Abstract
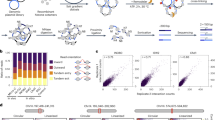
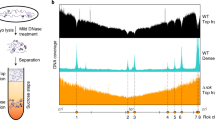
Layered on top of information conveyed by DNA sequence and chromatin are higher order structures that encompass portions of chromosomes, entire chromosomes, and even whole genomes1,2,3. Interphase chromosomes are not positioned randomly within the nucleus, but instead adopt preferred conformations4,5,6,7. Disparate DNA elements co-localize into functionally defined aggregates or ‘factories’ for transcription8 and DNA replication9. In budding yeast, Drosophila and many other eukaryotes, chromosomes adopt a Rabl configuration, with arms extending from centromeres adjacent to the spindle pole body to telomeres that abut the nuclear envelope10,11,12. Nonetheless, the topologies and spatial relationships of chromosomes remain poorly understood. Here we developed a method to globally capture intra- and inter-chromosomal interactions, and applied it to generate a map at kilobase resolution of the haploid genome of Saccharomyces cerevisiae. The map recapitulates known features of genome organization, thereby validating the method, and identifies new features. Extensive regional and higher order folding of individual chromosomes is observed. Chromosome XII exhibits a striking conformation that implicates the nucleolus as a formidable barrier to interaction between DNA sequences at either end. Inter-chromosomal contacts are anchored by centromeres and include interactions among transfer RNA genes, among origins of early DNA replication and among sites where chromosomal breakpoints occur. Finally, we constructed a three-dimensional model of the yeast genome. Our findings provide a glimpse of the interface between the form and function of a eukaryotic genome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Data deposits
Sequencing data have been deposited in the Sequence Read Archive under accession number SRP002120. An interactive website for yeast chromosomal interactions can be found at http://noble.gs.washington.edu/proj/yeast-architecture.
References
Misteli, T. Beyond the sequence: cellular organization of genome function. Cell 128, 787–800 (2007)
Lanctôt, C., Cheutin, T., Cremer, M., Cavalli, G. & Cremer, T. Dynamic genome architecture in the nuclear space: regulation of gene expression in three dimensions. Nature Rev. Genet. 8, 104–115 (2007)
Zhao, R., Bodnar, M. S. & Spector, D. L. Nuclear neighborhoods and gene expression. Curr. Opin. Genet. Dev. 19, 172–179 (2009)
Heun, P., Laroche, T., Shimada, K., Furrer, P. & Gasser, S. M. Chromosome dynamics in the yeast interphase nucleus. Science 294, 2181–2186 (2001)
Gasser, S. M. Visualizing chromatin dynamics in interphase nuclei. Science 296, 1412–1416 (2002)
Stone, E. M., Heun, P., Laroche, T., Pillus, L. & Gasser, S. M. MAP kinase signaling induces nuclear reorganization in budding yeast. Curr. Biol. 10, 373–382 (2000)
Casolari, J. M., Brown, C. R., Drubin, D. A., Rando, O. J. & Silver, P. A. Developmentally induced changes in transcriptional program alter spatial organization across chromosomes. Genes Dev. 19, 1188–1198 (2005)
Osborne, C. S. et al. Active genes dynamically colocalize to shared sites of ongoing transcription. Nature Genet. 36, 1065–1071 (2004)
Kitamura, E., Blow, J. J. & Tanaka, T. U. Live-cell imaging reveals replication of individual replicons in eukaryotic replication factories. Cell 125, 1297–1308 (2006)
Bystricky, K., Laroche, T., van Houwe, G., Blaszczyk, M. & Gasser, S. M. Chromosome looping in yeast: telomere pairing and coordinated movement reflect anchoring efficiency and territorial organization. J. Cell Biol. 168, 375–387 (2005)
Schober, H. et al. Controlled exchange of chromosomal arms reveals principles driving telomere interactions in yeast. Genome Res. 18, 261–271 (2008)
Berger, A. B. et al. High-resolution statistical mapping reveals gene territories in live yeast. Nature Methods 5, 1031–1037 (2008)
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002)
Simonis, M. et al. Nuclear organization of active and inactive chromatin domains uncovered by chromosome conformation capture-on-chip (4C). Nature Genet. 38, 1348–1354 (2006)
Murrell, A., Heeson, S. & Reik, W. Interaction between differentially methylated regions partitions the imprinted genes Igf2 and H19 into parent-specific chromatin loops. Nature Genet. 36, 889–893 (2004)
Spilianakis, C. G., Lalioti, M. D., Town, T., Lee, G. R. & Flavell, R. A. Interchromosomal associations between alternatively expressed loci. Nature 435, 637–645 (2005)
Zhao, Z. et al. Circular chromosome conformation capture (4C) uncovers extensive networks of epigenetically regulated intra- and interchromosomal interactions. Nature Genet. 38, 1341–1347 (2006)
Fullwood, M. J. & Ruan, Y. ChIP-based methods for the identification of long-range chromatin interactions. J. Cell. Biochem. 107, 30–39 (2009)
Fullwood, M. J. et al. An oestrogen-receptor-α-bound human chromatin interactome. Nature 462, 58–64 (2009)
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009)
Simonis, M., Kooren, J. & de Laat, W. An evaluation of 3C-based methods to capture DNA interactions. Nature Methods 4, 895–901 (2007)
