Abstract
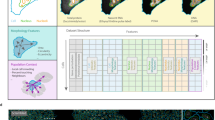
Despite our rapidly growing knowledge about the human genome, we do not know all of the genes required for some of the most basic functions of life. To start to fill this gap we developed a high-throughput phenotypic screening platform combining potent gene silencing by RNA interference, time-lapse microscopy and computational image processing. We carried out a genome-wide phenotypic profiling of each of the ∼21,000 human protein-coding genes by two-day live imaging of fluorescently labelled chromosomes. Phenotypes were scored quantitatively by computational image processing, which allowed us to identify hundreds of human genes involved in diverse biological functions including cell division, migration and survival. As part of the Mitocheck consortium, this study provides an in-depth analysis of cell division phenotypes and makes the entire high-content data set available as a resource to the community.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Neumann, B. et al. High-throughput RNAi screening by time-lapse imaging of live human cells. Nature Methods 3, 385–390 (2006)
Walter, T. et al. Automatic identification and clustering of chromosome phenotypes in a genome wide RNAi screen by time-lapse imaging. J. Struct. Biol. (in the press)
Boland, M. V., Markey, M. K. & Murphy, R. F. Automated recognition of patterns characteristic of subcellular structures in fluorescence microscopy images. Cytometry 33, 366–375 (1998)
Conrad, C. et al. Automatic identification of subcellular phenotypes on human cell arrays. Genome Res. 14, 1130–1136 (2004)
Glory, E. & Murphy, R. F. Automated subcellular location determination and high-throughput microscopy. Dev. Cell 12, 7–16 (2007)
Wang, M. et al. Novel cell segmentation and online SVM for cell cycle phase identification in automated microscopy. Bioinformatics 24, 94–101 (2008)
Sönnichsen, B. et al. Full-genome RNAi profiling of early embryogenesis in Caenorhabditis elegans . Nature 434, 462–469 (2005)
Björklund, M. et al. Identification of pathways regulating cell size and cell-cycle progression by RNAi. Nature 439, 1009–1013 (2006)
Kittler, R. et al. Genome-scale RNAi profiling of cell division in human tissue culture cells. Nature Cell Biol. 9, 1401–1412 (2007)
Kittler, R. et al. RNA interference rescue by bacterial artificial chromosome transgenesis in mammalian tissue culture cells. Proc. Natl Acad. Sci. USA 102, 2396–2401 (2005)
Poser, I. et al. BAC TransgeneOmics: a high-throughput method for exploration of protein function in mammals. Nature Methods 5, 409–415 (2008)
Raemaekers, T. et al. NuSAP, a novel microtubule-associated protein involved in mitotic spindle organization. J. Cell Biol. 162, 1017–1029 (2003)
Sumara, I. et al. Roles of polo-like kinase 1 in the assembly of functional mitotic spindles. Curr. Biol. 14, 1712–1722 (2004)
Gruss, O. J. et al. Chromosome-induced microtubule assembly mediated by TPX2 is required for spindle formation in HeLa cells. Nature Cell Biol. 4, 871–879 (2002)
Cassimeris, L. & Morabito, J. TOGp, the human homolog of XMAP215/Dis1, is required for centrosome integrity, spindle pole organization, and bipolar spindle assembly. Mol. Biol. Cell 15, 1580–1590 (2004)
Zhu, C. et al. Functional analysis of human microtubule-based motor proteins, the kinesins and dyneins, in mitosis/cytokinesis using RNA interference. Mol. Biol. Cell 16, 3187–3199 (2005)
Beaudouin, J., Gerlich, D., Daigle, N., Eils, R. & Ellenberg, J. Nuclear envelope breakdown proceeds by microtubule-induced tearing of the lamina. Cell 108, 83–96 (2002)
Glotzer, M. The molecular requirements for cytokinesis. Science 307, 1735–1739 (2005)
Steigemann, P. et al. Aurora B-mediated abscission checkpoint protects against tetraploidization. Cell 136, 473–484 (2009)
Mora-Bermúdez, F., Gerlich, D. & Ellenberg, J. Maximal chromosome compaction occurs by axial shortening in anaphase and depends on Aurora kinase. Nature Cell Biol. 9, 822–831 (2007)
Erfle, H. et al. Reverse transfection on cell arrays for high content screening microscopy. Nature Protocols 2, 392–399 (2007)
