Abstract
Arising from: Z. Shcheprova, S. Baldi, S. B. Frei, G. Gonnet & Y. Barral Nature 454, 728–734 (2008)10.1038/nature07212
During mitosis in Saccharomyces cerevisiae, senescence factors such as extrachromosomal ribosomal DNA circles (ERCs) are retained in the mother cell and excluded from the bud/daughter cell1. Shcheprova et al.2 proposed a model suggesting segregation of ERCs through their association with nuclear pore complexes (NPCs) and retention of pre-existing NPCs in the mother cell during mitosis. However, this model is inconsistent with previous data3 and we demonstrate here that NPCs do efficiently migrate from the mother into the bud. Therefore, binding to NPCs does not seem to explain the retention of ERCs in the mother cell.
Similar content being viewed by others
Main
On the basis of photobleaching experiments, Shcheprova et al.2 concluded that NPCs are prevented from migrating through the bud neck in anaphase and that new NPCs are inserted into the bud (but not the mother) nuclear envelope. Two testable predictions arise from this model: first, the number of NPCs in the mother nuclear envelope should remain constant throughout anaphase; and second, the total number of NPCs in the nuclear envelope should increase abruptly during anaphase.
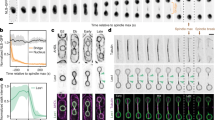
Using fluorescence time-lapse microscopy, we found that the intensity of GFP-tagged core NPC components at the nuclear envelope increased continuously during G1 phase, S phase and early mitosis but remained constant during nuclear migration through the bud neck in anaphase. Upon movement of the nucleus into the bud, the appearance of the GFP signal in the bud was associated with a pronounced GFP intensity drop in the mother (Fig. 1a, b). These results contradict the predictions of the model of Shcheprova et al.2.
a, Nuclear division followed with GFP-tagged Nup49. GFP intensity levels in sum projections (top panel) are colour-coded (bottom panel). Scale bar, 5 μm. b, Fluorescence intensity of GFP-tagged NPC components at the nuclear envelope in the mother, in the bud and their sum as a function of cell cycle progression (5 min time point corresponds to the onset of nuclear movement across the bud neck). Mean ± s.d. values (n = 5) are normalized to total intensity at time zero. c, Movement of NPCs analysed using photoconversion of Nic96 tagged with 2xDendra2. Nic96–2xDendra2 was photoconverted (green to red) at the nuclear envelope facing the bud neck before nuclear migration. Movement of red-labelled NPCs was followed by single-plane time-lapse microscopy. Scale bar, 5 μm.
A serial thin-sectioning electron microscopy study3, not cited by Shcheprova et al.2, showed that the number of NPCs in the nuclear envelope increases continuously from G1 (86 ± 16) to S phase (117 ± 17) to early mitosis (142 ± 17) and remains constant during anaphase (137 ± 29) (Table 1 in ref. 3). These results are incompatible with the model of Shcheprova et al.2 but consistent with our data (Fig. 1a, b).
Shcheprova et al.2 investigated NPC movement between mother and bud using fluorescence loss in photobleaching (FLIP) experiments with GFP-tagged NPC components. Repeated photobleaching of GFP in the mother nuclear envelope induced a decrease of the nuclear envelope fluorescence in the mother whereas the signal in the bud remained constant (Fig. 1a, b in ref. 2). Shcheprova et al.2 assumed that the signal in the bud was not affected by photobleaching and concluded that NPCs do not move from the mother to the bud during anaphase. However, Fig. 1a, b shows that the signal in the bud strongly increases during anaphase. If the GFP signal of bud NPCs was independent of the pre-existing pores in the mother cell, the FLIP experiments should have shown an increase in the bud signal; instead Shcheprova et al.2 reported a constant signal. This inconsistency, which would have become apparent if the FLIP experiments had been conducted with respect to unbleached control cells, suggests that substantial numbers of NPCs migrate from the mother into the bud during anaphase.
To investigate this, we used strains expressing NPC components fused to the photoconvertible fluorescent protein Dendra24. Upon photoconversion in the mother, movement of marked (red-fluorescent) NPCs to the bud could be observed (Fig. 1c; see also http://www.embl.de/SegregationMovie for a movie of this photoconversion experiment; movie also available from the authors). Analogous results were also obtained using iFRAP (see Appendix).
