Abstract
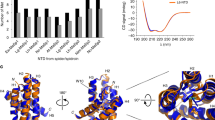
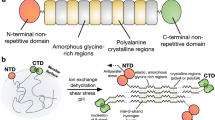
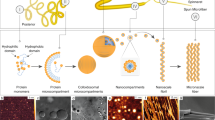
Nature’s high-performance polymer, spider silk, consists of specific proteins, spidroins, with repetitive segments flanked by conserved non-repetitive domains1,2. Spidroins are stored as a highly concentrated fluid dope. On silk formation, intermolecular interactions between repeat regions are established that provide strength and elasticity3,4. How spiders manage to avoid premature spidroin aggregation before self-assembly is not yet established. A pH drop to 6.3 along the spider’s spinning apparatus, altered salt composition and shear forces are believed to trigger the conversion to solid silk, but no molecular details are known. Miniature spidroins consisting of a few repetitive spidroin segments capped by the carboxy-terminal domain form metre-long silk-like fibres irrespective of pH5. We discovered that incorporation of the amino-terminal domain of major ampullate spidroin 1 from the dragline of the nursery web spider Euprosthenops australis (NT) into mini-spidroins enables immediate, charge-dependent self-assembly at pH values around 6.3, but delays aggregation above pH 7. The X-ray structure of NT, determined to 1.7 Å resolution, shows a homodimer of dipolar, antiparallel five-helix bundle subunits that lack homologues. The overall dimeric structure and observed charge distribution of NT is expected to be conserved through spider evolution and in all types of spidroins. Our results indicate a relay-like mechanism through which the N-terminal domain regulates spidroin assembly by inhibiting precocious aggregation during storage, and accelerating and directing self-assembly as the pH is lowered along the spider’s silk extrusion duct.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Candelas, G. C. & Cintron, J. A spider fibroin and its synthesis. J. Exp. Zool. 216, 1–6 (1981)
Ayoub, N. A., Garb, J. E., Tinghitella, R. M., Collin, M. A. & Hayashi, C. Y. Blueprint for a high-performance biomaterial: full-length spider dragline silk genes. PLoS ONE 2, e514
Gosline, J. M., Guerette, P. A., Ortlepp, C. S. & Savage, K. N. The mechanical design of spider silks: From fibroin sequence to mechanical function. J. Exp. Biol. 202, 3295–3303 (1999)
Ittah, S., Cohen, S., Garty, S., Cohn, D. & Gat, U. An essential role for the C-terminal domain of a dragline spider silk protein in directing fiber formation. Biomacromolecules 7, 1790–1795 (2006)
Stark, M. et al. Macroscopic fibers self-assembled from recombinant miniature spider silk proteins. Biomacromolecules 8, 1695–1701 (2007)
Hedhammar, M. et al. Structural properties of recombinant nonrepetitive and repetitive parts of major ampullate spidroin 1 from Euprosthenops australis: implications for fiber formation. Biochemistry 47, 3407–3417 (2008)
Holm, L. & Sander, C. Dali: a network tool for protein structure comparison. Trends Biochem. Sci. 20, 478–480 (1995)
Lin, Z., Huang, W., Zhang, J., Fan, J. S. & Yang, D. Solution structure of eggcase silk protein and its implications for silk fiber formation. Proc. Natl Acad. Sci. USA 106, 8906–8911 (2009)
Pace, C. N., Grimsley, G. R. & Scholtz, J. M. Protein ionizable groups: pK values and their contribution to protein stability and solubility. J. Biol. Chem. 284, 13285–13289 (2009)
Flocco, M. M. & Mowbray, S. L. Strange bedfellows: interactions between acidic side-chains in proteins. J. Mol. Biol. 254, 96–105 (1995)
Li, H., Robertson, A. D. & Jensen, J. H. Very fast empirical prediction and rationalization of protein pKa values. Proteins 61, 704–721 (2005)
Gordon, J.C. et al. H++: a server for estimating pK as and adding missing hydrogens to macromolecules. Nucleic Acids Res. 33, (Web Server issue) W368–W371 (2005)
Kabsch, W. A solution for the best rotation to relate two sets of vectors. Acta Crystallogr. A 32, 922–923 (1976)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Leslie, A. G. W. Recent changes to the MOSFLM package for processing film and image plate data. Joint CCP4 and ESF-EACBM Newsletter on Protein Crystallography no. 26, (1992)
Evans, P. R. SCALA. Joint CCP4 ESF-EACBM Newsletter Protein Crystallography 33, 22–24 (1997)
Weeks, C. M. et al. Automatic solution of heavy-atom substructures. Methods Enzymol. 374, 37–83 (2003)
Sheldrick, G. M. & Schneider, T. R. SHELXL: high-resolution refinement. Methods Enzymol. 277, 319–343 (1997)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D 58, 1948–1954 (2002)
Zwart, P. H. et al. Automated structure solution with the PHENIX suite. Methods Mol. Biol. 426, 419–435 (2008)
McCoy, A. J., Grosse-Kunstleve, R. W., Storoni, L. C. & Read, R. J. Likelihood-enhanced fast translation functions. Acta Crystallogr. D 61, 458–464 (2005)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Jones, T. A., Zou, J. Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Painter, J. & Merritt, E. A. Optimal description of a protein structure in terms of multiple groups undergoing TLS motion. Acta Crystallogr. D 62, 439–450 (2006)
Painter, J. & Merritt, E. A. TLSMD web server for the generation of multi-group TLS models. J. Appl. Cryst. 39, 109–111 (2006)
Rising, A., Hjalm, G., Engstrom, W. & Johansson, J. N-terminal nonrepetitive domain common to dragline, flagelliform, and cylindriform spider silk proteins. Biomacromolecules 7, 3120–3124 (2006)
Acknowledgements
This work was supported by grants (to J.J., M.H. and S.D.K.) from the Swedish Research Council and FORMAS. We thank E. Andersson, K. Tars and D. Ericsson for collecting the high resolution wild-type X-ray diffraction data set, H. Bramfeldt and C. Aulin for help with collecting scanning electron microscopy images, L. Serpell, I. Andersson, A. Zavialov and T. Härd for comments on the manuscript, and a number of other colleagues for discussions. We thank the ESRF (Grenoble, France) beamline staff for help during data collection.
Author information
Authors and Affiliations
Contributions
G.A. and M.H. contributed equally to this work. G.A., M.H. performed experiments and wrote the paper, K.N., A.S. performed experiments, C.C., A.R., J.J., S.D.K. discussed experiments and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
M.H., K.N., A.R. and J.J. own stocks and are funded by Spiber Technologies AB, a company that aims to commercialize recombinant spider silk for biomedical applications.
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-8 with legends and Supplementary Tables 1-2. (PDF 607 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Askarieh, G., Hedhammar, M., Nordling, K. et al. Self-assembly of spider silk proteins is controlled by a pH-sensitive relay. Nature 465, 236–238 (2010). https://doi.org/10.1038/nature08962
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08962
This article is cited by
-
Replicating shear-mediated self-assembly of spider silk through microfluidics
Nature Communications (2024)
-
Regionalization of cell types in silk glands of Larinioides sclopetarius suggest that spider silk fibers are complex layered structures
Scientific Reports (2023)
-
Bi-terminal fusion of intrinsically-disordered mussel foot protein fragments boosts mechanical strength for protein fibers
Nature Communications (2023)
-
NMR assignment and dynamics of the dimeric form of soluble C-terminal domain major ampullate spidroin 2 from Latrodectus hesperus
Biomolecular NMR Assignments (2023)
-
Massive production of fibroin nano-fibrous biomaterial by turbulent co-flow
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.