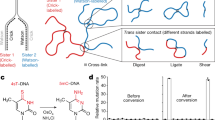
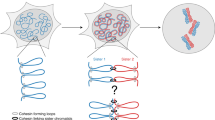
Abstract
It is generally assumed that sister chromatids are genetically and functionally identical and that segregation to daughter cells is a random process. However, functional differences between sister chromatids regulate daughter cell fate in yeast1 and sister chromatid segregation is not random in Escherichia coli2. Differentiated sister chromatids, coupled with non-random segregation, have been proposed to regulate cell fate during the development of multicellular organisms3. This hypothesis has not been tested because molecular features to reliably distinguish between sister chromatids are not obvious. Here we show that parental ‘Watson’ and ‘Crick’ DNA template strands can be identified in sister chromatids of murine metaphase chromosomes using CO-FISH (chromosome orientation fluorescence in situ hybridization4) with unidirectional probes specific for centromeric and telomeric repeats. All chromosomes were found to have a uniform orientation with the 5′ end of the short arm on the same strand as T-rich major satellite repeats. The invariable orientation of repetitive DNA was used to differentially label sister chromatids and directly study mitotic segregation patterns in different cell types. Whereas sister chromatids appeared to be randomly distributed between daughter cells in cultured lung fibroblasts and embryonic stem cells, significant non-random sister chromatid segregation was observed in a subset of colon crypt epithelial cells, including cells outside positions reported for colon stem cells5. Our results establish that DNA template sequences can be used to distinguish sister chromatids and follow their mitotic segregation in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Klar, A. J. Differentiated parental DNA strands confer developmental asymmetry on daughter cells in fission yeast. Nature 326, 466–470 (1987)
White, M. A., Eykelenboom, J. K., Lopez-Vernaza, M. A., Wilson, E. & Leach, D. R. Non-random segregation of sister chromosomes in Escherichia coli . Nature 455, 1248–1250 (2008)
Lansdorp, P. M. Immortal strands? Give me a break. Cell 129, 1244–1247 (2007)
Meyne, J. & Goodwin, E. H. Strand-specific fluorescence in situ hybridization for determining orientation and direction of DNA sequences. Methods Mol. Biol. 33, 141–145 (1994)
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5 . Nature 449, 1003–1007 (2007)
Garagna, S. et al. Pericentromeric organization at the fusion point of mouse Robertsonian translocation chromosomes. Proc. Natl Acad. Sci. USA 98, 171–175 (2001)
Lin, M. S. & Davidson, R. L. Centric fusion, satellite DNA, and DNA polarity in mouse chromosomes. Science 185, 1179–1181 (1974)
Bailey, S. M., Goodwin, E. H. & Cornforth, M. N. Strand-specific fluorescence in situ hybridization: the CO-FISH family. Cytogenet. Genome Res. 107, 14–17 (2004)
Alves, P. & Jonasson, J. New staining method for the detection of sister-chromatid exchanges in BrdU-labelled chromosomes. J. Cell Sci. 32, 185–195 (1978)
Potten, C. S., Hume, W. J., Reid, P. & Cairns, J. The segregation of DNA in epithelial stem cells. Cell 15, 899–906 (1978)
Schneider, E. L., Sternberg, H. & Tice, R. R. In vivo analysis of cellular replication. Proc. Natl Acad. Sci. USA 74, 2041–2044 (1977)
Armakolas, A. & Klar, A. J. Cell type regulates selective segregation of mouse chromosome 7 DNA strands in mitosis. Science 311, 1146–1149 (2006)
Bell, C. D. Is mitotic chromatid segregation random? Histol. Histopathol. 20, 1313–1320 (2005)
Karpowicz, P. et al. The germline stem cells of Drosophila melanogaster partition DNA non-randomly. Eur. J. Cell Biol. 88, 397–408 (2009)
Wang, X. et al. Asymmetric centrosome inheritance maintains neural progenitors in the neocortex. Nature 461, 947–955 (2009)
Yamashita, Y. M., Mahowald, A. P., Perlin, J. R. & Fuller, M. T. Asymmetric inheritance of mother versus daughter centrosome in stem cell division. Science 315, 518–521 (2007)
Etienne-Manneville, S. & Hall, A. Cdc42 regulates GSK-3β and adenomatous polyposis coli to control cell polarity. Nature 421, 753–756 (2003)
Hanson, C. A. & Miller, J. R. Non-traditional roles for the Adenomatous Polyposis Coli (APC) tumor suppressor protein. Gene 361, 1–12 (2005)
Kaplan, K. B. et al. A role for the Adenomatous Polyposis Coli protein in chromosome segregation. Nature Cell Biol. 3, 429–432 (2001)
Kita, K., Wittmann, T., Nathke, I. S. & Waterman-Storer, C. M. Adenomatous polyposis coli on microtubule plus ends in cell extensions can promote microtubule net growth with or without EB1. Mol. Biol. Cell 17, 2331–2345 (2006)
Yamashita, Y. M., Jones, D. L. & Fuller, M. T. Orientation of asymmetric stem cell division by the APC tumor suppressor and centrosome. Science 301, 1547–1550 (2003)
Jansen, L. E., Black, B. E., Foltz, D. R. & Cleveland, D. W. Propagation of centromeric chromatin requires exit from mitosis. J. Cell Biol. 176, 795–805 (2007)
Thorpe, P. H., Bruno, J. & Rothstein, R. Kinetochore asymmetry defines a single yeast lineage. Proc. Natl Acad. Sci. USA 106, 6673–6678 (2009)
Lew, D. J., Burke, D. J. & Dutta, A. The immortal strand hypothesis: how could it work? Cell 133, 21–23 (2008)
Luo, S. & Preuss, D. Strand-biased DNA methylation associated with centromeric regions in Arabidopsis . Proc. Natl Acad. Sci. USA 100, 11133–11138 (2003)
Kanellopoulou, C. et al. Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev. 19, 489–501 (2005)
