Abstract
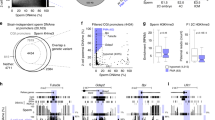
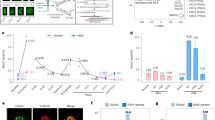
Epigenetic reprogramming including demethylation of DNA occurs in mammalian primordial germ cells (PGCs) and in early embryos, and is important for the erasure of imprints and epimutations, and the return to pluripotency1,2,3,4,5,6,7,8,9. The extent of this reprogramming and its molecular mechanisms are poorly understood. We previously showed that the cytidine deaminases AID and APOBEC1 can deaminate 5-methylcytosine in vitro and in Escherichia coli, and in the mouse are expressed in tissues in which demethylation occurs10. Here we profiled DNA methylation throughout the genome by unbiased bisulphite next generation sequencing11,12,13 in wild-type and AID-deficient mouse PGCs at embryonic day (E)13.5. Wild-type PGCs revealed marked genome-wide erasure of methylation to a level below that of methylation deficient (Np95-/- , also called Uhrf1-/- ) embryonic stem cells, with female PGCs being less methylated than male ones. By contrast, AID-deficient PGCs were up to three times more methylated than wild-type ones; this substantial difference occurred throughout the genome, with introns, intergenic regions and transposons being relatively more methylated than exons. Relative hypermethylation in AID-deficient PGCs was confirmed by analysis of individual loci in the genome. Our results reveal that erasure of DNA methylation in the germ line is a global process, hence limiting the potential for transgenerational epigenetic inheritance. AID deficiency interferes with genome-wide erasure of DNA methylation patterns, indicating that AID has a critical function in epigenetic reprogramming and potentially in restricting the inheritance of epimutations in mammals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Reik, W., Dean, W. & Walter, J. Epigenetic reprogramming in mammalian development. Science 293, 1089–1093 (2001)
Sasaki, H. & Matsui, Y. Epigenetic events in mammalian germ-cell development: reprogramming and beyond. Nature Rev. Genet. 9, 129–140 (2008)
Oswald, J. et al. Active demethylation of the paternal genome in the mouse zygote. Curr. Biol. 10, 475–478 (2000)
Mayer, W., Niveleau, A., Walter, J., Fundele, R. & Haaf, T. Demethylation of the zygotic paternal genome. Nature 403, 501–502 (2000)
Dean, W. et al. Conservation of methylation reprogramming in mammalian development: aberrant reprogramming in cloned embryos. Proc. Natl Acad. Sci. USA 98, 13734–13738 (2001)
Hajkova, P. et al. Epigenetic reprogramming in mouse primordial germ cells. Mech. Dev. 117, 15–23 (2002)
Lee, J. et al. Erasing genomic imprinting memory in mouse clone embryos produced from day 11.5 primordial germ cells. Development 129, 1807–1817 (2002)
Yamazaki, Y. et al. Reprogramming of primordial germ cells begins before migration into the genital ridge, making these cells inadequate donors for reproductive cloning. Proc. Natl Acad. Sci. USA 100, 12207–12212 (2003)
Hajkova, P. et al. Chromatin dynamics during epigenetic reprogramming in the mouse germ line. Nature 452, 877–881 (2008)
Morgan, H. D., Dean, W., Coker, H. A., Reik, W. & Petersen-Mahrt, S. K. Activation-induced cytidine deaminase deaminates 5-methylcytosine in DNA and is expressed in pluripotent tissues: implications for epigenetic reprogramming. J. Biol. Chem. 279, 52353–52360 (2004)
Cokus, S. J. et al. Shotgun bisulfite sequencing of the Arabidopsis genome reveals DNA methylation patterning. Nature 452, 215–219 (2008)
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008)
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009)
Gehring, M., Reik, W. & Henikoff, S. DNA demethylation by DNA repair. Trends Genet. 25, 82–90 (2009)
Gehring, M., Bubb, K. L. & Henikoff, S. Extensive demethylation of repetitive elements during seed development underlies gene imprinting. Science 324, 1447–1451 (2009)
Hsieh, T. F. et al. Genome-wide demethylation of Arabidopsis endosperm. Science 324, 1451–1454 (2009)
Rai, K. et al. DNA demethylation in zebrafish involves the coupling of a deaminase, a glycosylase and gadd45. Cell 135, 1201–1212 (2008)
