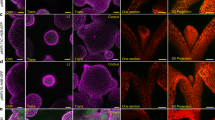
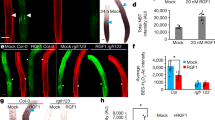
Abstract
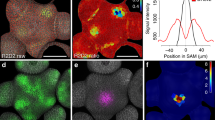
The classic phytohormones cytokinin and auxin play essential roles in the maintenance of stem-cell systems embedded in shoot and root meristems, and exhibit complex functional interactions1,2,3,4. Here we show that the activity of both hormones directly converges on the promoters of two A-type ARABIDOPSIS RESPONSE REGULATOR (ARR) genes, ARR7 and ARR15, which are negative regulators of cytokinin signalling5 and have important meristematic functions3. Whereas ARR7 and ARR15 expression in the shoot apical meristem (SAM) is induced by cytokinin, auxin has a negative effect, which is, at least in part, mediated by the AUXIN RESPONSE FACTOR5/MONOPTEROS (MP) transcription factor6. Our results provide a mechanistic framework for hormonal control of the apical stem-cell niche and demonstrate how root and shoot stem-cell systems differ in their response to phytohormones.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Müller, B. & Sheen, J. Cytokinin and auxin interaction in root stem-cell specification during early embryogenesis. Nature 453, 1094–1097 (2008)
Gordon, S. P., Chickarmane, V. S., Ohno, C. & Meyerowitz, E. M. Multiple feedback loops through cytokinin signaling control stem cell number within the Arabidopsis shoot meristem. Proc. Natl Acad. Sci. USA 106, 16529–16534 (2009)
Leibfried, A. et al. WUSCHEL controls meristem function by direct regulation of cytokinin-inducible response regulators. Nature 438, 1172–1175 (2005)
Heisler, M. G. et al. Patterns of auxin transport and gene expression during primordium development revealed by live imaging of the Arabidopsis inflorescence meristem. Curr. Biol. 15, 1899–1911 (2005)
To, J. P. et al. Type-A Arabidopsis response regulators are partially redundant negative regulators of cytokinin signaling. Plant Cell 16, 658–671 (2004)
Hardtke, C. S. & Berleth, T. The Arabidopsis gene MONOPTEROS encodes a transcription factor mediating embryo axis formation and vascular development. EMBO J. 17, 1405–1411 (1998)
Mayer, K. F. et al. Role of WUSCHEL in regulating stem cell fate in the Arabidopsis shoot meristem. Cell 95, 805–815 (1998)
Fletcher, J. C., Brand, U., Running, M. P., Simon, R. & Meyerowitz, E. M. Signaling of cell fate decisions by CLAVATA3 in Arabidopsis shoot meristems. Science 283, 1911–1914 (1999)
Schoof, H. et al. The stem cell population of Arabidopsis shoot meristems in maintained by a regulatory loop between the CLAVATA and WUSCHEL genes. Cell 100, 635–644 (2000)
Brand, U., Fletcher, J. C., Hobe, M., Meyerowitz, E. M. & Simon, R. Dependence of stem cell fate in Arabidopsis on a feedback loop regulated by CLV3 activity. Science 289, 617–619 (2000)
Ohyama, K., Shinohara, H., Ogawa-Ohnishi, M. & Matsubayashi, Y. A glycopeptide regulating stem cell fate in Arabidopsis thaliana. Nature Chem. Biol. 5, 578–580 (2009)
Reinhardt, D. et al. Regulation of phyllotaxis by polar auxin transport. Nature 426, 255–260 (2003)
Kurakawa, T. et al. Direct control of shoot meristem activity by a cytokinin-activating enzyme. Nature 445, 652–655 (2007)
Riou-Khamlichi, C., Huntley, R., Jacqmard, A. & Murray, J. A. Cytokinin activation of Arabidopsis cell division through a D-type cyclin. Science 283, 1541–1544 (1999)
Giulini, A., Wang, J. & Jackson, D. Control of phyllotaxy by the cytokinin-inducible response regulator homologue ABPHYL1. Nature 430, 1031–1034 (2004)
Dello Ioio, R. et al. A genetic framework for the control of cell division and differentiation in the root meristem. Science 322, 1380–1384 (2008)
Schwab, R., Ossowski, S., Riester, M., Warthmann, N. & Weigel, D. Highly specific gene silencing by artificial microRNAs in Arabidopsis. Plant Cell 18, 1121–1133 (2006)
Cheng, Y., Dai, X. & Zhao, Y. Auxin biosynthesis by the YUCCA flavin monooxygenases controls the formation of floral organs and vascular tissues in Arabidopsis. Genes Dev. 20, 1790–1799 (2006)
