Abstract
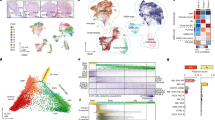
Medulloblastoma encompasses a collection of clinically and molecularly diverse tumour subtypes that together comprise the most common malignant childhood brain tumour1,2,3,4. These tumours are thought to arise within the cerebellum, with approximately 25% originating from granule neuron precursor cells (GNPCs) after aberrant activation of the Sonic Hedgehog pathway (hereafter, SHH subtype)3,4,5,6,7,8. The pathological processes that drive heterogeneity among the other medulloblastoma subtypes are not known, hindering the development of much needed new therapies. Here we provide evidence that a discrete subtype of medulloblastoma that contains activating mutations in the WNT pathway effector CTNNB1 (hereafter, WNT subtype)1,3,4 arises outside the cerebellum from cells of the dorsal brainstem. We found that genes marking human WNT-subtype medulloblastomas are more frequently expressed in the lower rhombic lip (LRL) and embryonic dorsal brainstem than in the upper rhombic lip (URL) and developing cerebellum. Magnetic resonance imaging (MRI) and intra-operative reports showed that human WNT-subtype tumours infiltrate the dorsal brainstem, whereas SHH-subtype tumours are located within the cerebellar hemispheres. Activating mutations in Ctnnb1 had little impact on progenitor cell populations in the cerebellum, but caused the abnormal accumulation of cells on the embryonic dorsal brainstem which included aberrantly proliferating Zic1+ precursor cells. These lesions persisted in all mutant adult mice; moreover, in 15% of cases in which Tp53 was concurrently deleted, they progressed to form medulloblastomas that recapitulated the anatomy and gene expression profiles of human WNT-subtype medulloblastoma. We provide the first evidence, to our knowledge, that subtypes of medulloblastoma have distinct cellular origins. Our data provide an explanation for the marked molecular and clinical differences between SHH- and WNT-subtype medulloblastomas and have profound implications for future research and treatment of this important childhood cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Ellison, D. W. et al. β-Catenin status predicts a favorable outcome in childhood medulloblastoma: the United Kingdom Children’s Cancer Study Group Brain Tumour Committee. J. Clin. Oncol. 23, 7951–7957 (2005)
Gajjar, A. et al. Risk-adapted craniospinal radiotherapy followed by high-dose chemotherapy and stem-cell rescue in children with newly diagnosed medulloblastoma (St Jude Medulloblastoma-96): long-term results from a prospective, multicentre trial. Lancet Oncol. 7, 813–820 (2006)
Thompson, M. C. et al. Genomics identifies medulloblastoma subgroups that are enriched for specific genetic alterations. J. Clin. Oncol. 24, 1924–1931 (2006)
Kool, M. et al. Integrated genomics identifies five medulloblastoma subtypes with distinct genetic profiles, pathway signatures and clinicopathological features. PLoS ONE 3, e3088 (2008)
Gilbertson, R. J. & Ellison, D. W. The origins of medulloblastoma subtypes. Annu. Rev. Pathol. 3, 341–365 (2008)
Goodrich, L. V., Milenkovic, L., Higgins, K. M. & Scott, M. P. Altered neural cell fates and medulloblastoma in mouse patched mutants. Science 277, 1109–1113 (1997)
Schuller, U. et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh-induced medulloblastoma. Cancer Cell 14, 123–134 (2008)
Yang, Z. J. et al. Medulloblastoma can be initiated by deletion of Patched in lineage-restricted progenitors or stem cells. Cancer Cell 14, 135–145 (2008)
Romer, J. T. et al. Suppression of the Shh pathway using a small molecule inhibitor eliminates medulloblastoma in Ptc1+/−p53−/− mice. Cancer Cell 6, 229–240 (2004)
Rudin, C. M. et al. Treatment of medulloblastoma with Hedgehog pathway inhibitor GDC-0449. N. Engl. J. Med. 2, 1173–1178 (2009)
Louis, D., Ohgaki, H., Wiestler, O., Cavenee, W. (eds) World Health Organization Classification of Tumours of the Central Nervous System (International Agency for Research on Cancer, 2007)
Huang, X. et al. Transventricular delivery of Sonic hedgehog is essential to cerebellar ventricular zone development. Proc Natl Acad. Sci. USA 107, 8422–8427 (2010)
Johnson, R. A. et al. Cross-species genomics matches driver mutations and cell compartments to model ependymoma. Nature 466, 632–636 (2010)
Lee, Y. et al. A molecular fingerprint for medulloblastoma. Cancer Res. 63, 5428–5437 (2003)
Ray, R. S. & Dymecki, S. M. Rautenlippe Redux—toward a unified view of the precerebellar rhombic lip. Curr. Opin. Cell Biol. 21, 741–747 (2009)
Morales, D. & Hatten, M. E. Molecular markers of neuronal progenitors in the embryonic cerebellar anlage. J. Neurosci. 26, 12226–12236 (2006)
