Abstract
A key feature of RNA interference is its ability to spread from cell to cell. Such non-cell-autonomous gene silencing has been characterized extensively in both plants and animals, but the identity of the mobile silencing signal has remained elusive. Several recent studies now shed light on the identity of this signal in plants, and indicate that small RNA molecules—from short-interfering RNAs to microRNAs—are capable of moving between cells and through the vasculature. The movement of small, 21–24-nucleotide RNA species has implications for biological processes ranging from developmental patterning and stress responses to epigenetic inheritance.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Voinnet, O. & Baulcombe, D. C. Systemic signaling in gene silencing. Nature 389, 553 (1997)
Palauqui, J. C., Elmayan, T., Pollien, J. M. & Vaucheret, H. Systemic acquired silencing: transgene-specific post-transcriptional silencing is transmitted by grafting from silenced stocks to non-silenced scions. EMBO J. 16, 4738–4745 (1997)
Fire, A. et al. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans . Nature 391, 806–811 (1998)
Timmons, L. & Fire, A. Specific interference by ingested dsRNA. Nature 395, 854 (1998)
Schwach, F., Vaistij, F. E., Jones, L. & Baulcombe, D. C. An RNA-dependent RNA polymerase prevents meristem invasion by potato virus X and is required for the activity but not the production of a systemic silencing signal. Plant Physiol. 138, 1842–1852 (2005)
Pant, B. D., Buhtz, A., Kehr, J. & Scheible, W. R. MicroRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J. 53, 731–738 (2008)
Buhtz, A., Springer, F., Chappell, L., Baulcombe, D. C. & Kehr, J. Identification and characterization of small RNAs from the phloem of Brassica napus . Plant J. 53, 739–749 (2008)
Chitwood, D. H. et al. Pattern formation via small RNA mobility. Genes Dev. 23, 549–554 (2009)This work documents the movement of endogenous ta-siRNAs by observing their accumulation outside the region of their biogenesis and proposes that gradients of small RNAs pattern gene expression through dose-dependent activity.
Carlsbecker, A. et al. Cell signaling by microRNA165/6 directs gene dose-dependent root cell fate. Nature 465, 316–321 (2010)By localizing the accumulation and activity of miR165/6 and studying the expression of its precursors and targets, this work proposes that radial patterning in the root results from cell to cell movement of an miRNA and a transcription factor.
Himber, C., Dunoyer, P., Moissiard, G., Ritzenthaler, C. & Voinnet, O. Transitivity-dependent and -independent cell-to-cell movement of RNA silencing. EMBO J. 22, 4523–4533 (2003)
Dunoyer, P., Himber, C. & Voinnet, O. DICER-LIKE4 is required for RNA interference and produces the 21-nucleotide small interfering RNA component of the plant cell-to-cell silencing signal. Nature Genet. 37, 1356–1360 (2005)
Dunoyer, D., Himber, C., Ruiz-Ferrer, V., Alioua, A. & Voinnet, O. Intra- and intercellular RNA interference in Arabidopsis thaliana requires components of the microRNA and heterochromatic silencing pathways. Nature Genet. 39, 848–856 (2007)
Smith, L. M. et al. An SNF2 protein associated with nuclear RNA silencing and the spread of a silencing signal between cells in Arabidopsis . Plant Cell 19, 1507–1521 (2007)
Dalmay, T., Hamilton, A., Rudd, S., Angell, S. & Baulcombe, D. C. An RNA-dependent RNA polymerase gene in Arabidopsis is required for posttranscriptional gene silencing mediated by a transgene but not by a virus. Cell 101, 543–553 (2000)
Mourrain, P. et al. Arabidopsis SGS2 and SGS3 genes are required for posttranscriptional gene silencing and natural virus resistance. Cell 101, 533–542 (2000)
Kobayashi, K. & Zambryski, P. RNA silencing and its cell-to-cell spread during Arabidopsis embryogenesis. Plant J. 50, 597–604 (2007)
Dunoyer, P. et al. Small RNA duplexes function as mobile silencing signals between plant cells. Science 328, 912–916 (2010)The authors use transgenic chimaeras to show that small RNAs form the mobile silencing signal, and on the basis of bombardment assays they conclude that the signal consists specifically of siRNA duplexes.
