Abstract
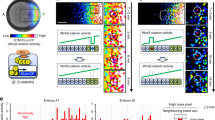
The Notch–Delta signalling pathway allows communication between neighbouring cells during development1. It has a critical role in the formation of ‘fine-grained’ patterns, generating distinct cell fates among groups of initially equivalent neighbouring cells and sharply delineating neighbouring regions in developing tissues2,3,4,5. The Delta ligand has been shown to have two activities: it transactivates Notch in neighbouring cells and cis-inhibits Notch in its own cell. However, it remains unclear how Notch integrates these two activities and how the resulting system facilitates pattern formation. Here we report the development of a quantitative time-lapse microscopy platform for analysing Notch–Delta signalling dynamics in individual mammalian cells, with the aim of addressing these issues. By controlling both cis- and trans-Delta concentrations, and monitoring the dynamics of a Notch reporter, we measured the combined cis–trans input–output relationship in the Notch–Delta system. The data revealed a striking difference between the responses of Notch to trans- and cis-Delta: whereas the response to trans-Delta is graded, the response to cis-Delta is sharp and occurs at a fixed threshold, independent of trans-Delta. We developed a simple mathematical model that shows how these behaviours emerge from the mutual inactivation of Notch and Delta proteins in the same cell. This interaction generates an ultrasensitive switch between mutually exclusive sending (high Delta/low Notch) and receiving (high Notch/low Delta) signalling states. At the multicellular level, this switch can amplify small differences between neighbouring cells even without transcription-mediated feedback. This Notch–Delta signalling switch facilitates the formation of sharp boundaries and lateral-inhibition patterns in models of development, and provides insight into previously unexplained mutant behaviours.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Artavanis-Tsakonas, S., Rand, M. D. & Lake, R. J. Notch signaling: cell fate control and signal integration in development. Science 284, 770–776 (1999)
Goodyear, R. & Richardson, G. Pattern formation in the basilar papilla: evidence for cell rearrangement. J. Neurosci. 17, 6289–6301 (1997)
Heitzler, P. & Simpson, P. The choice of cell fate in the epidermis of Drosophila. Cell 64, 1083–1092 (1991)
Huppert, S. S., Jacobsen, T. L. & Muskavitch, M. A. Feedback regulation is central to Delta-Notch signalling required for Drosophila wing vein morphogenesis. Development 124, 3283–3291 (1997)
de Celis, J. F., Bray, S. & Garcia-Bellido, A. Notch signalling regulates veinlet expression and establishes boundaries between veins and interveins in the Drosophila wing. Development 124, 1919–1928 (1997)
Bray, S. J. Notch signalling: a simple pathway becomes complex. Nature Rev. Mol. Cell Biol. 7, 678–689 (2006)
de Celis, J. F. & Bray, S. Feed-back mechanisms affecting Notch activation at the dorsoventral boundary in the Drosophila wing. Development 124, 3241–3251 (1997)
Micchelli, C. A., Rulifson, E. J. & Blair, S. S. The function and regulation of cut expression on the wing margin of Drosophila: Notch, Wingless and a dominant negative role for Delta and Serrate. Development 124, 1485–1495 (1997)
Klein, T., Brennan, K. & Arias, A. M. An intrinsic dominant negative activity of serrate that is modulated during wing development in Drosophila. Dev. Biol. 189, 123–134 (1997)
Miller, A. C., Lyons, E. L. & Herman, T. G. cis-Inhibition of notch by endogenous delta biases the outcome of lateral inhibition. Curr. Biol. 19, 1378–1383 (2009)
Cordle, J. et al. A conserved face of the Jagged/Serrate DSL domain is involved in Notch trans-activation and cis-inhibition. Nature Struct. Mol. Biol. 15, 849–857 (2008)
Matsuda, M. & Chitnis, A. B. Interaction with Notch determines endocytosis of specific Delta ligands in zebrafish neural tissue. Development 136, 197–206 (2009)
Kakidani, H. & Ptashne, M. GAL4 activates gene expression in mammalian cells. Cell 52, 161–167 (1988)
Struhl, G. & Adachi, A. Nuclear access and action of notch in vivo. Cell 93, 649–660 (1998)
Aster, J. C. et al. Essential roles for ankyrin repeat and transactivation domains in induction of T-cell leukemia by notch1. Mol. Cell. Biol. 20, 7505–7515 (2000)
Yang, L. T. et al. Fringe glycosyltransferases differentially modulate Notch1 proteolysis induced by Delta1 and Jagged1. Mol. Biol. Cell 16, 927–942 (2005)
Rothenberg, E. V., Moore, J. E. & Yui, M. A. Launching the T-cell-lineage developmental programme. Nature Rev. Immunol. 8, 9–21 (2008)
Varnum-Finney, B. et al. Immobilization of Notch ligand, Delta-1, is required for induction of notch signaling. J. Cell Sci. 113, 4313–4318 (2000)
