Abstract
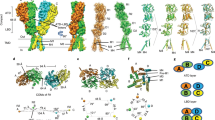
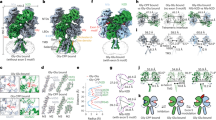
Ionotropic glutamate receptors mediate most excitatory neurotransmission in the central nervous system and function by opening a transmembrane ion channel upon binding of glutamate. Despite their crucial role in neurobiology, the architecture and atomic structure of an intact ionotropic glutamate receptor are unknown. Here we report the crystal structure of the α-amino-3-hydroxy-5-methyl-4-isoxazole propionic acid (AMPA)-sensitive, homotetrameric, rat GluA2 receptor at 3.6 Å resolution in complex with a competitive antagonist. The receptor harbours an overall axis of two-fold symmetry with the extracellular domains organized as pairs of local dimers and with the ion channel domain exhibiting four-fold symmetry. A symmetry mismatch between the extracellular and ion channel domains is mediated by two pairs of conformationally distinct subunits, A/C and B/D. Therefore, the stereochemical manner in which the A/C subunits are coupled to the ion channel gate is different from the B/D subunits. Guided by the GluA2 structure and site-directed cysteine mutagenesis, we suggest that GluN1 and GluN2A NMDA (N-methyl-d-aspartate) receptors have a similar architecture, with subunits arranged in a 1-2-1-2 pattern. We exploit the GluA2 structure to develop mechanisms of ion channel activation, desensitization and inhibition by non-competitive antagonists and pore blockers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout












Similar content being viewed by others
References
Cowan, W. M., Sudhof, T. C. & Stevens, C. F. eds. Synapses (The Johns Hopkins Univ. Press, 2001)
Hayashi, T. Effects of sodium glutamate on the nervous system. Keio J. Med. 3, 183–192 (1954)
Curtis, D. R., Phillis, J. W. & Watkins, J. C. Chemical excitation of spinal neurons. Nature 183, 611–612 (1959)
Sugiyama, H., Ito, I. & Watanabe, M. Glutamate receptor subtypes may be classified into two major categories: A study on Xenopus oocytes injected with rat brain mRNA. Neuron 3, 129–132 (1989)
Dingledine, R., Borges, K., Bowie, D. & Traynelis, S. F. The glutamate receptor ion channels. Pharmacol. Rev. 51, 7–61 (1999)
Jane, D. E., Lodge, D. & Collingridge, G. L. Kainate receptors: Pharmacology, function and therapeutic potential. Neuropharmacology 56, 90–113 (2009)
Lipton, S. A. Paradigm shift in neuroprotection by NMDA receptor blockage: memantine and beyond. Nature Rev. Drug Discov. 5, 160–170 (2006)
Alt, A., Nisenbaum, E. S., Bleakman, D. & Witkin, J. M. A role for AMPA receptors in mood disorders. Biochem. Pharmacol. 71, 1273–1288 (2006)
Labrie, V. & Roder, J. C. The involvement of the NMDA receptor d-serine/glycine site in the pathophysiology and treatment of schizophrenia. Neurosci. Biobehav. Rev. 10.1016/j.neubiorev.2009.08.002 (18 August 2009)
Boulter, J. et al. Molecular cloning and functional expression of glutamate receptor subunit genes. Science 249, 1033–1037 (1990)
Keinänen, K. et al. A family of AMPA-selective glutamate receptors. Science 249, 556–560 (1990)
Sommer, B. et al. A glutamate receptor channel with high affinity for domoate and kainate. EMBO J. 11, 1651–1656 (1992)
Moriyoshi, K. et al. Molecular cloning and characterization of the rat NMDA receptor. Nature 354, 31–37 (1991)
Monyer, H. et al. Heteromeric NMDA receptors: molecular and functional distinction of subtypes. Science 256, 1217–1221 (1992)
Rosenmund, C., Stern-Bach, Y. & Stevens, C. F. The tetrameric structure of a glutamate receptor channel. Science 280, 1596–1599 (1998)
Lu, W. et al. Subunit composition of synaptic AMPA receptors revealed by a single-cell genetic approach. Neuron 62, 254–268 (2009)
Mulle, C. et al. Subunit composition of kainate receptors in hippocampal interneurons. Neuron 28, 475–484 (2000)
Christensen, J. K., Paternain, A. V., Selak, S., Ahring, P. K. & Lerma, J. A mosaic of functional kainate receptors in hippocampal interneurons. J. Neurosci. 24, 8986–8993 (2004)
Wo, Z. G. & Oswald, R. E. Unraveling the modular design of glutamate-gated ion channels. Trends Neurosci. 18, 161–168 (1995)
