Abstract
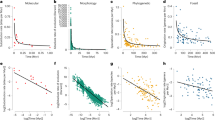
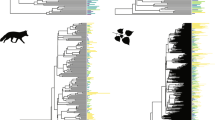
The Red Queen1 describes a view of nature in which species continually evolve but do not become better adapted. It is one of the more distinctive metaphors of evolutionary biology, but no test of its claim that speciation occurs at a constant rate2 has ever been made against competing models that can predict virtually identical outcomes, nor has any mechanism been proposed that could cause the constant-rate phenomenon. Here we use 101 phylogenies of animal, plant and fungal taxa to test the constant-rate claim against four competing models. Phylogenetic branch lengths record the amount of time or evolutionary change between successive events of speciation. The models predict the distribution of these lengths by specifying how factors combine to bring about speciation, or by describing how rates of speciation vary throughout a tree. We find that the hypotheses that speciation follows the accumulation of many small events that act either multiplicatively or additively found support in 8% and none of the trees, respectively. A further 8% of trees hinted that the probability of speciation changes according to the amount of divergence from the ancestral species, and 6% suggested speciation rates vary among taxa. By comparison, 78% of the trees fit the simplest model in which new species emerge from single events, each rare but individually sufficient to cause speciation. This model predicts a constant rate of speciation, and provides a new interpretation of the Red Queen: the metaphor of species losing a race against a deteriorating environment is replaced by a view linking speciation to rare stochastic events that cause reproductive isolation. Attempts to understand species-radiations3 or why some groups have more or fewer species should look to the size of the catalogue of potential causes of speciation shared by a group of closely related organisms rather than to how those causes combine.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Van Valen, L. A new evolutionary law. Evol. Theory 1, 1–30 (1973)
Stenseth, N. C. & Maynard Smith, J. Coevolution in ecosystems: Red Queen evolution or stasis? Evolution 38, 870–880 (1984)
Rabosky, D. L. & Lovette, I. J. Explosive evolutionary radiations: decreasing speciation or increasing extinction through time? Evolution 62, 1866–1875 (2008)
Benton, M. J. in Palaeobiology: A Synthesis (eds Briggs, D. E. G. & Crowther, P. R.) 119–124 (Blackwells, 1990)
Pearson, P. N. Survivorship analysis of fossil taxa when real-time extinction rates vary: the Paleogene planktonic foraminifera. Paleobiology 18, 115–131 (1992)
Nee, S., Mooers, A. O. & Harvey, P. H. Tempo and mode of evolution revealed from molecular phylogenies. Proc. Natl Acad. Sci. USA 89, 8322–8326 (1992)
Nee, S. Birth-death models in macroevolution. Annu. Rev. Ecol. Evol. Syst. 37, 1–17 (2006)
Pagel, M. & Meade, A. A phylogenetic mixture model for detecting pattern-heterogeneity in gene sequence or character-state data. Syst. Biol. 53, 571–581 (2004)
Venditti, C., Meade, A. & Pagel, M. Phylogenetic mixture models can reduce node-density artifacts. Syst. Biol. 57, 286–293 (2008)
Venditti, C., Meade, A. & Pagel, M. Detecting the node-density artifact in phylogeny reconstruction. Syst. Biol. 55, 637–643 (2006)
Pagel, M., Venditti, C. & Meade, A. Large punctuational contribution of speciation to evolutionary divergence at the molecular level. Science 314, 119–121 (2006)
Gillespie, D. J. H. The Causes of Molecular Evolution (Oxford Univ. Press, 1991)
Pagel, M., Meade, A. & Scott, D. Assembly rules for protein networks derived from phylogenetic-statistical analysis of whole genomes. BMC Evol. Biol. 7, S16 (2007)
Pagel, M. & Meade, A. Bayesian analysis of correlated evolution of discrete characters by reversible-jump Markov chain Monte Carlo. Am. Nat. 167, 808–825 (2006)