Vargason, J. M., Szittya, G., Burgyan, J. & Hall, T. M. Size selective recognition of siRNA by an RNA silencing suppressor. Cell 115, 799–811 (2003)
Montgomery, T. A. et al. Specificity of ARGONAUTE7-miR390 interaction and dual functionality in TAS3 trans-acting siRNA formation. Cell 133, 128–141 (2008)
Levine, E., McHale, P. & Levine, H. Small regulatory RNAs may sharpen spatial expression patterns. PLoS Comput. Biol. 3, e233 (2007)
Mi, S. et al. Sorting of small RNAs in Arabidopsis argonaute complexes is directed by the 5′ terminal nucleotide. Cell 133, 116–127 (2008)
Brodersen, P. et al. Widespread translational inhibition by plant miRNAs and siRNAs. Science 320, 1185–1190 (2008)
Chapman, E. J. & Carrington, J. C. Specialization and evolution of endogenous small RNA pathways. Nature Rev. Genet. 8, 884–896 (2007)
Deleris, A. et al. Hierarchical action and inhibition of plant Dicer-like proteins in antiviral defense. Science 313, 68–71 (2006)
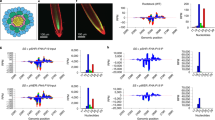
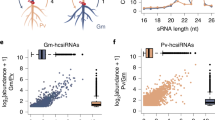
Molnar, A. et al. Small silencing RNAs in plants are mobile and direct epigenetic modification in recipient cells. Science 328, 872–875 (2010)Through the use of grafting and next-generation sequencing, this paper catalogues the movement of endogenous siRNAs, demonstrating that they can initiate methylation of target loci in recipient graft tissue.
Baurle, I., Smith, L., Baulcombe, D. C. & Dean, C. Widespread role for the flowering-time regulators FCA and FPA in RNA-mediated chromatin silencing. Science 318, 109–112 (2007)
Borsani, O., Zhu, J., Verslues, P. E., Sunkar, R. & Zhu, J. K. Endogenous siRNAs derived from a pair of natural cis-antisense transcripts regulate salt tolerance in Arabidopsis . Cell 123, 1279–1291 (2005)
Slotkin, R. K. et al. Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136, 461–472 (2009)
Malone, C. D. & Hannon, G. J. Small RNAs as guardians of the genome. Cell 136, 656–668 (2009)
Mosher, R. A. et al. Uniparental expression of PolIV-dependent siRNAs in developing endosperm of Arabidopsis . Nature 460, 283–286 (2009)
Dunoyer, P. et al. An endogenous, systemic RNAi pathway in plants. EMBO J. 29, 1699–1712 (2010)
Yoo, B. C. et al. A systemic small RNA signaling system in plants. Plant Cell 16, 1979–2000 (2004)
Alvarez, J. P. et al. Endogenous and synthetic microRNAs stimulate simultaneous, efficient, and localized regulation of multiple targets in diverse species. Plant Cell 18, 1134–1151 (2006)
Parizotto, E. A., Dunoyer, P., Rahm, N., Himber, C. & Voinnet, O. In vivo investigation of the transcription, processing, endonucleolytic activity, and functional relevance of the spatial distribution of a plant miRNA. Genes Dev. 18, 2237–2242 (2004)
Schwab, R., Ossowski, S., Riester, M., Warthmann, N. & Weigel, D. Highly specific gene silencing by artificial microRNAs in Arabidopsis . Plant Cell 18, 1121–1133 (2006)
Tretter, E. M., Alvarez, J. P., Eshed, Y. & Bowman, J. L. Activity range of Arabidopsis small RNAs derived from different biogenesis pathways. Plant Physiol. 147, 58–62 (2008)
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nature Cell Biol. 9, 654–659 (2007)
Ruvkun, G. The perfect storm of tiny RNAs. Nature Med. 14, 1041–1045 (2008)
Havecker, E. R. et al. The Arabidopsis RNA-directed DNA methylation ARGONAUTES functionally diverge based on their expression and interaction with target loci. Plant Cell 22, 321–334 (2010)
Tucker, M. R. et al. Vascular signaling mediated by ZWILLE potentiates WUSCHEL function during shoot meristem stem cell development in the Arabidopsis embryo. Development 135, 2839–2843 (2008)
Park, M. Y., Wu, G., Gonzalez-Sulser, A., Vaucheret, H. & Poethig, R. S. Nuclear processing and export of microRNAs in Arabidopsis . Proc. Natl Acad. Sci. USA 102, 3691–3696 (2005)
Liu, Q. et al. The ARGONAUTE10 gene modulates shoot apical meristem maintenance and leaf polarity establishment by repressing miR165/6 in Arabidopsis . Plant J. 58, 27–40 (2008)
Acknowledgements
We thank current and former members of the Timmermans laboratory for discussions. Research on small RNA mobility in the laboratory of M.C.P.T. is supported by a grant from the NSF (IOS-0615752).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Chitwood, D., Timmermans, M. Small RNAs are on the move. Nature 467, 415–419 (2010). https://doi.org/10.1038/nature09351
Issue Date:
DOI: https://doi.org/10.1038/nature09351
This article is cited by
-
Plasmodesmata mediate cell-to-cell transport of brassinosteroid hormones
Nature Chemical Biology (2023)
-
Implications of small RNAs in plant development, abiotic stress response and crop improvement in changing climate
The Nucleus (2023)
-
Small RNAs shoot for the root
Nature Plants (2021)
-
Roles of small RNAs in crop disease resistance
Stress Biology (2021)
-
Regulatory non-coding RNAs: a new frontier in regulation of plant biology
Functional & Integrative Genomics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.