Abstract
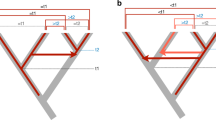
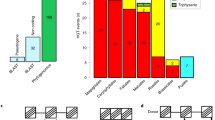
Horizontal transfer (HT), or the passage of genetic material between non-mating species, is increasingly recognized as an important force in the evolution of eukaryotic genomes1,2. Transposons, with their inherent ability to mobilize and amplify within genomes, may be especially prone to HT3,4,5,6,7. However, the means by which transposons can spread across widely diverged species remain elusive. Here we present evidence that host–parasite interactions have promoted the HT of four transposon families between invertebrates and vertebrates. We found that Rhodnius prolixus, a triatomine bug feeding on the blood of various tetrapods and vector of Chagas’ disease in humans, carries in its genome four distinct transposon families that also invaded the genomes of a diverse, but overlapping, set of tetrapods. The bug transposons are ∼98% identical and cluster phylogenetically with those of the opossum and squirrel monkey, two of its preferred mammalian hosts in South America. We also identified one of these transposon families in the pond snail Lymnaea stagnalis, a cosmopolitan vector of trematodes infecting diverse vertebrates, whose ancestral sequence is nearly identical and clusters with those found in Old World mammals. Together these data provide evidence for a previously hypothesized role of host–parasite interactions in facilitating HT among animals3,7. Furthermore, the large amount of DNA generated by the amplification of the horizontally transferred transposons supports the idea that the exchange of genetic material between hosts and parasites influences their genomic evolution.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Keeling, P. J. & Palmer, J. D. Horizontal transfer in eukaryotic evolution. Nature Rev. Genet. 9, 605–618 (2008)
Gladyshev, E. A., Meselson, M. & Arkhipova, I. R. Massive horizontal gene transfer in bdelloid rotifers. Science 320, 1210–1213 (2008)
Loreto, E. L., Carareto, C. M. & Capy, P. Revisiting horizontal transfer of transposable elements in Drosophila . Heredity 100, 545–554 (2008)
Diao, X., Freeling, M. & Lisch, D. Horizontal transfer of a plant transposon. PLoS Biol. 4, e5 (2006)
Pace, J. K., Gilbert, C., Clark, M. S. & Feschotte, C. Repeated horizontal transfer of a DNA transposon in mammals and other tetrapods. Proc. Natl Acad. Sci. USA 105, 17023–17028 (2008)
Novick, P. et al. Independent and parallel lateral transfer of DNA transposons in tetrapod genomes. Gene 449, 85–94 (2009)
Houck, M. A., Clark, J. B., Peterson, K. R. & Kidwell, M. G. Possible horizontal transfer of Drosophila genes by the mite Proctolaelaps regalis . Science 6, 1125–1129 (1991)
Lent, H. & Wygodzinsky, P. Revision of the Triatominae (Hemiptera, Reduviidae), and their significance as vectors of Chagas disease. Bull. Am. Mus. Nat. Hist. 163, 123–520 (1979)
Loy, C. & Haas, W. Prevalence of cercariae from Lymnaea stagnalis snails in a pond system in Southern Germany. Parasitol. Res. 87, 878–882 (2001)
Ray, D. A., Pagan, H. J. T., Thompson, M. L. & Stevens, R. D. Bats with hATs: Evidence for recent DNA transposon activity in genus Myotis . Mol. Biol. Evol. 24, 632–639 (2007)
Smit, A. F. A., Hubley, R. & Green, P. RepeatMasker Open-3.0 (1996–2004); 〈http://www.repeatmasker.org〉.