O’Hara, P. J. et al. The ligand-binding domain in metabotropic glutamate receptors is related to bacterial periplasmic binding proteins. Neuron 11, 41–52 (1993)
Stern-Bach, Y. et al. Agonist selectivity of glutamate receptors is specified by two domains structurally related to bacterial amino acid-binding proteins. Neuron 13, 1345–1357 (1994)
Wollmuth, L. P. & Sobolevsky, A. I. Structure and gating of the glutamate receptor ion channel. Trends Neurosci. 27, 321–328 (2004)
Soderling, T. R. & Derkach, V. A. Postsynaptic protein phosphorylation and LTP. Trends Neurosci. 23, 75–80 (2000)
Erreger, K., Chen, P. E., Wyllie, D. J. & Traynelis, S. F. Glutamate receptor gating. Crit. Rev. Neurobiol. 16, 187–224 (2004)
Hansen, K. B., Yuan, H. & Traynelis, S. F. Structural aspects of AMPA receptor activation, desensitization and deactivation. Curr. Opin. Neurobiol. 17, 281–288 (2007)
Paoletti, P. & Neyton, J. NMDA receptor subunits: function and pharmacology. Curr. Opin. Pharmacol. 7, 39–47 (2007)
Mayer, M. L., Westbrook, G. L. & Guthrie, P. B. Voltage-dependent block by Mg2+ of NMDA responses in spinal cord neurones. Nature 309, 261–263 (1984)
Johnson, J. W. & Ascher, P. Glycine potentiates the NMDA response in cultured mouse brain neurons. Nature 325, 529–531 (1987)
Smith, T. C. & Howe, J. R. Concentration-dependent substate behavior of native AMPA receptors. Nature Neurosci. 3, 992–997 (2000)
Klein, R. M. & Howe, J. R. Effects of the lurcher mutation on GluR1 desensitization and activation kinetics. J. Neurosci. 24, 4941–4951 (2004)
Vyklicky, L., Benveniste, M. & Mayer, M. L. Modulation of N-methyl-d-aspartic acid receptor desensitization by glycine in mouse cultured hippocampal neurones. J. Physiol. 428, 313–331 (1990)
Balannik, V., Menniti, F. S., Paternain, A. V., Lerma, J. & Stern-Bach, Y. Molecular mechanisms of AMPA receptor noncompetitive antagonism. Neuron 48, 279–288 (2005)
Mony, L., Kew, J. N., Gunthorpe, M. J. & Paoletti, P. Allosteric modulators of NR2B-containing NMDA receptors: molecular mechanisms and therapeutic potential. Br. J. Pharmacol. 157, 1301–1317 (2009)
Sun, Y. et al. Mechanism of glutamate receptor desensitization. Nature 417, 245–253 (2002)
Jin, R. et al. Mechanism of positive allosteric modulators acting on AMPA receptors. J. Neurosci. 25, 9027–9036 (2005)
Sobolevsky, A. I. Channel block of glutamate receptors. In Recent Research Developments in Physiology Vol. 1, Part I (ed. Pandalai, S. G.) 1–38 (Research Signpost, 2003)
Kuusinen, A., Abele, R., Madden, D. R. & Keinänen, K. Oligomerization and ligand-binding properties of the ectodomain of the α-amino-3-hydroxy-5-methyl-4-isoxazole propionic acid receptor subunit GluRD. J. Biol. Chem. 274, 28937–28943 (1999)
Armstrong, N. & Gouaux, E. Mechanisms for activation and antagonism of an AMPA-sensitive glutamate receptor: Crystal structures of the GluR2 ligand binding core. Neuron 28, 165–181 (2000)
Jin, R. et al. Crystal structure and association behavior of the GluR2 amino-terminal domain. EMBO J. 28, 1812–1823 (2009)
Kumar, J., Schuck, P., Jin, R. & Mayer, M. L. The N-terminal domain of GluR6-subtype glutamate receptor ion channels. Nature Struct. Mol. Biol. 16, 631–638 (2009)
Tichelaar, W., Safferling, M., Keinänen, K., Stark, H. & Madden, D. R. The three-dimensional structure of an ionotropic glutamate receptor reveals a dimer-of-dimers assembly. J. Mol. Biol. 344, 435–442 (2004)
Nakagawa, T., Cheng, Y., Ramm, E., Sheng, M. & Walz, T. Structure and different conformational states of native AMPA receptor complexes. Nature 433, 545–549 (2005)
Chen, G.-Q., Cui, C., Mayer, M. & Gouaux, E. Functional characterization of a potassium-selective prokaryotic glutamate receptor. Nature 402, 817–821 (1999)
Mansour, M., Nagarajan, N., Nehring, R. B., Clements, J. D. & Rosenmund, C. Heteromeric AMPA receptors assemble with a preferred subunit stoichiometry and spatial arrangement. Neuron 32, 841–853 (2001)
Schorge, S. & Colquhoun, D. Studies of NMDA receptor function and stoichiometry with truncated and tandem subunits. J. Neurosci. 23, 1151–1158 (2003)
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006)