Wang, S. et al. Notch receptor activation inhibits oligodendrocyte differentiation. Neuron 21, 63–75 (1998)
Rosenfeld, N., Young, J. W., Alon, U., Swain, P. S. & Elowitz, M. B. Gene regulation at the single-cell level. Science 307, 1962–1965 (2005)
Jacobsen, T. L., Brennan, K., Arias, A. M. & Muskavitch, M. A. Cis-interactions between Delta and Notch modulate neurogenic signalling in Drosophila. Development 125, 4531–4540 (1998)
Shaye, D. D. & Greenwald, I. LIN-12/Notch trafficking and regulation of DSL ligand activity during vulval induction in Caenorhabditis elegans. Development 132, 5081–5092 (2005)
de Celis, J. F. & Bray, S. J. The Abruptex domain of Notch regulates negative interactions between Notch, its ligands and Fringe. Development 127, 1291–1302 (2000)
Collier, J. R., Monk, N. A., Maini, P. K. & Lewis, J. H. Pattern formation by lateral inhibition with feedback: a mathematical model of Delta-Notch intercellular signalling. J. Theor. Biol. 183, 429–446 (1996)
Plahte, E. Pattern formation in discrete cell lattices. J. Math. Biol. 43, 411–445 (2001)
Melen, G. J., Levy, S., Barkai, N. & Shilo, B. Z. Threshold responses to morphogen gradients by zero-order ultrasensitivity. Mol. Syst. Biol. 1 10.1038/msb4100036 (2005)
Levine, E., Zhang, Z., Kuhlman, T. & Hwa, T. Quantitative characteristics of gene regulation by small RNA. PLoS Biol. 5, e229 (2007)
Buchler, N. E. & Louis, M. Molecular titration and ultrasensitivity in regulatory networks. J. Mol. Biol. 384, 1106–1119 (2008)
Lenz, D. H. et al. The small RNA chaperone Hfq and multiple small RNAs control quorum sensing in Vibrio harveyi and Vibrio cholerae. Cell 118, 69–82 (2004)
Acknowledgements
We would like to thank I. Bernstein for the IgG-Deltaext, U. Lendahl for the 12xCSL reporter construct, J. Aster for human NOTCH1 and other constructs, and G. Weinmaster for the rat DLL1 construct and advice. We also thank R. Tsien and K. Thorn for mCherry, S. Megason and S. Fraser for H2B–citrine and other constructs, R. Diamond and D. Perez for assistance with FACS, and F. Tan and J. Yong for help with cloning some of the constructs. We thank A. Eldar, J. Locke, G. Seelig, R. Kishony, B. Shraiman, A. C. Oates and members of the Elowitz laboratory for discussions and advice. This work was supported by the US National Institutes of Health Fellowship F32GM77014 (D.S.), the Caltech Center for Biological Circuit Design and the Packard Foundation. A.L. acknowledges support from the Fannie and John Hertz Foundation and the UCLA/Caltech Medical Scientist Training Program (NIH GM08042). J.G.O. acknowledges support from the Ministerio de Ciencia e Innovacion (Spain, project FIS2009-13360 and the I3 programme).
Author information
Authors and Affiliations
Contributions
D.S. and M.B.E. designed the research. D.S., L.A.S., M.E.F. and G.A.A. built cell lines and performed experiments. D.S., A.L., L.L., J.G.-O. and M.B.E. performed data analysis and mathematical modelling. D.S. and M.B.E. wrote the manuscript with substantial input from the other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S17 with legends, Supplementary Tables S1-S3, Supplementary Methods and Data Analysis and Data and References for the Supplementary Material. (PDF 3625 kb)
Supplementary Movie 1
This movie shows the trans-activation of hN1G4esn by plate bound Delta. Movie used to generate filmstrip in Figure 2B. (AVI 10479 kb)
Supplementary Movie 2
This movie shows the effect of cis-Delta on hN1G4esn activation. Movie used to generate filmstrip in Figure 3B. (AVI 22000 kb)
Supplementary Movie 3
This movie shows the trans-activation of hN1G4esn-No-Delta by co-culture with Delta-expressing cells. Movie used to generate filmstrip in Fig S5. (AVI 5340 kb)
Rights and permissions
About this article
Cite this article
Sprinzak, D., Lakhanpal, A., LeBon, L. et al. Cis-interactions between Notch and Delta generate mutually exclusive signalling states. Nature 465, 86–90 (2010). https://doi.org/10.1038/nature08959
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08959
This article is cited by
-
Cis inhibition of NOTCH1 through JAGGED1 sustains embryonic hematopoietic stem cell fate
Nature Communications (2024)
-
Genetic interference with HvNotch provides new insights into the role of the Notch-signalling pathway for developmental pattern formation in Hydra
Scientific Reports (2024)
-
The meaning of ubiquitylation of the DSL ligand Delta for the development of Drosophila
BMC Biology (2023)
-
Affinity-matured DLL4 ligands as broad-spectrum modulators of Notch signaling
Nature Chemical Biology (2023)
-
Molecular sensing of mechano- and ligand-dependent adhesion GPCR dissociation
Nature (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.