Abstract
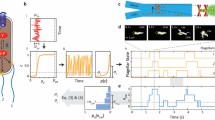
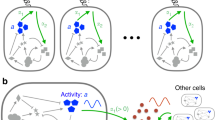
The chemotaxis signalling network in Escherichia coli that controls the locomotion of bacteria is a classic model system for signal transduction1,2. This pathway modulates the behaviour of flagellar motors to propel bacteria towards sources of chemical attractants. Although this system relaxes to a steady state in response to environmental changes, the signalling events within the chemotaxis network are noisy and cause large temporal variations of the motor behaviour even in the absence of stimulus3. That the same signalling network governs both behavioural variability and cellular response raises the question of whether these two traits are independent. Here, we experimentally establish a fluctuation–response relationship in the chemotaxis system of living bacteria. Using this relationship, we demonstrate the possibility of inferring the cellular response from the behavioural variability measured before stimulus. In monitoring the pre- and post-stimulus switching behaviour of individual bacterial motors, we found that variability scales linearly with the response time for different functioning states of the cell. This study highlights that the fundamental relationship between fluctuation and response is not constrained to physical systems at thermodynamic equilibrium4 but is extensible to living cells5. Such a relationship not only implies that behavioural variability and cellular response can be coupled traits, but it also provides a general framework within which we can examine how the selection of a network design shapes this interdependence.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
09 December 2010
The position of a sentence was changed in the first paragraph of the text.
References
Bourret, R. B., Borkovich, K. A. & Simon, M. I. Signal transduction pathways involving protein phosphorylation in prokaryotes. Annu. Rev. Biochem. 60, 401–441 (1991)
Bourret, R. B. & Stock, A. M. Molecular information processing: lessons from bacterial chemotaxis. J. Biol. Chem. 277, 9625–9628 (2002)
Korobkova, E. A., Emonet, T., Park, H. & Cluzel, P. Hidden stochastic nature of a single bacterial motor. Phys. Rev. Lett. 96, 058105 (2006)
Callen, H. B. & Welton, T. A. Irreversibility and generalized noise. Phys. Rev. 83, 34–40 (1951)
Prost, J., Joanny, J. F. & Parrondo, J. M. Generalized fluctuation-dissipation theorem for steady-state systems. Phys. Rev. Lett. 103, 090601 (2009)
Bustamante, C., Macosko, J. C. & Wuite, G. J. L. Grabbing the cat by the tail: manipulating molecules one by one. Nature Rev. Mol. Cell Biol. 1, 130–136 (2000)
Dorignac, J., Kalinowski, A., Erramilli, S. & Mohanty, P. Dynamical response of nanomechanical oscillators in immiscible viscous fluid for in vitro biomolecular recognition. Phys. Rev. Lett. 96, 186105 (2006)
Paulsson, J. Summing up the noise in gene networks. Nature 427, 415–418 (2004)
Ozbudak, E. M., Thattai, M., Kurtser, I., Grossman, A. D. & van Oudenaarden, A. Regulation of noise in the expression of a single gene. Nature Genet. 31, 69–73 (2002)
Cugliandolo, L. F., Dean, D. S. & Kurchan, J. Fluctuation-dissipation theorems and entropy production in relaxational systems. Phys. Rev. Lett. 79, 2168–2171 (1997)
Chetrite, R., Falkovich, G. & Gawedzki, K. Fluctuation relations in simple examples of non-equilibrium steady states. J. Stat. Mech.-Theory E 2008, P08005 (2008)
Speck, T. & Seifert, U. Restoring a fluctuation-dissipation theorem in a nonequilibrium steady state. Europhys. Lett. 74, 391–396 (2006)
Berg, H. C. Motile behavior of bacteria. Phys. Today 53, 24–29 (2000)
Korobkova, E., Emonet, T., Vilar, J. M., Shimizu, T. S. & Cluzel, P. From molecular noise to behavioural variability in a single bacterium. Nature 428, 574–578 (2004)
Cluzel, P., Surette, M. & Leibler, S. An ultrasensitive bacterial motor revealed by monitoring signaling proteins in single cells. Science 287, 1652–1655 (2000)
Sourjik, V. & Berg, H. C. Receptor sensitivity in bacterial chemotaxis. Proc. Natl Acad. Sci. USA 99, 123–127 (2002)
Barkai, N. & Leibler, S. Robustness in simple biochemical networks. Nature 387, 913–917 (1997)
Alon, U., Surette, M. G., Barkai, N. & Leibler, S. Robustness in bacterial chemotaxis. Nature 397, 168–171 (1999)
Anderson, R. L. Distribution of the serial correlation coefficient. Ann. Math. Stat. 13, 1–13 (1942)
Ratnam, R. & Nelson, M. E. Nonrenewal statistics of electrosensory afferent spike trains: implications for the detection of weak sensory signals. J. Neurosci. 20, 6672–6683 (2000)