Sommer, B., Köhler, M., Sprengel, R. & Seeburg, P. H. RNA editing in brain controls a determinant of ion flow in glutamate-gated channels. Cell 67, 11–19 (1991)
Sommer, B. et al. Flip and flop: A cell-specific functional switch in glutamate-operated channels of the CNS. Science 249, 1580–1585 (1990)
Turski, L. et al. ZK200775: a phosphonate quinoxalinedione AMPA antagonist for neuroprotection in stroke and trauma. Proc. Natl Acad. Sci. USA 95, 10960–10965 (1998)
Mayer, M. L. & Armstrong, N. Structure and function of glutamate receptor ion channels. Annu. Rev. Physiol. 66, 161–181 (2004)
Horning, M. & Mayer, M. Regulation of AMPA receptor gating by ligand binding core dimers. Neuron 41, 379–388 (2004)
Hollmann, M., Maron, C. & Heinemann, S. N-Glycosylation site tagging suggests a three transmembrane domain topology for the glutamate receptor GluR1. Neuron 13, 1331–1343 (1994)
Wo, Z. G. & Oswald, R. E. Transmembrane topology of two kainate receptor subunits revealed by N-glycosylation. Proc. Natl Acad. Sci. USA 91, 7154–7158 (1994)
Doyle, D. A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998)
Weston, M. C., Schuck, P., Ghosal, A., Rosenmund, C. & Mayer, M. L. Conformational restriction blocks glutamate receptor desensitization. Nature Struct. Mol. Biol. 13, 1120–1127 (2006)
Furukawa, H., Singh, S. K., Mancusso, R. & Gouaux, E. Subunit arrangement and function in NMDA receptors. Nature 438, 185–192 (2005)
Gielen, M. et al. Structural rearrangements of NR1/NR2A NMDA receptors during allosteric inhibition. Neuron 57, 80–93 (2008)
Armstrong, N., Jasti, J., Beich-Frandsen, M. & Gouaux, E. Measurement of conformational changes accompanying desensitization in an ionotropic glutamate receptor. Cell 127, 85–97 (2006)
Chang, H.-R. & Kuo, C.-C. The activation gate and gating mechanism of the NMDA receptor. J. Neurosci. 28, 1546–1556 (2008)
Kashiwabuchi, N. et al. Impairment of motor coordination, Purkinje cell synapse formation, and cerebellar long-term depression in GluRδ2 mutant mice. Cell 81, 245–252 (1995)
Zuo, J. et al. Neurodegeneration in Lurcher mice caused by mutation in δ2 glutamate receptor gene. Nature 388, 769–773 (1997)
Yelshansky, M. V., Sobolevsky, A. I., Jatzke, C. & Wollmuth, L. P. Block of AMPA receptor desensitization by a point mutation outside the ligand-binding domain. J. Neurosci. 24, 4728–4736 (2004)
Panchenko, V. A., Glasser, C. R. & Mayer, M. L. Structural similarities between glutamate receptor channels and K+ channels examined by scanning mutagenesis. J. Gen. Physiol. 117, 345–360 (2001)
Kuner, T., Seeburg, P. H. & Guy, H. R. A common architecture for K+ channels and ionotropic glutamate receptors? Trends Neurosci. 26, 27–32 (2003)
Long, S. B., Campbell, E. B. & MacKinnon, R. Crystal structure of a mammalian voltage-dependent Shaker family K+ channel. Science 309, 897–903 (2005)
Chen, L. et al. Stargazin regulates synaptic targeting of AMPA receptors by two distinct mechanisms. Nature 408, 936–943 (2000)
Schwenk, J. et al. Functional proteomics identify cornichon proteins as auxiliary subunits of AMPA receptors. Science 323, 1313–1319 (2009)
Soto, D., Coombs, I. D., Kelly, L., Farrant, M. & Cull-Candy, S. G. Stargazin attenuates intracellular polyamine block of calcium-permeable AMPA receptors. Nature Neurosci. 10, 1260–1267 (2007)
Sobolevsky, A. I., Yelshansky, M. V. & Wollmuth, L. P. Different gating mechanisms in glutamate receptor and K+ channels. J. Neurosci. 23, 7559–7568 (2003)
Marquez-Klaka, B., Rettinger, J., Bhargava, Y., Eisele, T. & Nicke, A. Identification of an intersubunit cross-link between substituted cysteine residues located in the putative ATP binding site of the P2X1 receptor. J. Neurosci. 27, 1456–1466 (2007)
Pedersen, S. E. & Cohen, J. B. d-Turbocurarine binding sites are located at α-γ and α-δ subunit interfaces of the nicotinic acetylcholine receptor. Proc. Natl Acad. Sci. USA 87, 2785–2789 (1990)
Kuusinen, A., Arvola, M. & Keinänen, K. Molecular dissection of the agonist binding site of an AMPA receptor. EMBO J. 14, 6327–6332 (1995)
Armstrong, N., Sun, Y., Chen, G.-Q. & Gouaux, E. Structure of a glutamate receptor ligand binding core in complex with kainate. Nature 395, 913–917 (1998)