Venema, J. & Tollervey, D. Ribosome synthesis in Saccharomyces cerevisiae . Annu. Rev. Genet. 33, 261–311 (1999)
Jin, Q., Trelles-Sticken, E., Scherthan, H. & Loidl, J. Yeast nuclei display prominent centromere clustering that is reduced in nondividing cells and in meiotic prophase. J. Cell Biol. 141, 21–29 (1998)
Gotta, M. et al. The clustering of telomeres and colocalization with Rap1, Sir3, and Sir4 proteins in wild-type Saccharomyces cerevisiae . J. Cell Biol. 134, 1349–1363 (1996)
Haeusler, R. A., Pratt-Hyatt, M., Good, P. D., Gipson, T. A. & Engelke, D. R. Clustering of yeast tRNA genes is mediated by specific association of condensin with tRNA gene transcription complexes. Genes Dev. 22, 2204–2214 (2008)
Thompson, M., Haeusler, R. A., Good, P. D. & Engelke, D. R. Nucleolar clustering of dispersed tRNA genes. Science 302, 1399–1401 (2003)
Di Rienzi, S. C., Collingwood, D., Raghuraman, M. K. & Brewer, B. J. Fragile genomic sites are associated with origins of replication. Genome. Biol. Evol. 2009, 350–363 (2009)
Haber, J. E. & Leung, W. Y. Lack of chromosome territoriality in yeast: promiscuous rejoining of broken chromosome ends. Proc. Natl Acad. Sci. USA 93, 13949–13954 (1996)
Lorenz, A., Fuchs, J., Trelles-Sticken, E., Scherthan, H. & Loidl, J. Spatial organisation and behaviour of the parental chromosome sets in the nuclei of Saccharomyces cerevisiae × S. paradoxus hybrids. J. Cell Sci. 115, 3829–3835 (2002)
Bystricky, K., Heun, P., Gehlen, L., Langowski, J. & Gasser, S. M. Long-range compaction and flexibility of interphase chromatin in budding yeast analyzed by high-resolution imaging techniques. Proc. Natl Acad. Sci. USA 101, 16495–16500 (2004)
Acknowledgements
We appreciate the advice and assistance of M. Dorschner, the comments of S. Di Rienzi, B. Brewer and B. Byers, and the assistance of L. Zhang and G. Schroth (Illumina Inc.) in performing sequencing. We thank A. Brown for help with the 3D model. Supported by NIH grants P01GM081619, P41RR0011823, a post-doctoral fellowship (to M.A.) from the Natural Sciences and Engineering Research Council of Canada, and the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
Z.D. devised the strategy for characterizing genome architecture, Z.D., J.S, S.F, C.A.B. and W.S.N. designed experiments, Z.D., K.S., Y.J.K., and C.L. performed experiments, Z.D., M.A., S.M., J.S., S.F., C.A.B. and W.S.N. analysed experimental data, M.A., K.S., J.S. and W.S.N. commented on the manuscript drafts, Z.D., S.F., and C.A.B. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Results, Methods, Data Analysis, References and Validation of Methods, Supplementary Tables 1- 4 and 14 -15 (for Supplementary Tables 5-13 see separate excel files) and Supplementary Figures 1-18 with legends. Minor errors in this file were corrected on 19 May 2010. (PDF 11280 kb)
Supplementary Table 5
This file contains a list of intra-chromosomal interactions identified from HindIII libraries at the threshold of FDR 1%. (XLS 6770 kb)
Supplementary Table 6
This file contains a list of inter-chromosomal interactions identified from HindIII libraries at the threshold of FDR 1%. (XLS 23326 kb)
Supplementary Table 7
This file contains a list of intra-chromosomal interactions identified from EcoRI libraries at the threshold of FDR 1%. (XLS 3164 kb)
Supplementary Table 8
This file contains a list of inter-chromosomal interactions identified from EcoRI libraries at the threshold of FDR 1%. (XLS 6910 kb)
Supplementary Table 9
This file contains the statistical data with respect to intra- and inter-chromosomal interactions-HindIII. (XLS 27 kb)
Supplementary Table 10
This file contains a list of intra-chromosomal interactions between the 20 and 30 kb regions of the ends of the chromosomes. (XLS 54 kb)
Supplementary Table 11
This file contains a list of Inter-chromosomal telomere pairing. (XLS 146 kb)
Supplementary Table 12
This file contains a list of primers used in this project. (XLS 27 kb)
Supplementary Table 13
This file contains a list of mappable HindIII and EcoRI fragments in each chromosome. (XLS 392 kb)
Supplementary Information
This file contains a 3d model of the yeast genome. This file can be opened using Rasmol (http://rasmol.org/). (PDB 1840 kb)
Rights and permissions
About this article
Cite this article
Duan, Z., Andronescu, M., Schutz, K. et al. A three-dimensional model of the yeast genome. Nature 465, 363–367 (2010). https://doi.org/10.1038/nature08973
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08973
This article is cited by
-
Computational methods for analysing multiscale 3D genome organization
Nature Reviews Genetics (2024)
-
In vitro reconstitution of chromatin domains shows a role for nucleosome positioning in 3D genome organization
Nature Genetics (2024)
-
Disordered C-terminal domain drives spatiotemporal confinement of RNAPII to enhance search for chromatin targets
Nature Cell Biology (2024)
-
Conserved chromatin and repetitive patterns reveal slow genome evolution in frogs
Nature Communications (2024)
-
Does multi-way, long-range chromatin contact data advance 3D genome reconstruction?
BMC Bioinformatics (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.