2000)
Acknowledgements
We thank D. Des Marais, J. Tung, S. Johnsen and T. Juenger for technical advice, D. Levin for assistance in locating natural populations, and M. Noor for comments on the manuscript. This work was supported by a National Science Foundation grant to M.D.R. and a National Science Foundation Doctoral Dissertation Research Improvement Grant to R.H. R.H. was supported in part by the National Science Foundation Graduate Research Fellowship Program.
Author information
Authors and Affiliations
Contributions
R.H. and M.D.R. designed the project; R.H. performed the experiments and the analyses; R.H. and M.D.R. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Tables 1-9 and Supplementary Figures 1-4 with legends. (PDF 311 kb)
Rights and permissions
About this article
Cite this article
Hopkins, R., Rausher, M. Identification of two genes causing reinforcement in the Texas wildflower Phlox drummondii. Nature 469, 411–414 (2011). https://doi.org/10.1038/nature09641
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09641
This article is cited by
-
Natural variation in Beauty Mark is associated with UV-based geographical adaptation in Gossypium species
BMC Biology (2023)
-
The Flower Colour Influences Spontaneous Nectaring in Butterflies: a Case Study with Twenty Subtropical Butterflies
Neotropical Entomology (2023)
-
The genomic and bulked segregant analysis of Curcuma alismatifolia revealed its diverse bract pigmentation
aBIOTECH (2022)
-
Transcriptome profiles of yellowish-white and fuchsia colored flowers in the Rheum palmatum complex reveal genes related to color polymorphism
Plant Molecular Biology (2022)
-
Neurogenomic divergence during speciation by reinforcement of mating behaviors in chorus frogs (Pseudacris)
BMC Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.