Yanai, O. et al. Arabidopsis KNOXI proteins activate cytokinin biosynthesis. Curr. Biol. 15, 1566–1571 (2005)
Jasinski, S. et al. KNOX action in Arabidopsis is mediated by coordinate regulation of cytokinin and gibberellin activities. Curr. Biol. 15, 1560–1565 (2005)
Furutani, M. et al. PIN-FORMED1 and PINOID regulate boundary formation and cotyledon development in Arabidopsis embryogenesis. Development 131, 5021–5030 (2004)
Kieber, J. J. in The Arabidopsis Book (American Society of Plant Biologists, 2002)
Hwang, I. & Sheen, J. Two-component circuitry in Arabidopsis cytokinin signal transduction. Nature 413, 383–389 (2001)
Schmid, M. et al. A gene expression map of Arabidopsis thaliana development. Nature Genet. 37, 501–506 (2005)
Guilfoyle, T. J. & Hagen, G. Auxin response factors. Curr. Opin. Plant Biol. 10, 453–460 (2007)
Schlereth, A. et al. MONOPTEROS controls embryonic root initiation by regulating a mobile transcription factor. Nature 464, 913–917 (2010)
Skoog, F. & Miller, C. O. Chemical regulation of growth and organ formation in plant tissues cultured in vitro. Symp. Soc. Exp. Biol. 54, 118–130 (1957)
Vidaurre, D. P., Ploense, S., Krogan, N. T. & Berleth, T. AMP1 and MP antagonistically regulate embryo and meristem development in Arabidopsis. Development 134, 2561–2567 (2007)
Andersen, S. U. et al. Requirement of B2-type cyclin-dependent kinases for meristem integrity in Arabidopsis thaliana. Plant Cell 20, 88–100 (2008)
Ulmasov, T., Hagen, G. & Guilfoyle, T. J. Dimerization and DNA binding of auxin response factors. Plant J. 19, 309–319 (1999)
Weigel, D. & Glazebrook, J. Arabidopsis: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 2002)
Wu, Z., Irizarry, R., Gentleman, R., Martinez-Murillo, F. & Spencer, F. A. Model-based background adjustment for oligonucleotide expression arrays. J. Am. Stat. Assoc. 99, 909–917 (2004)
Speed, T. P. Statistical Analysis of Gene Expression Microarray Data (CRC Press, 2003)
Nordstrom, A. et al. Derivatization for LC-electrospray ionization-MS: a tool for improving reversed-phase separation and ESI responses of bases, ribosides, and intact nucleotides. Anal. Chem. 76, 2869–2877 (2004)
Acknowledgements
We thank G. Jürgens, K. Schneitz and D. Weigel for reading the manuscript, and I. Weißwange for imaging. This work was supported by a Human Frontier Science Program Career Development Award, an EMBO Young Investigator Award, the SFB446 (all to J.U.L.), a Carlsberg Foundation fellowship (to S.U.A.), a Czech Ministry of Education grant (MSM 6198959216) (to K.D.) and the Max Planck Society.
Author information
Authors and Affiliations
Contributions
S.U.A. established the AM7/15 line and performed the microarray experiment, K.L. and K.D. quantified cytokinin content, A.M. performed ChIP experiments, S.J.S performed bioinformatic analyses and Z.Z. performed all other experiments. Z.Z and J.U.L. designed experiments and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-16 with legends, Supplementary Table 1 and a Reference. (PDF 4262 kb)
Rights and permissions
About this article
Cite this article
Zhao, Z., Andersen, S., Ljung, K. et al. Hormonal control of the shoot stem-cell niche. Nature 465, 1089–1092 (2010). https://doi.org/10.1038/nature09126
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature09126
This article is cited by
-
Transcriptome analysis of axillary buds in low phosphorus stress and functional analysis of TaWRKY74s in wheat
BMC Plant Biology (2024)
-
FRIZZLE PANICLE (FZP) regulates rice spikelets development through modulating cytokinin metabolism
BMC Plant Biology (2023)
-
LkARF7 and LkARF19 overexpression promote adventitious root formation in a heterologous poplar model by positively regulating LkBBM1
Communications Biology (2023)
-
Gene co-expression modules behind the three-pistil formation in wheat
Functional & Integrative Genomics (2023)
-
Promoting genotype-independent plant transformation by manipulating developmental regulatory genes and/or using nanoparticles
Plant Cell Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.