Photoconversion of all NPCs at the nuclear envelope in metaphase (using Nup49 or Nic96 tagged with Dendra2) and quantification of the red fluorescence in the two resulting nuclei after cell division indicated that 38 ± 12% (mean ± s.d., n = 50) of pre-existing NPCs moved into the bud during mitosis. The surface density of marked NPCs is roughly equal in both compartments, as the surface area of the bud nucleus is only ∼70% of the mother nucleus at the end of telophase (as determined by quantification of the NLS–GFP intensity in mother and bud nuclei in a separate experiment (n = 93), consistent with a previous report5). We conclude that NPCs effectively migrate into the bud along with the nucleus.
Our results and those of ref. 3 demonstrate that pre-existing NPCs are not retained in the mother but are partitioned between mother and bud nuclei proportionally to nuclear surface area. This questions the model for asymmetric segregation of ERCs during yeast mitosis based on the general retention of pre-existing NPCs in the mother, as proposed by Shcheprova et al.2 and highlighted elsewhere6,7,8,9,10. Therefore, how asymmetric segregation of ERCs is achieved remains an open question.
APPENDIX
An inverse fluorescence recovery after photobleaching (iFRAP) experiment demonstrates the appearance of NPC signal in the bud in the absence of a cytoplasmic labelled protein reservoir (Appendix Fig. 1a; quantification in Appendix Fig. 1b).
a, The cytoplasmic regions of cells expressing Nic96–GFP were photobleached using a 488-nm laser, as indicated by the red bleaching masks. As control, one entire cell was also bleached (marked with an asterisk). b, Quantification of the GFP signal intensity in mother and bud after photobleaching of the cytoplasmic GFP signal (mean ± s.d., n = 7).
Methods
NPC components were chromosomally tagged at the carboxy terminus using homologous recombination.
Imaging was performed on a Deltavision RT widefield microscope (Fig. 1) or a Leica SP-2 confocal microscope (see Appendix). For photoconversion, a 405-nm laser was used with a diffraction-limited spot, either on a Deltavision RT (Fig. 1c) or a Perkin Elmer Ultraview VoX spinning disc microscope (the photoconversion experiment quoted in the text). Flat-field, photobleaching and autofluorescence corrections were applied to all images.
References
Sinclair, D. A. & Guarente, L. Extrachromosomal rDNA circles—A cause of aging in yeast. Cell 91, 1033–1042 (1997)
Shcheprova, Z., Baldi, S., Frei, S. B., Gonnet, G. & Barral, Y. A mechanism for asymmetric segregation of age during yeast budding. Nature 454, 728–734 (2008)
Winey, M., Yarar, D., Giddings, T. H. & Mastronarde, D. N. Nuclear pore complex number and distribution throughout the Saccharomyces cerevisiae cell cycle by three-dimensional reconstruction from electron micrographs of nuclear envelopes. Mol. Biol. Cell 8, 2119–2132 (1997)
Chudakov, D. M., Lukyanov, S. & Lukyanov, K. A. Using photoactivatable fluorescent protein Dendra2 to track protein movement. Biotechniques 42, 553–563 (2007)
Heun, P., Laroche, T., Shimada, K., Furrer, P. & Gasser, S. M. Chromosome dynamics in the yeast interphase nucleus. Science 294, 2181–2186 (2001)
Barral, Y. & Knoblich, J. Cell division, growth and death. Curr. Opin. Cell Biol. 20, 647–649 (2008)
Caudron, F. & Barral, Y. Septins and the lateral compartmentalization of eukaryotic membranes. Dev. Cell 16, 493–506 (2009)
Henderson, K. A. & Gottschling, D. E. A mother’s sacrifice: what is she keeping for herself? Curr. Opin. Cell Biol. 20, 723–728 (2008)
Macara, I. G. & Mili, S. Polarity and differential inheritance—universal attributes of life? Cell 135, 801–812 (2008)
Güttinger, S., Laurell, E. & Kutay, U. Orchestrating nuclear envelope disassembly and reassembly during mitosis. Nature Rev. Mol. Cell Biol. 10, 178–191 (2009)
Author information
Authors and Affiliations
Ethics declarations
Competing interests
Competing financial interests: declared none.
PowerPoint slides
Rights and permissions
About this article
Cite this article
Khmelinskii, A., Keller, P., Lorenz, H. et al. Segregation of yeast nuclear pores. Nature 466, E1 (2010). https://doi.org/10.1038/nature09255
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature09255
This article is cited by
-
Analytical model for macromolecular partitioning during yeast cell division
BMC Biophysics (2014)
-
Organelle asymmetry for proper fitness, function, and fate
Chromosome Research (2013)
-
Tandem fluorescent protein timers for in vivo analysis of protein dynamics
Nature Biotechnology (2012)
-
Aging and TOR: interwoven in the fabric of life
Cellular and Molecular Life Sciences (2011)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.