Liebel, U. et al. A microscope-based screening platform for large-scale functional protein analysis in intact cells. FEBS Lett. 554, 394–398 (2003)
Acknowledgements
We thank J. Gagneur for suggestions on data processing; S. Berthoumieux for assistance in computation; Y. Sun for coordination support in Mitocheck; U. Ringeisen for help in preparing the figures; S. Winkler and L. Burger and EMBL’s electronic and mechanical workshops for support in microscope development; Olympus Soft Imaging Solutions (OSIS) and Olympus Europe for collaboration; Leica Microsystems for collaboration; Applied Biosystems for providing unpublished validation data of the siRNA library; and all our colleagues in the Mitocheck consortium for collaboration. This project was funded by grants to J.E. (within the Mitocheck consortium by the European Commission (LSHG-CT-2004-503464) as well as by the Federal Ministry of Education and Research (BMBF) in the framework of the National Genome Research Network (NGFN) (NGFN-2 SMP-RNAi, FKZ01GR0403)), to R.P. (BMBF NGFN2 SMP-Cell, FKZ01GR0423) as well as to J.E. and R.P. by the Landesstiftung Baden Wuerttemberg in the framework of the research programme ‘RNS/RNAi’. R.D. is supported by the Wellcome Trust.
Author Contributions B.N. developed the primary screen assay. P.R., H.E., B.N. and J.B. generated the data for the primary and validation screen. T.W. and M.H. developed the image processing software. T.W., J.-K.H., G.P., and W.H. analysed the data. J.-K.H., V.S. and R.S. performed the bioinformatics analysis. B.N., P.R., J.B., T.W. and C.Co. conducted the quality control and manual annotation. U.L., C.Co., F.S. and R.P. developed the HT microscope platform. H.E. and R.P. developed the HT transfection platform. J.-K.H. and R.D. created the Mitocheck database and website. C.Ce. and R.P. designed the manual annotation database. J.B., C.Co., B.N. and A.W. created the data for the spindle assay. R.K. and R.E. provided IT support. I.P. and A.A.H. provided the BAC cell pools. M.H.A.S., C.Ch. and D.W.G. provided reagents. J.-M.P. coordinated the Mitocheck consortium. J.E. coordinated and supervised the project and wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary References, Supplementary Figures 1-11 with legends and Legends for Supplementary Movies 1-14. (PDF 2252 kb)
Supplementary Tables
This zipped file contains Supplementary Tables 1-7. (ZIP 580 kb)
Supplementary Movies 1
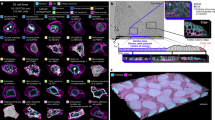
This zipped file contains Supplementary Movies 1-15 which show an overview of the representative morphology for each of the sixteen classes shown in Figure 1b. (ZIP 32444 kb)
Supplementary Movies 2
This zipped file contains Supplementary Movies 16-30 which show the highlighted single cell from movie 1-15 (Figure 1b). (ZIP 1991 kb)
Supplementary Movies 3
This zipped file contains Supplementary Movies 31-40 which show the spindle morphology phenotypes represented in Figure 5. (ZIP 26408 kb)
Rights and permissions
About this article
Cite this article
Neumann, B., Walter, T., Hériché, JK. et al. Phenotypic profiling of the human genome by time-lapse microscopy reveals cell division genes. Nature 464, 721–727 (2010). https://doi.org/10.1038/nature08869
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature08869
This article is cited by
-
Cell quantification in digital contrast microscopy images with convolutional neural networks algorithm
Scientific Reports (2023)
-
ALFI: Cell cycle phenotype annotations of label-free time-lapse imaging data from cultured human cells
Scientific Data (2023)
-
Effects of aneuploidy on cell behaviour and function
Nature Reviews Molecular Cell Biology (2022)
-
The N-terminal domain of TET1 promotes the formation of dense chromatin regions refractory to transcription
Chromosoma (2022)
-
2D-GolgiTrack—a semi-automated tracking system to quantify morphological changes and dynamics of the Golgi apparatus and Golgi-derived membrane tubules
Medical & Biological Engineering & Computing (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.