Murchison, E. P., Partridge, J. F., Tam, O. H., Cheloufi, S. & Hannon, G. J. Characterization of Dicer-deficient murine embryonic stem cells. Proc. Natl Acad. Sci. USA 102, 12135–12140 (2005)
Bouzinba-Segard, H., Guais, A. & Francastel, C. Accumulation of small murine minor satellite transcripts leads to impaired centromeric architecture and function. Proc. Natl Acad. Sci. USA 103, 8709–8714 (2006)
Rudert, F., Bronner, S., Garnier, J. M. & Dolle, P. Transcripts from opposite strands of gamma satellite DNA are differentially expressed during mouse development. Mamm. Genome 6, 76–83 (1995)
Bouck, D. C. & Bloom, K. Pericentric chromatin is an elastic component of the mitotic spindle. Curr. Biol. 17, 741–748 (2007)
Ding, H. et al. Regulation of murine telomere length by Rtel: an essential gene encoding a helicase-like protein. Cell 117, 873–886 (2004)
Gertsenstein, M., Lobe, C. & Nagy, A. ES cell-mediated conditional transgenesis. Methods Mol. Biol. 185, 285–307 (2002)
Hörz, W. & Altenburger, W. Nucleotide sequence of mouse satellite DNA. Nucleic Acids Res. 9, 683–696 (1981)
Johnson, G. D. et al. Fading of immunofluorescence during microscopy: a study of the phenomenon and its remedy. J. Immunol. Methods 55, 231–242 (1982)
Allen, J. W. & Latt, S. A. Analysis of sister chromatid exchange formation in vivo in mouse spermatogonia as a new test system for environmental mutagens. Nature 260, 449–451 (1976)
Acknowledgements
K. Lisaingo helped acquire images for this study, J. Tan helped with image analysis and C. J. S. Smith assisted with in vivo studies. We thank S. Selig, T. de Lange and R. C. Wang for help with CO-FISH protocols. S. Aparicio, L. Lefebvre, L. F. Lansdorp, B. M. Lansdorp and members of the Lansdorp laboratory are thanked for discussions and for critically reading the manuscript. This study was funded in part by grants from the Canadian Institutes of Health Research (RMF-92093), the Michael Smith Foundation for Health Research, the Canadian Cancer Society Research Institute and the Terry Fox Foundation. We are grateful to the British Columbia Cancer Agency, the Canada Foundation for Innovation, the British Columbia Knowledge Development Fund, the British Columbia Cancer Foundation, the Blossom Fund of the University of British Columbia and the Mahon family for funding the infrastructure that enabled this work.
Author Contributions E.F. helped with the design of the experiments, image acquisition, data analysis, interpretation of results and writing of the paper. E.A.C. performed most of the CO-FISH experiments. A.H. performed most of the mouse work. L.B. acquired some images for this study. S.S.S.P. performed analysis of digital data and helped with statistical analysis that was performed by S.M. D.G.H. helped with the design of the study and interpretation of results. P.M.L. conceived the study, helped with image acquisition, interpretation of results and writing of the paper.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-11 with Legends, Supplementary Notes, Supplementary Video Legends 1-4, Supplementary Data Legend and Supplementary Data Image Gallery Legend. (PDF 3547 kb)
Supplementary Video 1
This movie file shows the distribution of defined DNA template strands in paired daughter cells in mouse colon analyzed by CO-FISH (see Supplementary file for full Legend). (MOV 1067 kb)
Supplementary Video 2
This movie file shows the distribution of defined DNA template strands in paired metaphase chromosomes in the mouse colon analyzed by CO-FISH (see Supplementary file for full Legend). (MOV 1692 kb)
Supplementary Video 3
This movie file shows the mirror asymmetry of defined DNA template strands in a pair of post-mitotic mouse colon cells (see Supplementary Information file for full Legend). (MOV 2248 kb)
Supplementary Video 4
This movie file shows mirror asymmetry of defined DNA template strands in a pair of post-mitotic mouse colon cells (see Supplementary Information file for full Legend). (MOV 1347 kb)
Supplementary Data
This file contains an excel spreadsheet which shows the relative fluorescence measurements extracted from two-colour digital CO-FISH images of cell pairs present in colon tissue sections, colon suspension cells, fibroblast and ES cells (see Supplementary Information file for full Legend). (XLS 133 kb)
Supplementary Image Gallery
This file contains an image gallery which shows cell pairs and surrounding tissue from statistically significant asymmetric segregation events (see Supplementary Information file for full Legend). (PDF 589 kb)
Rights and permissions
About this article
Cite this article
Falconer, E., Chavez, E., Henderson, A. et al. Identification of sister chromatids by DNA template strand sequences. Nature 463, 93–97 (2010). https://doi.org/10.1038/nature08644
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08644
This article is cited by
-
Sister chromatid–sensitive Hi-C to map the conformation of replicated genomes
Nature Protocols (2022)
-
Detecting chromatin interactions between and along sister chromatids with SisterC
Nature Methods (2020)
-
A combinatorial approach to the restriction of a mouse genome
BMC Research Notes (2013)
-
BAIT: Organizing genomes and mapping rearrangements in single cells
Genome Medicine (2013)
-
Chromosome-specific nonrandom sister chromatid segregation during stem-cell division
Nature (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.