Harada, N. et al. Intestinal polyposis in mice with a dominant stable mutation of the beta-catenin gene. EMBO J. 18, 5931–5942 (1999)
Hegedus, B. et al. Neurofibromatosis-1 regulates neuronal and glial cell differentiation from neuroglial progenitors in vivo by both cAMP- and Ras-dependent mechanisms. Cell Stem Cell 1, 443–457 (2007)
Storm, R. et al. The bHLH transcription factor Olig3 marks the dorsal neuroepithelium of the hindbrain and is essential for the development of brainstem nuclei. Development 136, 295–305 (2009)
Jonkers, J. et al. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nature Genet. 29, 418–425 (2001)
Wetmore, C., Eberhart, D. E. & Curran, T. Loss of p53 but not ARF accelerates medulloblastoma in mice heterozygous for patched. Cancer Res. 61, 513–516 (2001)
Chenn, A. & Walsh, C. A. Regulation of cerebral cortical size by control of cell cycle exit in neural precursors. Science 297, 365–369 (2002)
Landsberg, R. L. et al. Hindbrain rhombic lip is comprised of discrete progenitor cell populations allocated by Pax6. Neuron 48, 933–947 (2005)
DiPietrantonio, H. J. & Dymecki, S. M. Zic1 levels regulate mossy fiber neuron position and axon laterality choice in the ventral brain stem. Neuroscience 162, 560–573 (2009)
Farago, A. F., Awatramani, R. B. & Dymecki, S. M. Assembly of the brainstem cochlear nuclear complex is revealed by intersectional and subtractive genetic fate maps. Neuron 50, 205–218 (2006)
Oliver, T. G. et al. Loss of patched and disruption of granule cell development in a pre-neoplastic stage of medulloblastoma. Development 132, 2425–2439 (2005)
Uziel, T. et al. The tumor suppressors Ink4c and p53 collaborate independently with Patched to suppress medulloblastoma formation. Genes Dev. 19, 2656–2667 (2005)
Frappart, P. O. et al. Recurrent genomic alterations characterize medulloblastoma arising from DNA double-strand break repair deficiency. Proc. Natl Acad. Sci. USA 106, 1880–1885 (2009)
Ikeda, A., Ikeda, S., Gridley, T., Nishina, P. M. & Naggert, J. K. Neural tube defects and neuroepithelial cell death in Tulp3 knockout mice. Hum. Mol. Genet. 10, 1325–1334 (2001)
Acknowledgements
R.J.G. holds the Howard C. Schott Research Chair from the Malia’s Cord Foundation, and is supported by grants from the National Institutes of Health (R01CA129541, P01CA96832 and P30CA021765), the Collaborative Ependymoma Research Network and by the American Lebanese Syrian Associated Charities. We are grateful to A. Chenn, J. Johnson and C. Birchmeier for their gifts of reagents and the staff of the Hartwell Center for Bioinformatics and Biotechnology and ARC at St Jude Children’s Research Hospital for technical assistance.
Author information
Authors and Affiliations
Contributions
R.J.G. conceived the research and planned experiments. P.G. also planned and conducted most of the experiments. Y.T., G.R., D.S.C., M.C.T., T.H., H.P., J.M., J.C.L., Y.L., F.Z., C.E., S.C.C., M.F.R., P.J.M. and R.W.-R. conducted experiments. D.F. and S.P. provided bioinformatic expertise. A.G., F.A.B. and R.A.S. provided clinical advice and tumour samples. D.H.G. provided the Blbp-Cre mouse and data. M.M.T. provided the Ctnnb1lox(ex3)/lox(ex3) mouse. Z.P. and R.O. reviewed and analysed the human MRI scans. D.W.E. provided pathology review. All authors contributed to writing the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Methods, additional references, Supplementary Figures 1-14 with legends and Supplementary Table 1. (PDF 12061 kb)
Supplementary Dataset 1
This dataset reports the expression distribution in the developing mouse hindbrain (rhombomeres 1-8) of 24 and 25 signature genes of human WNT-subgroup and SHH-subgroup medulloblastoma respectfully. (PDF 30323 kb)
Rights and permissions
About this article
Cite this article
Gibson, P., Tong, Y., Robinson, G. et al. Subtypes of medulloblastoma have distinct developmental origins. Nature 468, 1095–1099 (2010). https://doi.org/10.1038/nature09587
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09587
This article is cited by
-
Targeting the key players of phenotypic plasticity in cancer cells by phytochemicals
Cancer and Metastasis Reviews (2024)
-
Identifying new biomarkers of aggressive Group 3 and SHH medulloblastoma using 3D hydrogel models, single cell RNA sequencing and 3D OrbiSIMS imaging
Acta Neuropathologica Communications (2023)
-
A sellar presentation of a WNT-activated embryonal tumor: further evidence of an ectopic medulloblastoma
Acta Neuropathologica Communications (2023)
-
Neuron–oligodendroglial interactions in health and malignant disease
Nature Reviews Neuroscience (2023)
-
Molecular subgroup of medulloblastoma: evaluation of contribution to CSF diversion following tumour resection
Child's Nervous System (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.