Raftery, A. E. in Markov Chain Monte Carlo in Practice (eds Gilks, W. R., Richardson, S. & Spiegelhalter, D. J.) 145–161 (Chapman & Hall, 1996)
Colless, D. H. Phylogenetics: the theory and practice of phylogenetic systematics II (book review). Syst. Zool. 31, 100–104 (1982)
Yule, G. U. A mathematical theory of evolution, based on the conclusions of Dr. JC Willis, FRS. Phil. Trans. R. Soc. Lond. B. 213, 21–87 (1925)
Smith, S. A. & Donoghue, M. J. Rates of molecular evolution are linked to life history in flowering plants. Science 322, 86–89 (2008)
Gillespie, J. H. The molecular clock may be an episodic clock. Proc. Natl Acad. Sci. USA 81, 8009–8013 (1984)
Coyne, J. A. & Orr, H. A. Speciation 411–442 (Sinauer Associates, 2004)
Soltis, D. E., Soltis, P. S. & Tate, J. A. Advances in the study of polyploidy since plant speciation. New Phytol. 161, 173–191 (2003)
Mitsainas, G. P., Rovatsos, M. T. H., Rizou, E. I. & Giagia-Athanasopoulou, E. Sex chromosome variability outlines the pathway to the chromosomal evolution in Microtus thomasi (Rodentia, Arvicolinae). Biol. J. Linn. Soc. 96, 685–695 (2009)
Navarro, A. & Barton, N. H. Chromosomal speciation and molecular divergence — accelerated evolution in rearranged chromosomes. Science 300, 321–324 (2003)
Orr, H. A. & Turelli, M. The evolution of postzygotic isolation: accumulating Dobzhansky-Muller incompatibilities. Evolution 55, 1085–1094 (2001)
Seehausen, O. et al. Speciation through sensory drive in cichlid fish. Nature 455, 620–626 (2008)
Mallet, J. Hybrid speciation. Nature 446, 279–283 (2007)
Green, P. J. Reversible jump Markov chain Monte Carlo computation and Bayesian model determination. Biome 82, 711–732 (1995)
Castoe, T. A. & Parkinson, C. L. Bayesian mixed models and the phylogeny of pitvipers (Viperidae: Serpentes). Mol. Phylogenet. Evol. 39, 91–110 (2006)
Cameron, S. A., Hines, H. M. & Williams, P. H. A comprehensive phylogeny of the bumble bees (Bombus). Biol. J. Linn. Soc. 91, 161–188 (2007)
Fritsch, P. W., Cruz, B. C., Almeda, F., Wang, Y. & Shi, S. Phylogeny of Symplocos based on DNA sequences of the chloroplast trnC-trnD intergenic region. Syst. Bot. 31, 181–192 (2006)
Acknowledgements
We thank the Centre for Advanced Computing and Emerging Technologies (ACET) at the University of Reading for the use of the ThamesBlue supercomputer. We thank M. Turelli for calling our attention to Gillespie’s Poisson process model and for comments on earlier drafts of the paper. M. Steel pointed out the geometric distribution proof given in the Supplementary Information. This research was supported by grants to M.P. from the Natural Environment Research Council (NERC), UK, and the Leverhulme Trust.
Author Contributions C.V., A.M. and M.P. contributed to all aspects of this work.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Methods, Supplementary Data, Supplementary Figure S1 with Legend, Supplementary Tables S1-S3, a Supplementary List for Dataset sources and Supplementary References. (PDF 806 kb)
Rights and permissions
About this article
Cite this article
Venditti, C., Meade, A. & Pagel, M. Phylogenies reveal new interpretation of speciation and the Red Queen. Nature 463, 349–352 (2010). https://doi.org/10.1038/nature08630
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08630
This article is cited by
-
4xTrifolium ambiguum and 2xT. occidentale hybridise despite wide geographic separation and polyploidisation: implications for clover breeding
Theoretical and Applied Genetics (2019)
-
Dinosaurs reveal the geographical signature of an evolutionary radiation
Nature Ecology & Evolution (2018)
-
How Future Depends on Past and Rare Events in Systems of Life
Foundations of Science (2018)
-
Speciation: Goldschmidt’s Chromosomal Heresy, Once Supported by Gould and Dawkins, is Again Reinstated
Biological Theory (2017)
-
PhyInformR: phylogenetic experimental design and phylogenomic data exploration in R
BMC Evolutionary Biology (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.