Mayer, M. L. Crystal structures of the GluR5 and GluR6 ligand binding cores: molecular mechanisms underlying kainate receptor selectivity. Neuron 45, 539–552 (2005)
Inanobe, A., Furukawa, H. & Gouaux, E. Mechanism of partial agonist action at the NR1 subunit of NMDA receptors. Neuron 47, 71–84 (2005)
Yao, Y., Harrison, C. B., Freddolino, P. L., Schulten, K. & Mayer, M. L. Molecular mechanism of ligand recognition by NR3 subtype glutamate receptors. EMBO J. 27, 2158–2170 (2008)
Jin, R., Horning, M., Mayer, M. L. & Gouaux, E. Mechanism of activation and selectivity in a ligand-gated ion channel: structural and functional studies of GluR2 and quisqualate. Biochemistry 41, 15635–15643 (2002)
Sobolevsky, A. I., Yelshansky, M. V. & Wollmuth, L. P. The outer pore of the glutamate receptor channel has 2-fold rotational symmetry. Neuron 41, 367–378 (2004)
Plested, A. J. & Mayer, M. L. AMPA receptor ligand binding domain mobility revealed by functional cross linking. J. Neurosci. 29, 11912–11923 (2009)
Gielen, M., Siegler Retchless, B., Mony, L., Johnson, J. W. & Paoletti, P. Mechanism of differential control of NMDA receptor activity by NR2 subunits. Nature 459, 703–707 (2009)
Hollmann, M., O’Shea-Greenfield, A., Rogers, S. W. & Heinemann, S. Cloning by functional expression of a member of the glutamate receptor family. Nature 342, 643–648 (1989)
Cormack, B. P., Valdivia, R. H. & Falkow, S. FACS-optimized mutants of the green fluorescent protein (GFP). Gene 173, 33–38 (1996)
Miu, P. et al. Novel AMPA receptor potentiators LY392098 and LY404187: effects on recombinant human AMPA receptors in vitro . Neuropharm. 40, 976–983 (2001)
Matthews, B. W. Solvent content of protein crystals. J. Mol. Biol. 33, 491–497 (1968)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
McCoy, A. J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D. 63, 32–41 (2007)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. 60, 2126–2132 (2004)
Adams, P. D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D. 58, 1948–1954 (2002)
Chen, G. Q. & Gouaux, E. Overexpression of a glutamate receptor (GluR2) ligand binding domain in Escherichia coli: Application of a novel protein folding screen. Proc. Natl Acad. Sci. USA 94, 13431–13436 (1997)
Acknowledgements
We thank the personnel at beamlines 5.0.2, 8.2.1 and 8.2.2 of the Advanced Light Source and at beamline 24-ID-E of the Advanced Photon Source. We also thank T. Homrichhausen for help with cloning and FSEC screening; L. Vaskalis for assistance with illustrations; and Gouaux laboratory members for discussion. M.P.R. was supported by an individual NIH National Research Service Award. This work was supported by the NIH. E.G. is an investigator with the Howard Hughes Medical Institute.
Author Contributions All authors contributed to writing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Tables 1-2, Supplementary Figures 1-34 with Legends and Supplementary References. (PDF 5544 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Sobolevsky, A., Rosconi, M. & Gouaux, E. X-ray structure, symmetry and mechanism of an AMPA-subtype glutamate receptor. Nature 462, 745–756 (2009). https://doi.org/10.1038/nature08624
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08624
This article is cited by
-
The structural basis of divalent cation block in a tetrameric prokaryotic sodium channel
Nature Communications (2023)
-
GSG1L-containing AMPA receptor complexes are defined by their spatiotemporal expression, native interactome and allosteric sites
Nature Communications (2023)
-
Two gates mediate NMDA receptor activity and are under subunit-specific regulation
Nature Communications (2023)
-
Uncovering the Significance of STEP61 in Alzheimer’s Disease: Structure, Substrates, and Interactome
Cellular and Molecular Neurobiology (2023)
-
Novel neuroactive steroids as positive allosteric modulators of NMDA receptors: mechanism, site of action, and rescue pharmacology on GRIN variants associated with neurological conditions
Cellular and Molecular Life Sciences (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.