Lane, N. et al. Resistance of IAPs to methylation reprogramming may provide a mechanism for epigenetic inheritance in the mouse. Genesis 35, 88–93 (2003)
Zvetkova, I. et al. Global hypomethylation of the genome in XX embryonic stem cells. Nature Genet. 37, 1274–1279 (2005)
Seki, T. et al. Cellular dynamics associated with the genome-wide epigenetic reprogramming in migrating primordial germ cells in mice. Development 134, 2627–2638 (2007)
Muramatsu, M. et al. Class switch recombination and hypermutation require activation-induced cytidine deaminase (AID), a potent RNA editing enzyme. Cell 102, 553–563 (2000)
Whitelaw, N. C. & Whitelaw, E. Transgenerational epigenetic inheritance in health and disease. Curr. Opin. Genet. Dev. 18, 273–279 (2008)
Slotkin, R. K. et al. Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell 136, 461–472 (2009)
Teixeira, F. K. et al. A role for RNAi in the selective correction of DNA methylation defects. Science 323, 1600–1604 (2009)
Bhutani, N. et al. Reprogramming towards pluripotency requires AID-dependent DNA demethylation. Nature 10.1038/nature08752 (in the press)
Robbiani, D. F. et al. Aid produces DNA double-strand breaks in non-Ig genes and mature B cell lymphomas with reciprocal chromosome translocations. Mol. Cell 36, 631–641 (2009)
Neuberger, M. S., Harris, R. S., Di Noia, J. & Petersen-Mahrt, S. K. Immunity through DNA deamination. Trends Biochem. Sci. 28, 305–312 (2003)
Larijani, M. et al. Methylation protects cytidines from Aid-mediated deamination. Mol. Immunol. 42, 599–604 (2005)
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–930 (2009)
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009)
Acknowledgements
We thank H. Morgan for his contributions to some of the early analysis of Aid-/- mice, A. Segonds-Pichon for help with statistical evaluation, and J. Hetzel for assisting in preparing the Illumina Solexa libraries and their sequencing. We also thank S. Petersen-Mahrt, C. Rada and F. Santos for advice and discussions. C.P. was a Boehringer-Ingelheim predoctoral Fellow. S.F. is a Howard Hughes Medical Institute Fellow of the Life Science Research Foundation. S.E.J. is an investigator of the Howard Hughes Medical Institute. This work was supported by BBSRC, MRC, EU NoE The Epigenome, and CellCentric (to W.R.), and by HHMI, NSF Plant Genome Research Programme, and NIH (to S.E.J.).
Author Contributions C.P. and W.D. isolated tissue samples and PGCs, assessed the purity of the samples and prepared DNA. C.P. undertook genetic crosses, determined weights of mouse pups and carried out Sequenom EpiTYPER analysis. S.F. constructed bisulphite libraries and did Illumina Solexa sequencing. S.J.C., S.A. and M.P. carried out mapping, base-calling and computational analyses. C.P., W.D., S.F., S.J.C., S.A., M.P., S.E.J. and W.R. analysed data. C.P., W.D., S.F., S.E.J and W.R. designed experiments; S.E.J. and W.R. designed and directed the study. C.P. and W.R. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S4 with Legends, Supplementary Methods and a Supplementary Reference (PDF 795 kb)
Rights and permissions
About this article
Cite this article
Popp, C., Dean, W., Feng, S. et al. Genome-wide erasure of DNA methylation in mouse primordial germ cells is affected by AID deficiency. Nature 463, 1101–1105 (2010). https://doi.org/10.1038/nature08829
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08829
This article is cited by
-
DMRT1-mediated reprogramming drives development of cancer resembling human germ cell tumors with features of totipotency
Nature Communications (2021)
-
Activation-induced cytidine deaminase is a possible regulator of cross-talk between oocytes and granulosa cells through GDF-9 and SCF feedback system
Scientific Reports (2021)
-
Genetic and epigenetic determinants of diffuse large B-cell lymphoma
Blood Cancer Journal (2020)
-
Characterization of DNA methylation variations during fruit development and ripening of Vitis vinifera (cv. ‘Fujiminori’)
Physiology and Molecular Biology of Plants (2020)
-
Retention of paternal DNA methylome in the developing zebrafish germline
Nature Communications (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.