Abstract
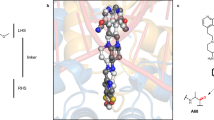
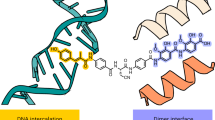
Despite the success of genomics in identifying new essential bacterial genes, there is a lack of sustainable leads in antibacterial drug discovery to address increasing multidrug resistance. Type IIA topoisomerases cleave and religate DNA to regulate DNA topology and are a major class of antibacterial and anticancer drug targets, yet there is no well developed structural basis for understanding drug action. Here we report the 2.1 Å crystal structure of a potent, new class, broad-spectrum antibacterial agent in complex with Staphylococcus aureus DNA gyrase and DNA, showing a new mode of inhibition that circumvents fluoroquinolone resistance in this clinically important drug target. The inhibitor ‘bridges’ the DNA and a transient non-catalytic pocket on the two-fold axis at the GyrA dimer interface, and is close to the active sites and fluoroquinolone binding sites. In the inhibitor complex the active site seems poised to cleave the DNA, with a single metal ion observed between the TOPRIM (topoisomerase/primase) domain and the scissile phosphate. This work provides new insights into the mechanism of topoisomerase action and a platform for structure-based drug design of a new class of antibacterial agents against a clinically proven, but conformationally flexible, enzyme class.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
19 August 2010
Author Andrew J. Theobald's middle initial was corrected for the print issue.
References
Mitscher, L. A. Bacterial topoisomerase inhibitors: quinolone and pyridone antibacterial agents. Chem. Rev. 105, 559–592 (2005)
Talbot, G. H. et al. Bad bugs need drugs: an update on the development pipeline from the Antimicrobial Availability Task Force of the Infectious Diseases Society of America. Clin. Infect. Dis. 42, 657–668 (2006)
Boucher, H. W. et al. Bad bugs, no drugs: no ESKAPE! An update from the Infectious Diseases Society of America. Clin. Infect. Dis. 48, 1–12 (2009)
Payne, D. J., Gwynn, M. N., Holmes, D. J. & Pompliano, D. L. Drugs for bad bugs: confronting the challenges of antibacterial discovery. Nature Rev. Drug Discov. 6, 29–40 (2007)
Schoeffler, A. J. & Berger, J. M. DNA topoisomerases: harnessing and constraining energy to govern chromosome topology. Q. Rev. Biophys. 41, 41–101 (2008)
Nöllmann, M. et al. Multiple modes of Escherichia coli DNA gyrase activity revealed by force and torque. Nature Struct. Mol. Biol. 14, 264–271 (2007)
Horowitz, D. S. & Wang, J. C. Mapping the active site tyrosine of Escherichia coli DNA gyrase. J. Biol. Chem. 262, 5339–5344 (1987)
Laponogov, I. et al. Structural insight into the quinolone–DNA cleavage complex of type IIA topoisomerases. Nature Struct. Biol. 16, 667–669 (2009)
Wigley, D. B., Davies, G. J., Dodson, E. J., Maxwell, A. & Dodson, G. Crystal structure of an N-terminal fragment of the DNA gyrase B protein. Nature 351, 624–629 (1991)
Berger, J. M., Gamblin, S. J., Harrison, S. C. & Wang, J. C. Structure and mechanism of DNA topoisomerase II. Nature 379, 225–232 (1996)
Morais Cabral, J. H. et al. Crystal structure of the breakage-reunion domain of DNA gyrase. Nature 388, 903–906 (1997)
Dong, K. C. & Berger, J. M. Structural basis for gate-DNA recognition and bending by type IIA topoisomerases. Nature 450, 1201–1205 (2007)
Gellert, M., Fisher, L. M. & O’Dea, M. H. DNA gyrase: purification and catalytic properties of a fragment of gyrase B protein. Proc. Natl Acad. Sci. USA 76, 6289–6293 (1979)
Reece, R. J. & Maxwell, A. Probing the limits of the DNA breakage-reunion domain of the Escherichia coli DNA gyrase A protein. J. Biol. Chem. 266, 3540–3546 (1991)
Liu, Q. & Wang, J. C. Identification of active site residues in the “GyrA” half of yeast DNA topoisomerase II. J. Biol. Chem. 273, 20252–20260 (1998)
Liu, Q. & Wang, J. C. Similarity in the catalysis of DNA breakage and rejoining by type IA and IIA DNA topoisomerases. Proc. Natl Acad. Sci. USA 96, 881–886 (1999)
Coates, W. J. et al. Preparation of Piperidinylalkylquinolines as Antibacterials. European patent 1051413 (1999)
Gomez, L. et al. Novel pyrazole derivatives as potent inhibitors of type II topoisomerases. Part 1: synthesis and preliminary SAR analysis. Bioorg. Med. Chem. Lett. 17, 2723–2727 (2007)
Wiener, J. J. M. et al. Tetrahydroindazole inhibitors of bacterial type II topoisomerases. Part 2: SAR development and potency against multidrug-resistant strains. Bioorg. Med. Chem. Lett. 17, 2718–2722 (2007)