Robertson, H. M. in Mobile DNA II (eds Craig, N. L., Craigie, R., Gellert, M. & Lambowitz, A. M.) 1093–1110 (ASM, 2002)
Marshall, E. et al. Calibration of the great American interchange. Science 204, 272–279 (1979)
Maia Da Silva, F. et al. Comparative phylogeography of Trypanosoma rangeli and Rhodnius (Hemiptera: Reduviidae) supports a long coexistence of parasite lineages and their sympatric vectors. Mol. Ecol. 16, 3361–3373 (2007)
Maia Da Silva, F. et al. Trypanosoma rangeli isolates of bats from Central Brazil: genotyping and phylogenetic analysis enable description of a new lineage using spliced-leader gene sequences. Acta Trop. 109, 199–207 (2009)
Hecht, M. M. et al. Inheritance of DNA transferred from American trypanosomes to human hosts. PLoS ONE 5, e9181 (2010)
Rogan, M. T. et al. The occurrence of the trematode Plagiorchis muris in the wood mouse Apodemus sylvaticus in North Yorkshire, UK. J. Helminthol. 81, 57–62 (2007)
Zbikowska, E. et al. Infestation of Lymnaea stagnalis by Digenea flukes in the Jeziorak lake. Parasitol. Res. 99, 434–439 (2006)
Piskurek, O. & Okada, N. Poxviruses as possible vectors of horizontal transfer of retroposons from reptiles to mammals. Proc. Natl Acad. Sci. USA 104, 12046–12051 (2007)
Fraser, M. J., Smith, G. E. & Summers, M. D. Acquisition of host cell DNA sequences by baculoviruses: relationship between host DNA insertions and FP mutants of Autographa californica and Galleria mellonella nuclear polyhedris viruses. J. Virol. 47, 287–300 (1983)
Friesen, P. D. & Nissen, M. S. Gene organization and transcription of TED, a lepidopteran retrotransposon integrated within the baculovirus genome. Mol. Cell. Biol. 10, 3067–3077 (1990)
Jehle, J. A., Nickel, A., Vlak, J. M. & Backhaus, H. J. Horizontal escape of the novel Tc1-like lepidopteran transposon TCp3.2 into Cydia pomonella granulovirus. J. Mol. Evol. 46, 215–224 (1998)
Heath, B. D., Butcher, R. D. J., Whitfield, W. G. F. & Hubbard, S. F. Horizontal transfer of Wolbachia between phylogenetically distant insect species by a naturally occurring mechanism. Curr. Biol. 9, 313–316 (1999)
Huigens, M. E. et al. Infectious parthenogenesis. Nature 405, 178–179 (2000)
Yoshiyama, M. et al. Possible horizontal transfer of a transposable element from host to parasitoid. Mol. Biol. Evol. 18, 1952–1958 (2001)
Mower, J. P., Stefanovic, S., Young, G. J. & Palmer, J. D. Gene transfer from parasitic to host plants. Nature 432, 165–166 (2004)
Davis, C. C. & Wurdack, K. J. Host to parasite gene transfer in flowering plants: phylogenetic evidence from Malpighiales. Science 305, 676–678 (2004)
Feschotte, C. & Pritham, E. J. DNA transposons and the evolution of eukaryotic genomes. Annu. Rev. Genet. 41, 331–368 (2007)
Cordaux, R. & Batzer, M. A. The impact of retrotransposons on human genome evolution. Nature Rev. Genet. 10, 691–703 (2009)
Kumar, S. & Subramanian, S. Mutation rates in mammalian genomes. Proc. Natl Acad. Sci. USA 99, 803–808 (2002)
Altschul, S. F. et al. Basic Local Alignment Search Tool. J. Mol. Biol. 215, 403–410 (1990)
Jurka, J. et al. Repbase Update, a database of eukaryotic repetitive elements. Cytogenet. Genome Res. 110, 462–467 (2005)
Gentles, A. J. & Jurka, J. HAT2_MD — an autonomous hAT transposon from Monodelphis domestica; consensus sequence. Repbase Report 5, 10 (2005)
Ray, D. A. et al. Multiple waves of recent DNA transposon activity in the bat, Myotis lucifugus . Genome Res. 18, 717–728 (2008)
Jukes, T. H. & Cantor, C. R. in Mammalian Protein Metabolism (ed. Munro, H. N.) 21–132 (Academic, 1969)
Tamura, K., Dudley, J., Nei, M. & Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, 1596–1599 (2007)
Hedges, S. B., Dudley, J. & Kumar, S. TimeTree: a public knowledge-base of divergence times among organisms. Bioinformatics 22, 2971–2972 (2006)
Yamanoue, Y. et al. The mitochondrial genome of spotted green pufferfish Tetraodon nigroviridis (Teleostei: Tetraodontiformes) and divergence time estimation among model organisms in fishes. Genes Genet. Syst. 81, 29–39 (2006)
Steinke, D., Salzburger, W. & Meyer, A. Novel relationships among ten fish model species revealed based on a phylogenomic analysis using ESTs. J. Mol. Evol. 62, 772–784 (2006)
Cutter, A. D. Divergence times in Caenorhabditis and Drosophila inferred from direct estimates of the neutral mutation rate. Mol. Biol. Evol. 25, 778–786 (2008)
De Giorgi, C. et al. Structural and evolutionary analysis of the ribosomal genes of the parasitic nematode Meloidogyne artiellia suggests its ancient origin. Mol. Biochem. Parasitol. 124, 91–94 (2002)
Morgan, J. A. T. et al. Schistosoma mansoni and Biomphalaria: past history and future trends. Parasitology 123, S211–S228 (2001)
Wiegmann, B. M. et al. Single-copy nuclear genes resolve the phylogeny of the holometabolous insects. BMC Biol. 7, 34 (2009)
Yoder, A. D. et al. Ancient origin for Malagasy primates. Proc. Natl Acad. Sci. USA 93, 5122–5126 (1996)
