Abstract
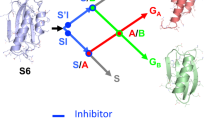
The three-dimensional structures of proteins often show a modular architecture comprised of discrete structural regions or domains. Cooperative communication between these regions is important for catalysis, regulation and efficient folding; lack of coupling has been implicated in the formation of fibrils and other misfolding pathologies1. How different structural regions of a protein communicate and contribute to a protein’s overall energetics and folding, however, is still poorly understood. Here we use a single-molecule optical tweezers approach to induce the selective unfolding of particular regions of T4 lysozyme and monitor the effect on other regions not directly acted on by force. We investigate how the topological organization of a protein (the order of structural elements along the sequence) affects the coupling and folding cooperativity between its domains. To probe the status of the regions not directly subjected to force, we determine the free energy changes during mechanical unfolding using Crooks’ fluctuation theorem. We pull on topological variants (circular permutants) and find that the topological organization of the polypeptide chain critically determines the folding cooperativity between domains and thus what parts of the folding/unfolding landscape are explored. We speculate that proteins may have evolved to select certain topologies that increase coupling between regions to avoid areas of the landscape that lead to kinetic trapping and misfolding.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Chiti, F. & Dobson, C. M. Protein misfolding, functional amyloid, and human disease. Annu. Rev. Biochem. 75, 333–366 (2006)
Ng, S. P., Randles, L. G. & Clarke, J. Single molecule studies of protein folding using atomic force microscopy. Methods Mol. Biol. 350, 139–167 (2007)
Borgia, A., Williams, P. M. & Clarke, J. Single-molecule studies of protein folding. Annu. Rev. Biochem. 77, 101–125 (2008)
Fisher, T. E., Marszalek, P. E. & Fernandez, J. M. Stretching single molecules into novel conformations using the atomic force microscope. Nature Struct. Biol. 7, 719–724 (2000)
Bustamante, C., Chemla, Y. R., Forde, N. R. & Izhaky, D. Mechanical processes in biochemistry. Annu. Rev. Biochem. 73, 705–748 (2004)
Brockwell, D. J. et al. Pulling geometry defines the mechanical resistance of a β-sheet protein. Nature Struct. Mol. Biol. 10, 731–737 (2003)
Carrion-Vazquez, M. et al. The mechanical stability of ubiquitin is linkage dependent. Nature Struct. Mol. Biol. 10, 738–743 (2003)
Dietz, H. & Rief, M. Protein structure by mechanical triangulation. Proc. Natl Acad. Sci. USA 103, 1244–1247 (2006)
Wetlaufer, D. B. Nucleation, rapid folding, and globular intrachain regions in proteins. Proc. Natl Acad. Sci. USA 70, 697–701 (1973)
Elwell, M. & Schellman, J. Phage T4 lysozyme. Physical properties and reversible unfolding. Biochim. Biophys. Acta 386, 309–323 (1975)
Mchaourab, H. S., Oh, K. J., Fang, C. J. & Hubbell, W. L. Conformation of T4 lysozyme in solution. Hinge-bending motion and the substrate-induced conformational transition studied by site-directed spin labeling. Biochemistry 36, 307–316 (1997)
Anderson, D. E., Lu, J., McIntosh, L. & Dahlquist, F. W. in NMR of Proteins (eds Clore, G. M. & Gronenborn, A. M.) ch. 9 (CRC Press, 1993)
Zhang, X. J., Wozniak, J. A. & Matthews, B. W. Protein flexibility and adaptability seen in 25 crystal forms of T4 lysozyme. J. Mol. Biol. 250, 527–552 (1995)
Llinás, M., Gillespie, B., Dahlquist, F. W. & Marqusee, S. The energetics of T4 lysozyme reveal a hierarchy of conformations. Nature Struct. Biol. 6, 1072–1078 (1999)