Black, M. T. et al. Mechanism of action of the antibiotic NXL101, a novel nonfluoroquinolone inhibitor of bacterial type II topoisomerases. Antimicrob. Agents Chemother. 52, 3339–3349 (2008)
Aravind, L., Leipe, D. D. & Koonin, E. V. Toprim—a conserved catalytic domain in type IA and II topoisomerases, DnaG-type primases, OLD family nucleases and RecR proteins. Nucleic Acids Res. 26, 4205–4213 (1998)
Pan, X.-S., Gould, K. A. & Fisher, L. M. Probing the differential interactions of quinazolinedione PD 0305970 and quinolones with gyrase and topoisomerase IV. Antimicrob. Agents Chemother. 53, 3822–3831 (2009)
Gmunder, H., Kuratli, K. & Keck, W. In the presence of subunit A inhibitors DNA gyrase cleaves DNA fragments as short as 20 bp at specific sites. Nucleic Acids Res. 25, 604–611 (1997)
Fu, G. et al. Crystal structure of DNA gyrase B′ domain sheds lights on the mechanism for T-segment navigation. Nucleic Acids Res. 37, 5908–5916 (2009)
Edwards, M. J. et al. A crystal structure of the bifunctional antibiotic simocyclinone D8, bound to DNA gyrase. Science 326, 1415–1418 (2009)
Tretter, E. M., Schoeffler, A. J., Weisfield, S. R. & Berger, J. M. Crystal structure of the DNA gyrase GyrA N-terminal domain from Mycobacterium tuberculosis. Proteins 78, 492–495 (2010)
Jeffrey, G. A. & Saenger, W. Hydrogen Bonding in Biological Structures. (Springer, 1991)
Tanaka, M., Wang, T., Onodera, Y., Uchida, Y. & Sato, K. Mechanism of quinolone resistance in Staphylococcus aureus. J. Infect. Chemother. 6, 131–139 (2000)
Bock, C. W., Katz, A. K., Markham, G. D. & Glusker, J. P. Manganese as a replacement for magnesium and zinc: functional comparison of the divalent ions. J. Am. Chem. Soc. 121, 7360–7372 (1999)
Deweese, J. E., Burgin, A. B. & Osheroff, N. Human topoisomerase IIα uses a two-metal-ion mechanism for DNA cleavage. Nucleic Acids Res. 36, 4883–4893 (2008)
Noble, C. G. & Maxwell, A. The role of GyrB in the DNA cleavage-religation reaction of DNA gyrase: a proposed two metal-ion mechanism. J. Mol. Biol. 318, 361–371 (2002)
Sissi, C. & Palumbo, M. Effects of magnesium and related divalent metal ions in topoisomerase structure and function. Nucleic Acids Res. 37, 702–711 (2009)
Oezguen, N. et al. A “Moving Metal Mechanism” for substrate cleavage by the DNA repair endonuclease APE-1. Proteins 68, 313–323 (2007)
Berger, J. M., Fass, D., Wang, J. C. & Harrison, S. C. Structural similarities between topoisomerases that cleave one or both DNA strands. Proc. Natl Acad. Sci. USA 95, 7876–7881 (1998)
Changela, A., DiGate, R. J. & Mondragon, A. Crystal structure of a complex of a type IA DNA topoisomerase with a single-stranded DNA molecule. Nature 411, 1077–1081 (2001)
Changela, A., DiGate, R. J. & Mondragon, A. Structural studies of E. coli topoisomerase III-DNA complexes reveal a novel type IA topoisomerase-DNA conformational intermediate. J. Mol. Biol. 368, 105–118 (2007)
Mueller-Planitz, F. & Herschlag, D. Coupling between ATP binding and DNA cleavage by topoisomerase II: a unifying kinetic and structural mechanism. J. Biol. Chem. 283, 17463–17476 (2008)
Nitiss, J. L. Targeting DNA topoisomerase II in cancer chemotherapy. Nature Rev. Cancer 9, 338–350 (2009)
Schmidt, B. H., Burgin, A. B., Deweese, J. E., Osheroff, O. & Berger, J. M. A novel and unified two-metal mechanism for DNA cleavage by type II and 1A topoisomerases. Nature 465, 641–645 (2010)
DeLano, W. L. The PyMOL Molecular Graphics System. 〈http://www.pymol.org〉 (2008)
Miller, W. H. & Seefeld, M. A. Preparation of Naphthalenes, Quinolines, Quinoxalines and Naphthyridines as Antibacterial Agents. United States patent 7,691,850 B2 (2010)
Clinical and Laboratory Standards Institute. Methods for Dilution Antimicrobial Susceptibility Tests for Bacteria that Grow Aerobically; Approved Standard 8th edn (Clinical and Laboratory Standards Institute, 2008)
Lavasani, L. S. & Hiasa, H. A ParE–ParC fusion protein is a functional topoisomerase. Biochemistry 40, 8438–8443 (2001)
Trigueros, S. & Roca, J. A GyrB–GyrA fusion protein expressed in yeast cells is able to remove DNA supercoils but cannot substitute eukaryotic topoisomerase II. Genes Cells 7, 249–257 (2002)
Leslie, A. G. W. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 + ESF-EAMCB Newsl. Protein Crystallogr. 26 (1992)