Evans, B. J. et al. A mitochondrial DNA phylogeny of African clawed frogs: phylogeography and implication for polyploidy evolution. Mol. Phylogenet. Evol. 33, 197–213 (2004)
Wilson, D. E. & Reeder, D. M. Mammal Species of the World: A Taxonomic and Geographic Reference (Smithsonian Institution Press, 2005)
Poux, C. et al. Asynchronous colonization of Madagascar by the four endemic clades of primates, tenrecs, carnivores, and rodents as inferred from nuclear genes. Syst. Biol. 54, 719–730 (2005)
Gebo, D. L. et al. The oldest known anthropoid postcranial fossils and the early evolution of higher primates. Nature 404, 276–278 (2000)
Rossie, J. B., Ni, X. J. & Beard, K. C. Cranial remains of an Eocene tarsier. Proc. Natl Acad. Sci. USA 103, 4381–4385 (2006)
Steiner, C., Tilak, M., Douzery, E. J. P. & Catzeflis, F. M. New DNA data from a transthyretin nuclear intron suggest an Oligocene to Miocene diversification of living South America opossums (Marsupialia: Didelphidae). Mol. Phylogenet. Evol. 35, 363–379 (2005)
Roughgarden, J. Ed. Anolis Lizards of the Caribbean. Ecology, Evolution and Plate Tectonics (Oxford Univ. Press, 1995)
Stadelmann, B., Lin, L. K., Kunz, T. H. & Ruedi, M. Molecular phylogeny of New World Myotis (Chiroptera, Vespertilionidae) inferred from mitochondrial and nuclear DNA genes. Mol. Phylogenet. Evol. 43, 32–48 (2007)
Steppan, S. J., Adkins, R. M. & Anderson, J. Phylogeny and divergence-date estimates of rapid radiation in muroid rodents based on multiple nuclear genes. Syst. Biol. 53, 533–553 (2004)
Poux, C. et al. Arrival and diversification of caviomorph rodents and platyrrhine primates in South America. Syst. Biol. 55, 228–244 (2006)
Gaunt, M. W. & Miles, M. A. An insect molecular clock dates the origin of the insects and accords with palaeontological and biogeographic landmarks. Mol. Biol. Evol. 19, 748–761 (2002)
Remigio, E. A. Molecular phylogenetic relationships in the aquatic snail genus Lymnaea, the intermediate host of the causative agent of fascioliasis: insights from broader taxon sampling. Parasitol. Res. 88, 687–696 (2002)
Guindon, S. & Gascuel, O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, 696–704 (2003)
Hall, T. BioEdit version 5.0.6. 〈http://www.mbio.ncsu.edu/BioEdit/bioedit.html/〉 (2004)
Posada, D. & Crandall, K. A. MODELTEST: testing the model of DNA substitution. Bioinformatics 14, 817–818 (1998)
Jones, D. T., Taylor, W. R. & Thornton, J. M. The rapid generation of mutation data matrices from protein sequences. Comput. Appl. Biosci. 8, 275–282 (1992)
Librado, P. & Rozas, J. DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25, 1451–1452 (2009)
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994)
Acknowledgements
We thank E. Betrán, J. Demuth, T. Fondon, B. Koskella, J. Meik, E. Pritham, Q. Wang and members of the Feschotte laboratory for comments and suggestions during the preparation of the manuscript; M. Batzer, E. Dotson, S. Goodman, A. Prelat, T. Robinson, A. Ropiquet and the Grosell and Sánchez laboratories for the gifts of tissue samples used in this study; and J. Spieth and The Genome Center at Washington University School of Medicine in St Louis for permission to use the R. prolixus assembly before publication. C.F. is funded by the National Institutes of Health and S.S. by the National Science Foundation.
Author information
Authors and Affiliations
Contributions
C.G., S.S. and C.F. designed research, performed research, and analysed data. J.K.P. contributed data and perl scripts. P.J.B. contributed reagents and materials. C.G., S.S. and C.F. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-5 and Supplementary Figures 1-6 with legends and references. (PDF 1978 kb)
Supplementary Data
This file contains Supplementary Datasets 1-3. (PDF 761 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Gilbert, C., Schaack, S., Pace II, J. et al. A role for host–parasite interactions in the horizontal transfer of transposons across phyla. Nature 464, 1347–1350 (2010). https://doi.org/10.1038/nature08939
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08939
This article is cited by
-
Diversity of short interspersed nuclear elements (SINEs) in lepidopteran insects and evidence of horizontal SINE transfer between baculovirus and lepidopteran hosts
BMC Genomics (2021)
-
Mining histone methyltransferases and demethylases from whole genome sequence
Journal of Biosciences (2020)
-
Characterization of a novel Helitron family in insect genomes: insights into classification, evolution and horizontal transfer
Mobile DNA (2019)
-
High frequency of horizontal transfer in Jockey families (LINE order) of drosophilids
Mobile DNA (2019)
-
Horizontal transfer of a retrotransposon between parasitic nematodes and the common shrew
Mobile DNA (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.