Emonet, T. & Cluzel, P. Relationship between cellular response and behavioral variability in bacterial chemotaxis. Proc. Natl Acad. Sci. USA 105, 3304–3309 (2008)
Shibata, T. & Fujimoto, K. Noisy signal amplification in ultrasensitive signal transduction. Proc. Natl Acad. Sci. USA 102, 331–336 (2005)
Bray, D., Levin, M. D. & Morton-Firth, C. J. Receptor clustering as a cellular mechanism to control sensitivity. Nature 393, 85–88 (1998)
Sourjik, V. & Berg, H. C. Functional interactions between receptors in bacterial chemotaxis. Nature 428, 437–441 (2004)
Rao, C. V., Wolf, D. M. & Arkin, A. P. Control, exploitation and tolerance of intracellular noise. Nature 420, 231–237 (2002)
Bialek, W. & Setayeshgar, S. Physical limits to biochemical signaling. Proc. Natl Acad. Sci. USA 102, 10040–10045 (2005)
Andrews, B. W., Yi, T. M. & Iglesias, P. A. Optimal noise filtering in the chemotactic response of Escherichia coli . PLOS Comput. Biol. 2, e154 (2006)
Goldbeter, A. & Koshland, D. E. Jr. An amplified sensitivity arising from covalent modification in biological systems. Proc. Natl Acad. Sci. USA 78, 6840–6844 (1981)
Detwiler, P. B., Ramanathan, S., Sengupta, A. & Shraiman, B. I. Engineering aspects of enzymatic signal transduction: photoreceptors in the retina. Biophys. J. 79, 2801–2817 (2000)
Shinar, G. & Feinberg, M. Structural sources of robustness in biochemical reaction networks. Science 327, 1389–1391 (2010)
Parkinson, J. S. & Houts, S. E. Isolation and behavior of Escherichia coli deletion mutants lacking chemotaxis functions. J. Bacteriol. 151, 106–113 (1982)
Lutz, R. & Bujard, H. Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements. Nucleic Acids Res. 25, 1203–1210 (1997)
Jasuja, R., Yu-Lin, Trentham, D. R. & Khan, S. Response tuning in bacterial chemotaxis. Proc. Natl Acad. Sci. USA 96, 11346–11351 (1999)
Adler, J. A method for measuring chemotaxis and use of the method to determine optimum conditions for chemotaxis by Escherichia coli . J. Gen. Microbiol. 74, 77–91 (1973)
Bainer, R., Park, H. & Cluzel, P. A high-throughput capillary assay for bacterial chemotaxis. J. Microbiol. Methods 55, 315–319 (2003)
Acknowledgements
This research was funded by an NSF DMR award 0213745 to the Materials Research Science and Engineering Center at the University of Chicago, and NIH award R01AI059195-03 (to P.C.). W.P. and T.E. were supported by NSF CCF0829836, an Alfred P. Sloan Research Fellowship, and a National Academies Keck Futures Initiative award (to T.E.). J.F.M. was supported by NSF awards PHY-0852130 and DMR-0715099 and NIH grant 1U54CA143869-01. This work was also supported by the Chicago Biomedical Consortium with support from The Searle Funds at The Chicago Community Trust. D. Trentham supplied caged l-aspartate. We thank J. S. Parkinson for ΔCheB mutant strains RP4972 and RP4992. We thank T. Shimizu for discussions and sharing unpublished work. We thank H. Lee for help with the HPLC measurements. We thank J. Moffitt and K. Wood for comments on the manuscript and all members of the Cluzel laboratory for many discussions. W. Grus provided editorial assistance.
Author information
Authors and Affiliations
Contributions
P.C. conceived and designed the research. H.P. performed all the experiments. H.P., P.C., T.E., W.P. and J.F.M. analysed the data. H.P., P.C., J.F.M. and T.E. wrote the paper. C.C.G. constructed the pZE21-CheR plasmid.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
The file contains Supplementary Figures 1-9 with legends and additional references. (PDF 1313 kb)
Rights and permissions
About this article
Cite this article
Park, H., Pontius, W., Guet, C. et al. Interdependence of behavioural variability and response to small stimuli in bacteria. Nature 468, 819–823 (2010). https://doi.org/10.1038/nature09551
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09551
This article is cited by
-
Flagellar dynamics reveal fluctuations and kinetic limit in the Escherichia coli chemotaxis network
Scientific Reports (2023)
-
Chemotactic behaviour of Escherichia coli at high cell density
Nature Communications (2019)
-
Bacterial chemotaxis in a microfluidic T-maze reveals strong phenotypic heterogeneity in chemotactic sensitivity
Nature Communications (2019)
-
Stochastic effects as a force to increase the complexity of signaling networks
Scientific Reports (2013)
-
Noise characteristics of the Escherichia coli rotary motor
BMC Systems Biology (2011)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.