Collaborative Computational Project, Number 4. The CCP4 suite: programmes for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Brünger, A. T. et al. Crystallography and NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Hansen, C. L., Skordalakes, E., Berger, J. M. & Quake, S. R. A robust and scalable microfluidic metering method that allows protein crystal growth by free interface diffusion. Proc. Natl Acad. Sci. USA 99, 16531–16536 (2002)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Kleywegt, G. J. & Jones, T. A. Where freedom is given liberties are taken. Structure 3, 535–540 (1995)
Critchlow, S. E. & Maxwell, A. DNA cleavage is not required for the binding of quinolone drugs to the DNA gyrase–DNA complex. Biochemistry 35, 7387–7393 (1996)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Acknowledgements
We thank H. Hiasa, A. Maxwell, J. C. Wang and D. B. Wigley for discussions. We thank S. Erskine, A. West and S. Brooks for experimental work that helped initiate this project. We thank P. Rowland for help with data collection and processing, the antimicrobial profiling group at GlaxoSmithKline (GSK) for antibacterial data, and K. Smith and D. Payne and members of the project team for discussions. A.W. was supported by the Wellcome Trust Seeding Drug Discovery Initiative and contract HDTRA1-07-9-0002 with the US Department of Defense (DoD) Joint Science and Technology Office for Chemical and Biological Defense (JSTO-CBD), and the Defense Threat Reduction Agency (DTRA) Transformational Medical Technologies (TMT). The views expressed in this article are those of the authors and do not necessarily reflect the official policy or position of the DoD or the US Government.
Author information
Authors and Affiliations
Contributions
M.N.G. and D.R.G. defined the relevant domains for crystallography by mutational analysis. K.K.B. performed gene sequence analysis and designed B27–A56 translation fusion. J.H. and E.W.M. made the initial fusion constructs, and tested them for activity. J.H., C.S. and P.F.C. performed studies with GSK299423 on target potency and the inhibition mechanism and demonstrated the lack of cross-resistance with quinolones. B.D.B. and M.R.S. designed the Greek key deletion. A.F. made the Greek-key-deletion and catalytic Tyr-to-Phe constructs. J.J. optimized expression in fermentors and grew cells for purification. M.H., E.J., A.S. and A.F.T. purified apo protein and complexes of various constructs for crystallography and assay. I.G. and A.H. designed and synthesized the compound. C.E.S. performed analytical size exclusion chromatography experiments that were used to find a stable particle that could be crystallized. F.G. crystallized apo and GSK299423 complex in fluidigm chips. O.S. and A.W. grew quinolone crystals by microbatch. B.D.B. grew apo and GSK299423 complex crystals by vapour diffusion, solved and refined structures. C.J.L. lead biochemistry. N.D.P. led chemistry and provided compound. D.S.E., M.N.G. and N.D.P. initiated and led the project. B.D.B. and M.N.G. wrote the manuscript with the assistance of M.M.H. and the other authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-4, Supplementary Figures 1-15 with legends, a Supplementary Discussion and References. (PDF 8970 kb)
Rights and permissions
About this article
Cite this article
Bax, B., Chan, P., Eggleston, D. et al. Type IIA topoisomerase inhibition by a new class of antibacterial agents. Nature 466, 935–940 (2010). https://doi.org/10.1038/nature09197
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09197
This article is cited by
-
Two classes of DNA gyrase inhibitors elicit distinct evolutionary trajectories toward resistance in gram-negative pathogens
npj Antimicrobials and Resistance (2024)
-
In Vitro and In Silico Evaluations of Cardanol and Derivatives from Cashew Nut-Shell
Chemistry Africa (2024)
-
Synthesis, characterization, antimicrobial activity and molecular docking study of transition metal complexes based on azo coumarin and thiosemicarbazone derivative
Journal of the Iranian Chemical Society (2024)
-
Efficacy and Safety of Gepotidacin as Treatment of Uncomplicated Urogenital Gonorrhea (EAGLE-1): Design of a Randomized, Comparator-Controlled, Phase 3 Study
Infectious Diseases and Therapy (2023)
-
Synthesis and molecular docking of new N4-piperazinyl ciprofloxacin hybrids as antimicrobial DNA gyrase inhibitors
Molecular Diversity (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.