Llinás, M. & Marqusee, S. Subdomain interactions as a determinant in the folding and stability of T4 lysozyme. Protein Sci. 7, 96–104 (1998)
Cellitti, J. et al. Exploring subdomain cooperativity in T4 lysozyme I: Structural and energetic studies of a circular permutant and protein fragment. Protein Sci. 16, 842–851 (2007)
Cecconi, C., Shank, E. A., Bustamante, C. & Marqusee, S. Direct observation of the three-state folding of a single protein molecule. Science 309, 2057–2060 (2005)
Cecconi, C., Shank, E. A., Dahlquist, F. W., Marqusee, S. & Bustamante, C. Protein-DNA chimeras for single molecule mechanical folding studies with the optical tweezers. Eur. Biophys. J. 37, 729–738 (2008)
Bustamante, C., Marko, J. F., Siggia, E. D. & Smith, S. Entropic elasticity of lambda-phage DNA. Science 265, 1599–1600 (1994)
Yang, G. et al. Solid-state synthesis and mechanical unfolding of polymers of T4 lysozyme. Proc. Natl Acad. Sci. USA 97, 139–144 (2000)
Peng, Q. & Li, H. Atomic force microscopy reveals parallel mechanical unfolding pathways of T4 lysozyme: Evidence for a kinetic partitioning mechanism. Proc. Natl Acad. Sci. USA 105, 1885–1890 (2008)
Crooks, G. E. Entropy production fluctuation theorem and the nonequilibrium work relation for free energy differences. Phys. Rev. E 60, 2721–2726 (1999)
Collin, D. et al. Verification of the Crooks fluctuation theorem and recovery of RNA folding free energies. Nature 437, 231–234 (2005)
Bornschlögl, T. & Rief, M. Single-molecule dynamics of mechanical coiled-coil unzipping. Langmuir 24, 1338–1342 (2008)
Booth, D. R. et al. Instability, unfolding and aggregation of human lysozyme variants underlying amyloid fibrillogenesis. Nature 385, 787–793 (1997)
Dumoulin, M. et al. A camelid antibody fragment inhibits the formation of amyloid fibrils by human lysozyme. Nature 424, 783–788 (2003)
Lo, W. C., Lee, C. C., Lee, C. Y. & Lyu, P. C. CPDB: a database of circular permutation in proteins. Nucleic Acids Res. 37, D328–D332 (2009)
Smith, S. B., Cui, Y. & Bustamante, C. Overstretching B-DNA: the elastic response of individual double-stranded and single-stranded DNA molecules. Science 271, 795–799 (1996)
Acknowledgements
We would like to thank G. Crooks for assistance in use of Crooks fluctuation analysis, R. Dahlquist and B. Matthews for help in initiating this study, and the entire Bustamante and Marqusee labs for advice and technical help. We would particularly like to thank E. Kwon for her assistance in reagent preparation and data collection. This work was supported in part by NIH grants GM 32543 (C.B.), GM 50945 (S.M.) and a grant from the NSF (S.M.).
Author information
Authors and Affiliations
Contributions
E.A.S., C.C., J.W.D. conducted the experiments and analysed the data. J.W.D. carried out the Crooks fluctuation analysis. E.A.S., J.W.D., S.M. and C.B. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-6 with legends, Supplementary Tables 1-2 and References. (PDF 836 kb)
Rights and permissions
About this article
Cite this article
Shank, E., Cecconi, C., Dill, J. et al. The folding cooperativity of a protein is controlled by its chain topology. Nature 465, 637–640 (2010). https://doi.org/10.1038/nature09021
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature09021
This article is cited by
-
Folding pathway of a discontinuous two-domain protein
Nature Communications (2024)
-
The role of single-protein elasticity in mechanobiology
Nature Reviews Materials (2022)
-
A topology framework for macromolecular complexes and condensates
Nano Research (2022)
-
Forces of Change: Optical Tweezers in Membrane Remodeling Studies
The Journal of Membrane Biology (2022)
-
Optical tweezers in single-molecule biophysics
Nature Reviews Methods Primers (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.